|
|
| (14 intermediate revisions not shown) |
| Line 137: |
Line 137: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | June 24, 2009
| + | JUNE 24, 2009 |
| | </div> | | </div> |
| | <br> | | <br> |
| Line 157: |
Line 157: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You’re Doing
| + | Sequencing |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | * Sent both mutated and non-mutated <i>luxCDABE</i> for sequencing using Sp6 and T7 primers. The mutation site was sequenced using a primer that was designed to detect the mutated site. |
| | + | |
| | <html> | | <html> |
| | </div> | | </div> |
| Line 187: |
Line 188: |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | <br>Bridging the Gap between Stochastic and Deterministic Regimes in the Kinetic Simulations of the Biochemical Reaction Networks
| + | *Bridging the Gap between Stochastic and Deterministic Regimes in the Kinetic Simulations of the Biochemical Reaction Networks |
| - | <br>Efficient Exact Stochastic Simulation of Chemical Systems with Many Species and Many
| + | *Efficient Exact Stochastic Simulation of Chemical Systems with Many Species and Many |
| | Channels | | Channels |
| - | <br>Modelling and Simulation of IntraCellular
| + | *Modelling and Simulation of IntraCellular |
| | Dynamics: Choosing an Appropriate Framework | | Dynamics: Choosing an Appropriate Framework |
| - |
| |
| | | | |
| | <html> | | <html> |
| Line 221: |
Line 221: |
| | *Restriction digest products were mixed with 10x Orange Dye and visualized on a 1% agarose gel with Generuler 1.0 KB+ DNA Ladder and uncut plasmid of the same colonies as a negative conttrol. Gel photo is below. | | *Restriction digest products were mixed with 10x Orange Dye and visualized on a 1% agarose gel with Generuler 1.0 KB+ DNA Ladder and uncut plasmid of the same colonies as a negative conttrol. Gel photo is below. |
| | *Lanes 1,3,5 and 7 are the restriction digest products for colonies 1-4 and lanes 2,4,6 and 8 are uncut plasmid for colonies 1-4. | | *Lanes 1,3,5 and 7 are the restriction digest products for colonies 1-4 and lanes 2,4,6 and 8 are uncut plasmid for colonies 1-4. |
| - | *Analysis: There seems to have been no cutting with NotI as the band sin the digested lanes are the same as the bands in the uncut lanes. There are also some strange bands at about 12 KB which are much too big to be LuxOD47E. It looks like my gene may be lacking the NotI restriction sites, which indiactes that the BBK Ampliification Gradient PCR to Biobrick my gene if interest may not have been successful. We will proceed by returning to this PCR nad repeating it. | + | <Br> |
| | + | [[Image:2009.06.24VerDig.jpg|350px]] |
| | + | *Analysis: There seems to have been no cutting with NotI as the band sin the digested lanes are the same as the bands in the uncut lanes. There are also some strange bands at about 12 KB which are much too big to be LuxOD47E. It looks like my gene may be lacking the NotI restriction sites, which indicates that the BBK Ampliification Gradient PCR to Biobrick my gene if interest may not have been successful. We will proceed by returning to this PCR and repeating it. |
| | *Repeated PCR, see protocol on June 6th 2009. | | *Repeated PCR, see protocol on June 6th 2009. |
| | *PCR Products were visualized on a 1% agarose gel run at 120 V with 1.0 KB GeneRuler DNA Ladder. See gel photo below. | | *PCR Products were visualized on a 1% agarose gel run at 120 V with 1.0 KB GeneRuler DNA Ladder. See gel photo below. |
| | *Lanes 1-12 contain LuxOD47E in pCR2.1-TOPO vector. Lane 13 is a negative control with only Mastermix + ddH2O. | | *Lanes 1-12 contain LuxOD47E in pCR2.1-TOPO vector. Lane 13 is a negative control with only Mastermix + ddH2O. |
| | + | <Br> |
| | + | [[Image:2009.06.25BBKAmpPCR.jpg|350px]] |
| | *From this gel, it looks like our gene if interest, LuxOD47E is there as we see bads around 1.4 KB, as expected (gene is 1362 base pairs). We will proceed by isoltaing plasmid from the PCR product and performing a NotI Verification digest. | | *From this gel, it looks like our gene if interest, LuxOD47E is there as we see bads around 1.4 KB, as expected (gene is 1362 base pairs). We will proceed by isoltaing plasmid from the PCR product and performing a NotI Verification digest. |
| - | | + | *So I pooled the DNA from the BBK amplification PCR. Lanes: 2,3,9,11 and 12 went into tube 1 and lanes 4,5,6,7 and 8 went into tube 2. I did a PCR product isolation (Sigma) according to the manufacturer's instructions and took the concentration using the Nanodrop Spectrophotometer. |
| - | <html>
| + | |
| - | </div>
| + | |
| - | </td>
| + | |
| - | </tr>
| + | |
| - | | + | |
| - | <tr>
| + | |
| - | <td height="10px" bgcolor="#222222">
| + | |
| - | </td>
| + | |
| - | </tr>
| + | |
| - | | + | |
| - | <tr>
| + | |
| - | <td>
| + | |
| - | <a name="Fahd"></a>
| + | |
| - | <br>
| + | |
| - | <div class="heading">
| + | |
| - | FAHD
| + | |
| - | </div>
| + | |
| - | <br>
| + | |
| - | <div class="heading">
| + | |
| - | Descriptive Title of What You're Doing
| + | |
| - | </div>
| + | |
| - | <br>
| + | |
| - | <div class="desc">
| + | |
| - | </html>
| + | |
| - | WIKI CODING HERE
| + | |
| - | | + | |
| - | <html>
| + | |
| - | </div>
| + | |
| - | </td>
| + | |
| - | </tr>
| + | |
| - | | + | |
| - | <tr>
| + | |
| - | <td height="10px" bgcolor="#222222">
| + | |
| - | </td>
| + | |
| - | </tr>
| + | |
| - | | + | |
| - | <tr>
| + | |
| - | <td>
| + | |
| - | <a name="Iman"></a>
| + | |
| - | <br>
| + | |
| - | <div class="heading">
| + | |
| - | IMAN
| + | |
| - | </div>
| + | |
| - | <br>
| + | |
| - | <div class="heading">
| + | |
| - | Descriptive Title of What You're Doing
| + | |
| - | </div>
| + | |
| - | <br>
| + | |
| - | <div class="desc">
| + | |
| - | </html>
| + | |
| - | WIKI CODING HERE
| + | |
| | | | |
| | <html> | | <html> |
| Line 298: |
Line 251: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Construction of B0015-R0040-luxOU-B0015 |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | Final construction was carried out using BBa_B0015-BBa_R0040 C1 in psBAK3 and luxOU(XbaI)-BBa_B0015 in psB1AC3 C4. B0015-R0040 was digested using ''EcoR''I with ''Spe''I and luxOU(XbaI)-B0015 was digested using ''EcoR''I with ''Xba''I (Invitrogen, CA). Standard phosphatase treatment, ligation and transformation. |
| | | | |
| | <html> | | <html> |
| Line 324: |
Line 277: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Descriptive Title of What You're Doing
| + | Sending LuxPQ in psB1AK3 for DNA Sequencing |
| | </div> | | </div> |
| | <br> | | <br> |
| | <div class="desc"> | | <div class="desc"> |
| | </html> | | </html> |
| - | WIKI CODING HERE
| + | Purpose: To verify presence of LuxPQ in psB1AK3. LuxPQ-C4-2 was sent to University of Calgary DNA Sequencing with the BBK-CP-F/R primers. |
| - | | + | |
| - | <html>
| + | |
| - | </div>
| + | |
| - | </td>
| + | |
| - | </tr>
| + | |
| - | | + | |
| - | <tr>
| + | |
| - | <td height="10px" bgcolor="#222222">
| + | |
| - | </td>
| + | |
| - | </tr>
| + | |
| - | | + | |
| - | <tr>
| + | |
| - | <td>
| + | |
| - | <a name="Katie"></a>
| + | |
| - | <br>
| + | |
| - | <div class="heading">
| + | |
| - | KATIE
| + | |
| - | </div>
| + | |
| - | <br>
| + | |
| - | <div class="heading">
| + | |
| - | Descriptive Title of What You're Doing
| + | |
| - | </div>
| + | |
| - | <br>
| + | |
| - | <div class="desc">
| + | |
| - | </html>
| + | |
| - | WIKI CODING HERE
| + | |
| | | | |
| | <html> | | <html> |
| Line 384: |
Line 311: |
| | <br><br> | | <br><br> |
| | [[Image:Calgary_2009.06.24.KT1144_pBLUE1+2.png|500px]] | | [[Image:Calgary_2009.06.24.KT1144_pBLUE1+2.png|500px]] |
| - | <html>
| |
| - | </div>
| |
| - | </td>
| |
| - | </tr>
| |
| - |
| |
| - | <tr>
| |
| - | <td height="10px" bgcolor="#222222">
| |
| - | </td>
| |
| - | </tr>
| |
| - |
| |
| - | <tr>
| |
| - | <td>
| |
| - | <a name="Mandy"></a>
| |
| - | <br>
| |
| - | <div class="heading">
| |
| - | MANDY
| |
| - | </div>
| |
| - | <br>
| |
| - | <div class="heading">
| |
| - | Descriptive Title of What You're Doing
| |
| - | </div>
| |
| - | <br>
| |
| - | <div class="desc">
| |
| - | </html>
| |
| - | WIKI CODING HERE
| |
| - |
| |
| - | <html>
| |
| - | </div>
| |
| - | </td>
| |
| - | </tr>
| |
| - |
| |
| - | <tr>
| |
| - | <td height="10px" bgcolor="#222222">
| |
| - | </td>
| |
| - | </tr>
| |
| - |
| |
| - | <tr>
| |
| - | <td>
| |
| - | <a name="Patrick"></a>
| |
| - | <br>
| |
| - | <div class="heading">
| |
| - | PATRICK
| |
| - | </div>
| |
| - | <br>
| |
| - | <div class="heading">
| |
| - | Descriptive Title of What You're Doing
| |
| - | </div>
| |
| - | <br>
| |
| - | <div class="desc">
| |
| - | </html>
| |
| - | WIKI CODING HERE
| |
| | | | |
| | <html> | | <html> |
| Line 455: |
Line 331: |
| | <br> | | <br> |
| | <div class="heading"> | | <div class="heading"> |
| - | Bacterial Transformation | + | Bacterial Transformation & Response circuit |
| | </div> | | </div> |
| | <br> | | <br> |
| Line 471: |
Line 347: |
| | </tr> | | </tr> |
| | | | |
| - | <tr>
| |
| - | <td height="10px" bgcolor="#222222">
| |
| - | </td>
| |
| - | </tr>
| |
| - |
| |
| - | <tr>
| |
| - | <td>
| |
| - | <a name="Stefan"></a>
| |
| - | <br>
| |
| - | <div class="heading">
| |
| - | STEFAN
| |
| - | </div>
| |
| - | <br>
| |
| - | <div class="heading">
| |
| - | Descriptive Title of What You're Doing
| |
| - | </div>
| |
| - | <br>
| |
| - | <div class="desc">
| |
| - | </html>
| |
| - | WIKI CODING HERE
| |
| - |
| |
| - | <html>
| |
| - | </div>
| |
| - | </td>
| |
| - | </tr>
| |
| | | | |
| | <tr> | | <tr> |
|
|
CAROL
Sequencing
- Sent both mutated and non-mutated luxCDABE for sequencing using Sp6 and T7 primers. The mutation site was sequenced using a primer that was designed to detect the mutated site.
|
|
|
CHINMOYEE
More Research into Stochasticity
- Bridging the Gap between Stochastic and Deterministic Regimes in the Kinetic Simulations of the Biochemical Reaction Networks
- Efficient Exact Stochastic Simulation of Chemical Systems with Many Species and Many
Channels
- Modelling and Simulation of IntraCellular
Dynamics: Choosing an Appropriate Framework
|
|
|
EMILY
Going Back to Biobricking LuxOD47E
- Restriction digest products were mixed with 10x Orange Dye and visualized on a 1% agarose gel with Generuler 1.0 KB+ DNA Ladder and uncut plasmid of the same colonies as a negative conttrol. Gel photo is below.
- Lanes 1,3,5 and 7 are the restriction digest products for colonies 1-4 and lanes 2,4,6 and 8 are uncut plasmid for colonies 1-4.
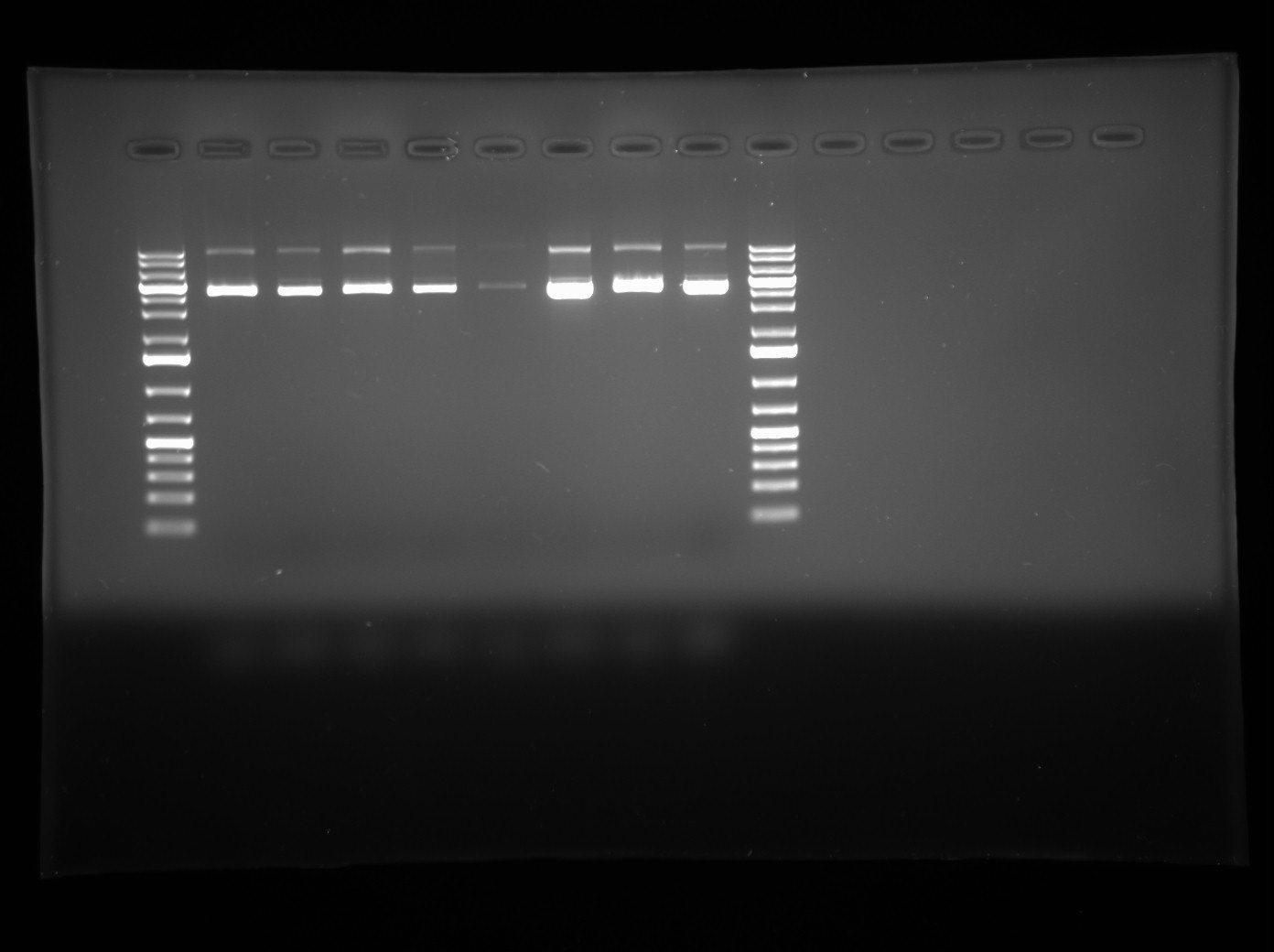
- Analysis: There seems to have been no cutting with NotI as the band sin the digested lanes are the same as the bands in the uncut lanes. There are also some strange bands at about 12 KB which are much too big to be LuxOD47E. It looks like my gene may be lacking the NotI restriction sites, which indicates that the BBK Ampliification Gradient PCR to Biobrick my gene if interest may not have been successful. We will proceed by returning to this PCR and repeating it.
- Repeated PCR, see protocol on June 6th 2009.
- PCR Products were visualized on a 1% agarose gel run at 120 V with 1.0 KB GeneRuler DNA Ladder. See gel photo below.
- Lanes 1-12 contain LuxOD47E in pCR2.1-TOPO vector. Lane 13 is a negative control with only Mastermix + ddH2O.
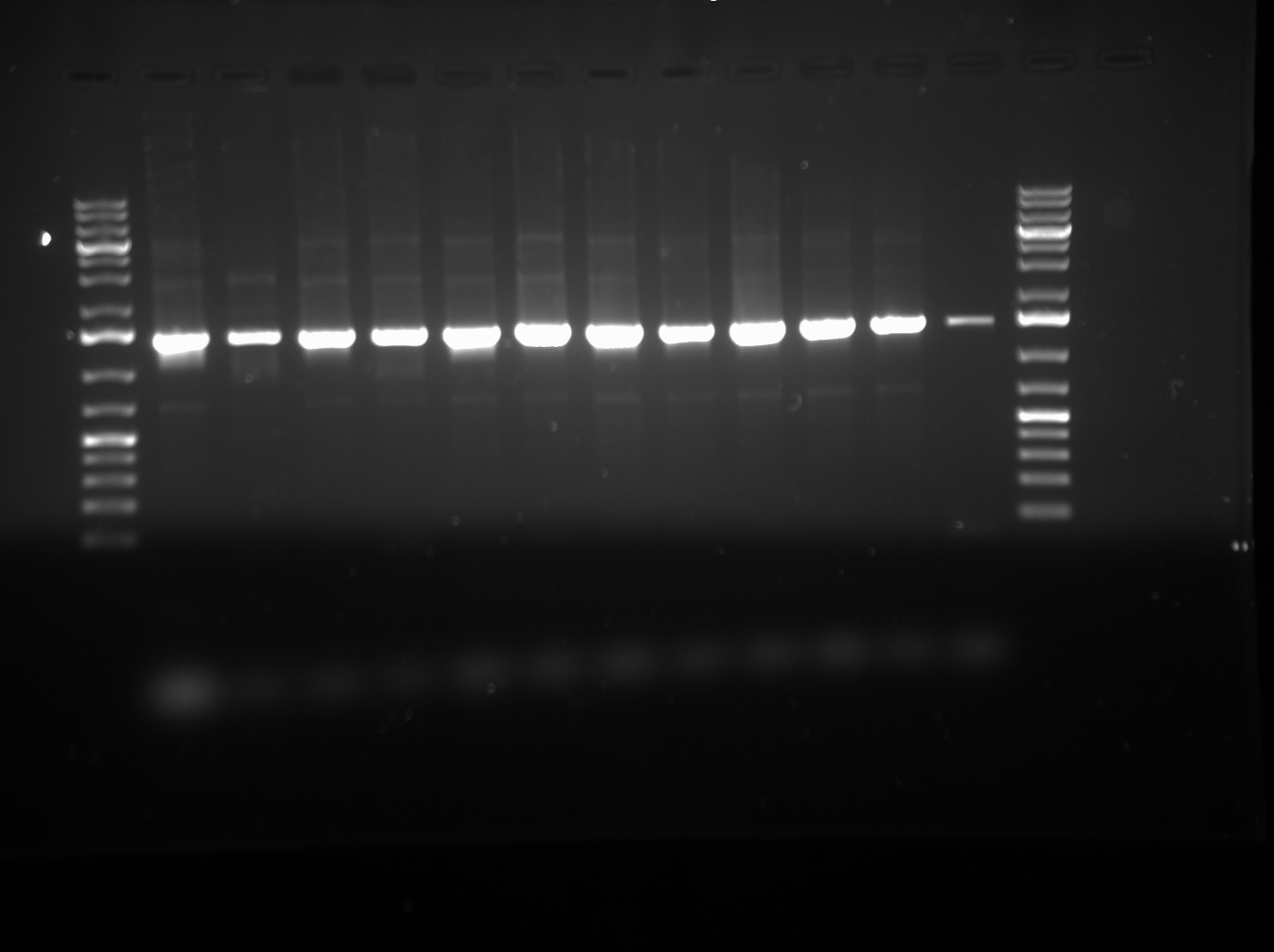
- From this gel, it looks like our gene if interest, LuxOD47E is there as we see bads around 1.4 KB, as expected (gene is 1362 base pairs). We will proceed by isoltaing plasmid from the PCR product and performing a NotI Verification digest.
- So I pooled the DNA from the BBK amplification PCR. Lanes: 2,3,9,11 and 12 went into tube 1 and lanes 4,5,6,7 and 8 went into tube 2. I did a PCR product isolation (Sigma) according to the manufacturer's instructions and took the concentration using the Nanodrop Spectrophotometer.
|
|
|
JAMIE
Construction of B0015-R0040-luxOU-B0015
Final construction was carried out using BBa_B0015-BBa_R0040 C1 in psBAK3 and luxOU(XbaI)-BBa_B0015 in psB1AC3 C4. B0015-R0040 was digested using EcoRI with SpeI and luxOU(XbaI)-B0015 was digested using EcoRI with XbaI (Invitrogen, CA). Standard phosphatase treatment, ligation and transformation.
|
|
|
JEREMY
Sending LuxPQ in psB1AK3 for DNA Sequencing
Purpose: To verify presence of LuxPQ in psB1AK3. LuxPQ-C4-2 was sent to University of Calgary DNA Sequencing with the BBK-CP-F/R primers.
|
|
|
KEVIN
Transformation
In order to verify the competency of KT1144 cells, I have transformed pBluescript into them. Competent KT1144 cells are needed to test the mutant circuits.

|
|
|
PRIMA
Bacterial Transformation & Response circuit
I shadowed Jeremy today. He was transforming the BBK ligated vector to competent cells (Top 10). I'm not sure exactly what he was transforming but I just hung out to learn the procedures and purposes of transformation.
Jamie and I were working on possible applications of the response circuit. We proposed that we should have bacteria produce wintergreen smell in response to stinky odor. The Wintergreen gene already exists in the registry so we could use a sulfur sensitive promoter in bacteria which would bind to sigma 54 and together bind to Pqrr4 promoter. Since Pqrr4 initiates the response circuit, further down the line, we could add the Winter green gene which would then be activated as sigma 54 latches on the Pqrr4.
I followed up with a few companies. BioAlberta has generously donated $1000.00 to the team. I wrote a Thank-You letter on behalf of the team and sent it to the President of BioAlberta.
|
|
|
VICKI
A sexy circuit discussion
Fahd and I worked together to propose a possible response circuit. We investigated the possiblity of using parts from the registry that can form a bioplastic, which could be useful for encapsulating cells that need to be inserted into the human body to - say - degrade a biofilm, but need to resist an immune and inflammatory response. Some considerations for our next discussion on this front will be whether or not this part works, and if so, what form the bioplastic comes in.
|

 "
"













