Team:Washington/Project/Display
From 2009.igem.org
(→Streptavidin-Biotin Binding Is Not Visualized By Fluorescence Microscopy) |
|||
| (286 intermediate revisions not shown) | |||
| Line 2: | Line 2: | ||
{{Template:Team:Washington/Templates/Header}} | {{Template:Team:Washington/Templates/Header}} | ||
| - | + | <html><style> | |
| - | + | #uw_himg { | |
| - | + | display:none; | |
| - | + | } | |
| - | + | </style></html> | |
| + | <div> | ||
| + | <div style="float:left;">'''[https://2009.igem.org/Team:Washington/Project < Project Description]'''</div> | ||
| + | <div style="float:right;">'''[https://2009.igem.org/Team:Washington/Project/Release Release >]'''</div> | ||
| + | </div> | ||
| + | [[Image:Main_graphic3_display_banner.png|center]] | ||
| + | ='''Background'''= | ||
| + | Our system hinged on finding a protein which could bind other proteins to the outside of the cell, but whose interaction with these proteins was weak enough to be disrupted by another small molecule. streptavidin presented itself as a logical choice. Several protein tags (such as the Nano-Tag<sup>1</sup>) have been developed to bind streptavidin at the biotin binding site, but can be released when biotin is added to the system and binds to streptavidin. The ability for peptides to bind streptavidin and be released upon the addition of biotin is a technology currently used on a daily basis world-wide by labs purifying proteins. The major difference between the traditional system and our system is that streptavidin is attached to beads which are used in a column format in the traditional protein purification method. Our system will have streptavidin attached to the surface of the cell. Combining the display system with our target and secretion vector our vision is complete. A single cell can then produce, secrete, bind, and release any protein of interest. | ||
| + | {| cellpadding="1" cellspacing="8" | ||
| + | |- | ||
| + | |[[Image:Strept-biotin.png | 175px]] | ||
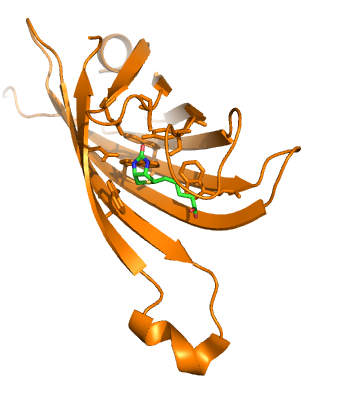
| + | |The Nano-Tag binds streptavidin with 4nM binding affinity and has been used to purify multiple proteins. The ability of biotin to compete off the Nano-Tag in biding assays indicates that the binding occurs at the same site of streptavidin. Of great importance for incorporating streptavidin into our display system is that streptavidin is a common protein used in bio-chemical assays. Its ability to bind biotin is one of the strongest non-covalent interactions known. This allows for the utilization of biotin and streptavidin to act as molecular connectors in protein systems. Furthermore the availability of biotinylated and streptavidin linked fluorophores allows for an easy assay of their presence, using fluorescence microscopy<sup>2</sup>. | ||
| + | |- | ||
| + | |} | ||
| - | [[Image: | + | {| cellpadding="1" cellspacing="8" |
| + | |- | ||
| + | |Next we needed a way to anchor the streptavidin to the outer membrane. The obvious choice was to use the LPP signal peptide and Outer Membrane Protein A (OmpA) because this system has been extensively characterized as an expression system to display proteins on the cell surface. This LPP-OmpA system has been used to display a diverse group of proteins, including Green Fluorescent Protein (GFP)<sup>3</sup>, Organophosphorus Hydrolase (OPH)<sup>4</sup>, Cyclodextrin Glucanotransferase (CGTase)<sup>5</sup>, Methyl Parathion Hydrolase (MPH)<sup>6</sup>, and Enhanced Green Fluorescent Protein (EGFP)<sup>6</sup>. Thus, an appropriate display construct for our system would be a Lpp-OmpA-streptavidin fusion protein, which would be anchored in the outer membrane and display streptavidin on the surface of the cell. Streptavidin would bind the nanotag on our target protein in an interaction that could be disrupted by adding biotin. | ||
| + | |[[Image:Display image.jpg|400px|right]] | ||
| + | |- | ||
| + | |}<br> | ||
| + | ==== Harvard 2006 Surface Display System: Legacy Parts ==== | ||
| - | + | When we checked the Parts Registry to see if someone had already built this or sometime similar that we could use as a starting part for our Idealized Protein Purification Display System, we found the Lpp-OmpA-streptavidin construct submitted by the 2006 Harvard iGEM team<sup>7</sup>. In fact, Harvard had submitted four biobrick parts which were variations of the same function. All four are fusion proteins which have a LPP signal peptide, either one or five trans-membrane ompA, and either monomeric or a single-chain dimeric streptavidin. All the variations and the resulting biobricks are shown below. | |
| - | == | + | {|cellpadding="3" cellspacing="1" border="2" width="75%" align="center" |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
! Bio Brick | ! Bio Brick | ||
! OmpA trans-membrane domains | ! OmpA trans-membrane domains | ||
| - | ! Type of | + | ! Type of Streptavidin |
| - | |- | + | |- align="center" |
|J36848 | |J36848 | ||
|1 | |1 | ||
|monomeric | |monomeric | ||
| - | |- | + | |- align="center" |
|J36849 | |J36849 | ||
|1 | |1 | ||
|dimeric | |dimeric | ||
| - | |- | + | |- align="center" |
|J36850 | |J36850 | ||
|5 | |5 | ||
|monomeric | |monomeric | ||
| - | |- | + | |- align="center" |
|J36851 | |J36851 | ||
|5 | |5 | ||
| Line 56: | Line 56: | ||
|- | |- | ||
|} | |} | ||
| + | <br> | ||
| + | ==== Harvard 2006 Surface Display System: Documented Data ==== | ||
| - | + | Data regarding the activity of the Harvard iGEM 2006 Lpp-OmpA-Streptavidin construct has been published in IET Synthetic Biology in 2007<sup>7</sup>. The author showed, by Western blot against the His tag fused to this construct, that their protein was being expressed within the cell. In order to demonstrate binding of their cell-surface-anchored streptavidin to biotin, the author incubated cells expressing Lpp-OmpA-streptavidin with an aptamer that linked biotin to a fluorophore using an oligonucleotide linker (see figure below). Any cells that express streptavidin on the surface of the cell should fluoresce in their assay - concordantly, when cells were incubated with this biotin-oligo-fluorescein aptamer, the author saw fluorescence on cells expressing Lpp-OmpA-Streptavidin, but not on Lpp-OmpA negative control. However, cells that expressed streptavidin on the surface also bound to the negative aptamer control (oligo-fluorescein) that lacked biotin, indicating that the oligo aptamer itself (and not biotin) may have been responsible for the association of the fluorophore to the cell surface. Thus, the Harvard 2006 iGEM team was never able to definitively show specific affinity of their Lpp-OmpA-streptavidin construct. | |
| - | [[Image: | + | {| |
| + | |- | ||
| + | |[[Image:HarvardIETSYNTH.jpg|400px|center]] | ||
| + | |'' | ||
| + | L01W: Lpp-OmpA-Streptavidin monomer, no His Tag.<br> | ||
| + | L01H: Lpp-OmpA-Streptavidin monomer with His tag.<br> | ||
| + | L01S: Lpp-OmpA-Streptavidin dimer.<br> | ||
| + | L01: Lpp-OmpA alone).<br> | ||
| + | A: no aptamer added.<br> | ||
| + | B: With only ‘F’ 50 -fluorescently-tagged oligonucleotide.<br> | ||
| + | C: With fluorescently tagged streptavidin aptamer.<br> | ||
| + | D: With ‘B-F’ hybrid of 50 -biotinylated oligonucleotide annealed with 50-fluorescently-tagged oligonucleotide<sup>7</sup>.'' | ||
| + | |- | ||
| + | |} | ||
| + | |||
| - | + | Based on these experiments, we decided to try using this display system for our project. But since binding of the system to streptavidin had not be definitively shown, we wanted to verify binding of the Lpp-OmpA-streptavidin construct to biotin for ourselves, and characterize this interaction. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | = | + | ='''Experiments'''= |
| + | == Harvard's Legacy 2006 Cell Surface Display Parts: Do They Work As Predicted? == | ||
| - | + | In order to determine whether the Harvard 2006 iGEM streptavidin cell surface display parts were, first, expressing properly, and, second, binding biotin on the surface of the cell, we decided to do the following tests: | |
| - | + | ||
| - | + | # Western blot to verify expression of Harvard 2006 surface display parts | |
| - | # | + | # Assess biotin binding to cell surface by fluorescence microscopy |
| - | # | + | # Assess biotin binding to cell surface by flow cytometry |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | # | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | === | + | === Western Blot Demonstrates that Harvard 2006 iGEM Parts Are Expressed === |
| + | The goal of this experiment was to make sure that the proteins were being expressed in our cell strains, and also to make sure that they were the correct length. This was crucial to ascertain before we moved on and began to test the parts. Even though we had the individual parts sequenced confirmed we needed to make sure that the proteins were being expressed correctly in the cell. Since all the Harvard parts are His tagged, we used a Western blot reagent (horseradish peroxidase) that was conjugated to nickel-NTA which binds His tags.<sup>8</sup>. | ||
| - | |||
| - | |||
| - | The expected | + | ==== Data ==== |
| - | {| | + | |
| + | [[Image:HarvardExpressionWestern.jpg|700px|center]] | ||
| + | <br> | ||
| + | |||
| + | The expected sizes of each of these proteins are shown in the table below: | ||
| + | {|cellpadding="3" cellspacing="1" border="2" width="50%" | ||
! Bio Brick | ! Bio Brick | ||
| - | ! Length (in | + | ! Length (in Da) |
| - | |- | + | |- align="center" |
|J36848 | |J36848 | ||
|21478.7 | |21478.7 | ||
| - | |- | + | |- align="center" |
|J36849 | |J36849 | ||
|34600.9 | |34600.9 | ||
| - | |- | + | |- align="center" |
|J36850 | |J36850 | ||
|31215.4 | |31215.4 | ||
| - | |- | + | |- align="center" |
|J36851 | |J36851 | ||
|44972.3 | |44972.3 | ||
| Line 121: | Line 117: | ||
|} | |} | ||
| - | + | <br> | |
| + | Based on the table above, we were able to verify expression of the Harvard 2006 iGEM surface display parts. The varying intensity of the bands does not indicate the strength of expression because the protein amount was not normalized before it was inserted into the gel. Next we wanted to determine whether functional, biotin-binding streptavidin was displayed on the cell surface in cells expressing these parts. To test this, we used two different methods: visualization by fluorescence microscopy and by flow cytometer. <br> | ||
| + | === Streptavidin-Biotin Binding Is Not Visualized By Fluorescence Microscopy === | ||
| - | + | The goal of this experiment was to confirm the display of streptavidin on the surface of the cell. To do this, we incubated cells expressing the Harvard 2006 iGEM streptavidin surface display parts with a biotinylated fluorophore<sup>9</sup>. If cells are expressing functional streptavidin on the surface of the cell, they should bind the biotinylated fluorophore, and this binding should be detectable in our florescence microscope as a halo of fluorescence surrounding each individual cell. As a positive control for streptavidin-biotin binding we incubated the biotinylated fluorophore with streptavidin-coated beads that were roughly the same size (with respect to volume) as our cells (except spherical) (see [https://2009.igem.org/Team:Washington/Notebook/Microscope Notebook] page for protocols). | |
| - | The goal of this experiment was to | + | |
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
==== Data ==== | ==== Data ==== | ||
| - | |||
| - | + | <table> | |
| + | <tr> | ||
| + | <td><p style="font-size:18px"> Positive Control </p> </td> | ||
| + | <td><p style="font-size:18px"> Negative Control </p> </td> | ||
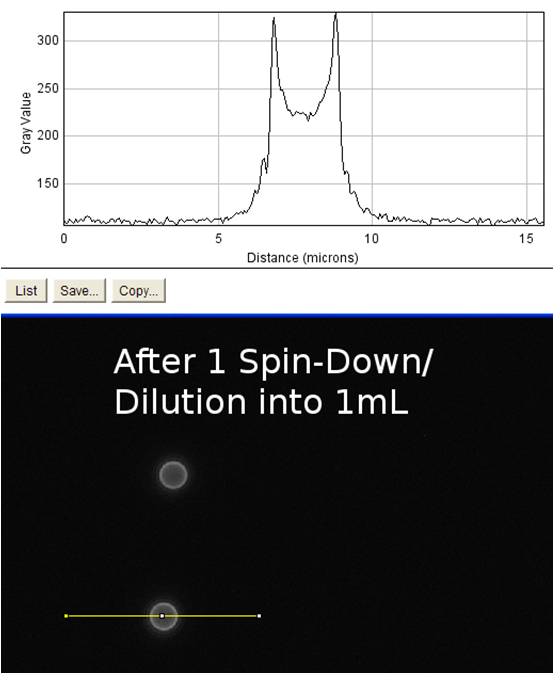
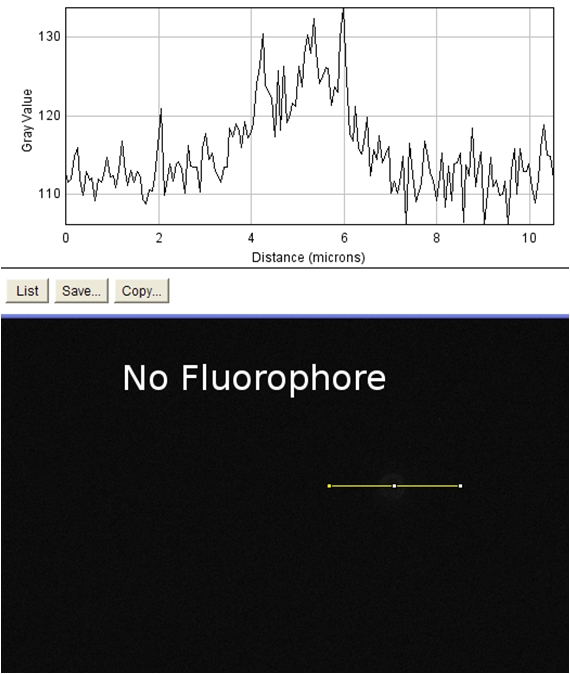
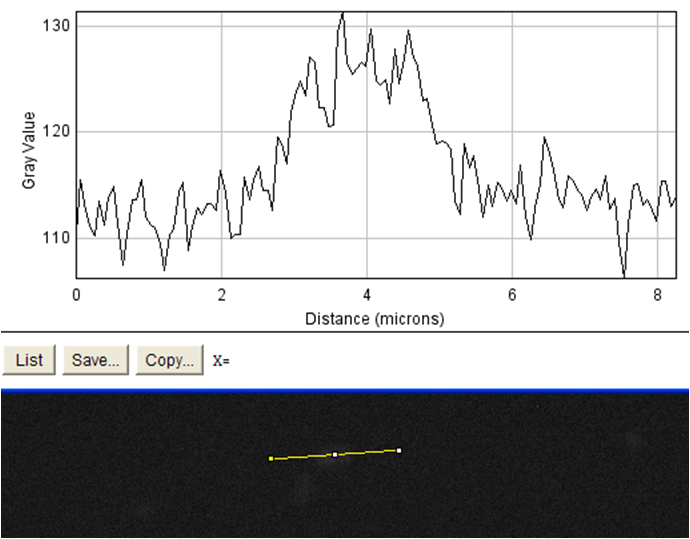
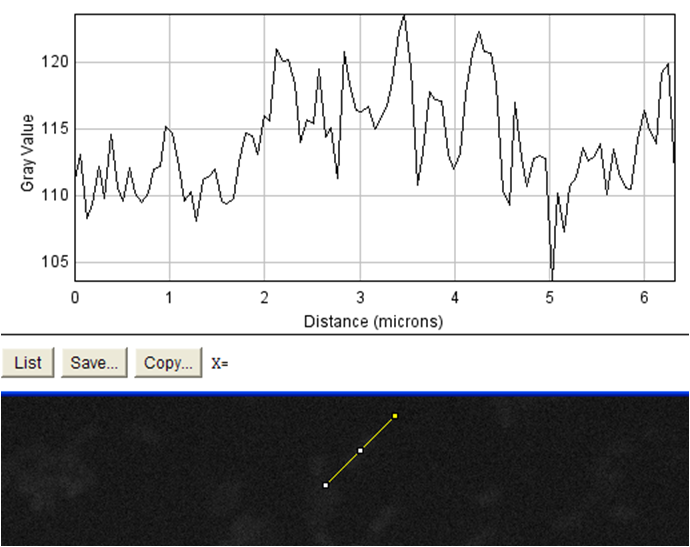
| + | <td rowspan="4">These images were analyzed using imageJ<sup>10</sup>. The image on the left shows an image of the beads with fluorophore added to them. These beads were diluted and spun diluted and spun down until the background level of fluorescence was low enough to get an accurate reading. We then used imageJ to analyze the intensity of a line going through the bead, which is demonstrated by the above schematic. On the positive control the edges of the streptavidin-coated beads show spikes in fluorescence, indicating binding of the biotinylated fluorophore to the beads. The negative control (beads without fluorophore) showed no such increase. This meant that our biotinylated fluorophore binds to streptavidin in a detectable manner. When our cells were examined in the same manner, no difference could be seen in biotinylated fluorophore binding between cells that had induced expression of surface streptavidin (left) and uninduced cells which should not express the surface display protein (right). This was evidence that the streptavidin surface display part binding to biotin was very low / nonexistent. In order to verify this, we measured the binding of the biotinylated fluorophore to entire populations of streptavidin-expressing cells by flow cytometry.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>[[Image:M_beads1.png| 210px]]</td> | ||
| + | <td>[[Image:M_beads2.png| 210px]]</td> | ||
| + | </tr> | ||
| + | <td><p style="font-size:18px"> Induced Cells </p></td> | ||
| + | <td><p style="font-size:18px"> Uninduced Cells </p></td> | ||
| + | <tr> | ||
| + | <td>[[Image:M_cells1.png| 210px]]</td> | ||
| + | <td>[[Image:M_cells2.png| 210px]]</td> | ||
| + | </tr> | ||
| + | </table> | ||
| - | === Flow Cytometry === | + | === Streptavidin-Biotin Binding Is Not Visualized By Flow Cytometry === |
| - | The goal of this experiment was to | + | The goal of this experiment was to visualize biotin binding to streptavidin over a whole population of cells. Whereas the Microscope experiment looked at a few localized cells, we set the cytometer to look 50,000 streptavidin-expressing cells after incubation (and washing) with the biotinylated fluorophore and read the resulting fluorescence ([https://2009.igem.org/Team:Washington/Notebook/Flow_Cytometry see Notebook page for protocols]). Like in the microscopy assay, we used streptavidin-coated beads as a positive control. The cytometer reads the fluorescence of each individual particle (cell or bead) passing through it, which allows for accurate readings, and monitoring of a much larger sample size than we visualized under the microscope. The results of this experiment are described below. |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
==== Data ==== | ==== Data ==== | ||
| - | + | <gallery heights=300px widths=350> | |
| - | + | Image:48.png|'''BBa_J36848''' ''This image shows both the induced and uninduced cells for part 48 in varying levels of fluorophore (0nM to 100nM). The cells were induced at 1mM IPTG. This data shows that there is no appreciable difference between the induced and uninduced cells at any given level of fluorophore. All curves appear to have the same amount of fluorescence. We found similar results using a microscopy assay.'' | |
| + | Image:StreptBead cyto.png|'''+ Control''' ''We used the streptavidin coated beads<sup>11</sup> to show us what the magnitude of fluorescence increase we should see with increased fluorophore levels. We were able to see that as the level of fluorophore was increased we could see increased retention between the beads and the fluorophore. The black is beads with no fluorophore, the red is with 10 nM, and the purple is 100 nM. These showed a clear difference between the beads without fluorophore, and the beads with fluorophore. The cells shown at right, matched the readings of the beads when they had no fluorophore added.'' | ||
| + | </gallery> | ||
| + | |||
| + | In the flow cytometer, our positive control, streptavidin-coated beads, showed a clear distinction between beads pre-incubated with biotinylated fluorophore and the beads without fluorophore, indicating that this assay is capable of visualizing streptavidin binding to the biotinylated fluorophore. Like in the microscopy assay, we did not observe appreciable binding occurring with any of the ompA-streptavidin parts to biotin at the population level. This data only shows part number 48. However this data is applicable to all of the parts we observed. All streptavidin-expressing cells demonstrated low levels of fluorescence comparable to those of beads not pre-incubated with fluorophore. | ||
| + | |||
| + | === Conclusion === | ||
| - | |||
| - | + | In conclusion, we demonstrated expression of the Harvard 2006 streptavidin surface display parts via Western Blot. We were unable, however, to visualize binding between a biotinylated fluorophore and the cells expressing these proteins. This indicates that the streptavidin display system is likely not binding biotin correctly. We hypothesize that streptavidin might not having enough room on the cell surface to form tetramers (which is its native state), and thus may not be binding to biotin with high efficiency. In order to generate a construct that demonstrates tight binding to to biotin, we propose to design a generalized cell surface display system, and to computationally design a biotin-binding streptavidin monomer using software developed at the University of Washington.<br> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| + | '''These solutions are described in the [https://2009.igem.org/wiki/index.php?title=Team:Washington/Future future directions section].''' | ||
| + | ---- | ||
| + | === Citations === | ||
| + | #The Nano-tag, a streptavidin-binding peptide for the purification and detection of recombinant proteins [http://www.ncbi.nlm.nih.gov/pubmed/14680960 doi:10.1016/j.physletb.2003.10.071] | ||
| + | #Structural origins of high-affinity biotin binding to streptavidin. PC Weber, DH Ohlendorf, JJ Wendoloski, and FR Salemme (6 January 1989). Science 243 (4887), 85. [http://www.ncbi.nlm.nih.gov/pubmed/2911722 DOI: 10.1126/science.2911722] | ||
| + | #Display of green fluorescent protein on ''Escherichia coli'' cell surface. Shi H, Wen Su W. Department of Biological & Agricultural Engineering, University of Missouri-Columbia, 65211, Columbia, MO, USA [http://www.ncbi.nlm.nih.gov/pubmed/11118595 Enzyme Microb Technol. 2001 Jan 2;28(1):25-34] | ||
| + | #Cell surface display of organophosphorus hydrolase using ice nucleation protein. Shimazu M, Mulchandani A, Chen W.Department of Chemical and Environmental Engineering, and Environmental Toxicology Program, University of California, Riverside, CA 92521, USA. [http://www.ncbi.nlm.nih.gov/pubmed/11170483 Biotechnol Prog. 2001 Jan-Feb;17(1):76-80] | ||
| + | #Anchorage of cyclodextrin glucanotransferase on the outer membrane of ''Escherichia coli''. Wan HM, Chang BY, Lin SC. Department of Chemical Engineering, National Chung Hsing University, Taichung, 402, Taiwan. [http://www.ncbi.nlm.nih.gov/pubmed/12115409 Biotechnol Bioeng. 2002 Aug 20;79(4):457-64] | ||
| + | #Cell surface display of functional macromolecule fusions on ''Escherichia coli'' for development of an autofluorescent whole-cell biocatalyst. Yang C, Zhao Q, Liu Z, Li Q, Qiao C, Mulchandani A, Chen W. State Key Laboratory of Integrated Management of Pest Insects and Rodents, Institute of Zoology, Chinese Academy of Sciences, Beijing 100101, China. [http://www.ncbi.nlm.nih.gov/pubmed/18767673 Environ Sci Technol. 2008 Aug 15;42(16):6105-10] | ||
| + | #Cell surface streptavidin. Tsai P. [http://dx.doi.org/10.1049/iet-stb:20070002 IET Synth Biol. 2007;1(1.2):32] | ||
| + | #[http://www.kpl.com/catalog/productdetail.cfm?catalog_ID=17&Category_ID=448&Product_ID=1085 HisDetector Nickel-HRP] | ||
| + | #Biotin-5-fluorescein, [http://www.sigmaaldrich.com/catalog/ProductDetail.do?lang=en&N4=53608|SIGMA&N5=SEARCH_CONCAT_PNO|BRAND_KEY&F=SPEC Sigma Aldrich] | ||
| + | #[http://rsbweb.nih.gov/ij/docs/index.html ImageJ] | ||
| + | #SVP-15-5 streptavidin coated polstyerene spheres (1.5-1.9 µm), [http://www.spherotech.com/coa_pol_par.htm Spherotech] | ||
{{Template:Team:Washington/Templates/Footer}} | {{Template:Team:Washington/Templates/Footer}} | ||
Latest revision as of 02:46, 22 October 2009
Background
Our system hinged on finding a protein which could bind other proteins to the outside of the cell, but whose interaction with these proteins was weak enough to be disrupted by another small molecule. streptavidin presented itself as a logical choice. Several protein tags (such as the Nano-Tag1) have been developed to bind streptavidin at the biotin binding site, but can be released when biotin is added to the system and binds to streptavidin. The ability for peptides to bind streptavidin and be released upon the addition of biotin is a technology currently used on a daily basis world-wide by labs purifying proteins. The major difference between the traditional system and our system is that streptavidin is attached to beads which are used in a column format in the traditional protein purification method. Our system will have streptavidin attached to the surface of the cell. Combining the display system with our target and secretion vector our vision is complete. A single cell can then produce, secrete, bind, and release any protein of interest.
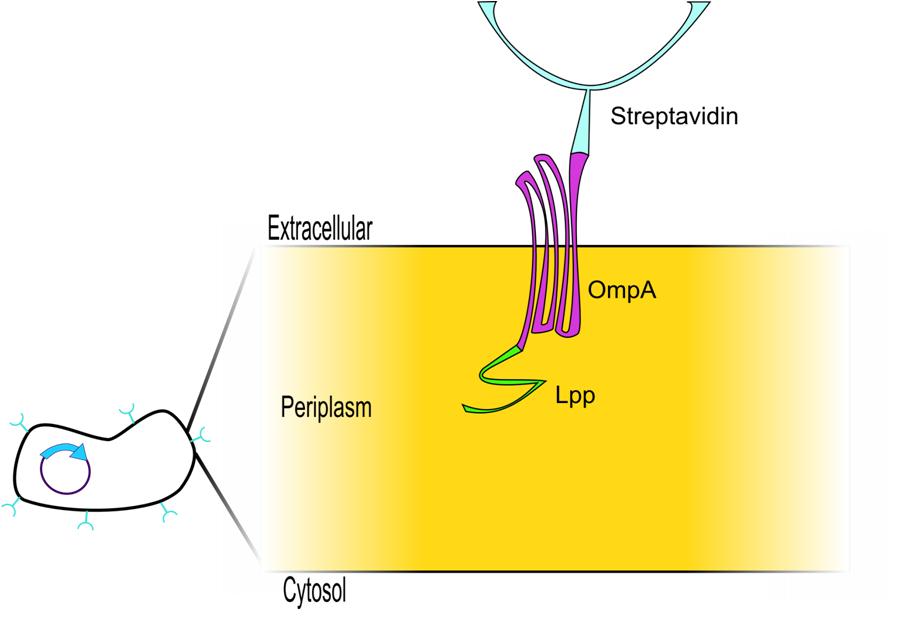
| Next we needed a way to anchor the streptavidin to the outer membrane. The obvious choice was to use the LPP signal peptide and Outer Membrane Protein A (OmpA) because this system has been extensively characterized as an expression system to display proteins on the cell surface. This LPP-OmpA system has been used to display a diverse group of proteins, including Green Fluorescent Protein (GFP)3, Organophosphorus Hydrolase (OPH)4, Cyclodextrin Glucanotransferase (CGTase)5, Methyl Parathion Hydrolase (MPH)6, and Enhanced Green Fluorescent Protein (EGFP)6. Thus, an appropriate display construct for our system would be a Lpp-OmpA-streptavidin fusion protein, which would be anchored in the outer membrane and display streptavidin on the surface of the cell. Streptavidin would bind the nanotag on our target protein in an interaction that could be disrupted by adding biotin. |
Harvard 2006 Surface Display System: Legacy Parts
When we checked the Parts Registry to see if someone had already built this or sometime similar that we could use as a starting part for our Idealized Protein Purification Display System, we found the Lpp-OmpA-streptavidin construct submitted by the 2006 Harvard iGEM team7. In fact, Harvard had submitted four biobrick parts which were variations of the same function. All four are fusion proteins which have a LPP signal peptide, either one or five trans-membrane ompA, and either monomeric or a single-chain dimeric streptavidin. All the variations and the resulting biobricks are shown below.
| Bio Brick | OmpA trans-membrane domains | Type of Streptavidin |
|---|---|---|
| J36848 | 1 | monomeric |
| J36849 | 1 | dimeric |
| J36850 | 5 | monomeric |
| J36851 | 5 | dimeric |
Harvard 2006 Surface Display System: Documented Data
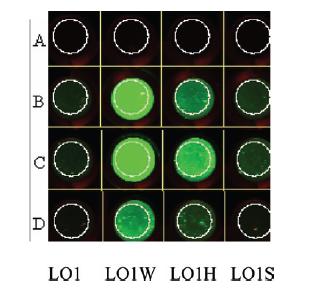
Data regarding the activity of the Harvard iGEM 2006 Lpp-OmpA-Streptavidin construct has been published in IET Synthetic Biology in 20077. The author showed, by Western blot against the His tag fused to this construct, that their protein was being expressed within the cell. In order to demonstrate binding of their cell-surface-anchored streptavidin to biotin, the author incubated cells expressing Lpp-OmpA-streptavidin with an aptamer that linked biotin to a fluorophore using an oligonucleotide linker (see figure below). Any cells that express streptavidin on the surface of the cell should fluoresce in their assay - concordantly, when cells were incubated with this biotin-oligo-fluorescein aptamer, the author saw fluorescence on cells expressing Lpp-OmpA-Streptavidin, but not on Lpp-OmpA negative control. However, cells that expressed streptavidin on the surface also bound to the negative aptamer control (oligo-fluorescein) that lacked biotin, indicating that the oligo aptamer itself (and not biotin) may have been responsible for the association of the fluorophore to the cell surface. Thus, the Harvard 2006 iGEM team was never able to definitively show specific affinity of their Lpp-OmpA-streptavidin construct.
|
L01W: Lpp-OmpA-Streptavidin monomer, no His Tag. |
Based on these experiments, we decided to try using this display system for our project. But since binding of the system to streptavidin had not be definitively shown, we wanted to verify binding of the Lpp-OmpA-streptavidin construct to biotin for ourselves, and characterize this interaction.
Experiments
Harvard's Legacy 2006 Cell Surface Display Parts: Do They Work As Predicted?
In order to determine whether the Harvard 2006 iGEM streptavidin cell surface display parts were, first, expressing properly, and, second, binding biotin on the surface of the cell, we decided to do the following tests:
- Western blot to verify expression of Harvard 2006 surface display parts
- Assess biotin binding to cell surface by fluorescence microscopy
- Assess biotin binding to cell surface by flow cytometry
Western Blot Demonstrates that Harvard 2006 iGEM Parts Are Expressed
The goal of this experiment was to make sure that the proteins were being expressed in our cell strains, and also to make sure that they were the correct length. This was crucial to ascertain before we moved on and began to test the parts. Even though we had the individual parts sequenced confirmed we needed to make sure that the proteins were being expressed correctly in the cell. Since all the Harvard parts are His tagged, we used a Western blot reagent (horseradish peroxidase) that was conjugated to nickel-NTA which binds His tags.8.
Data
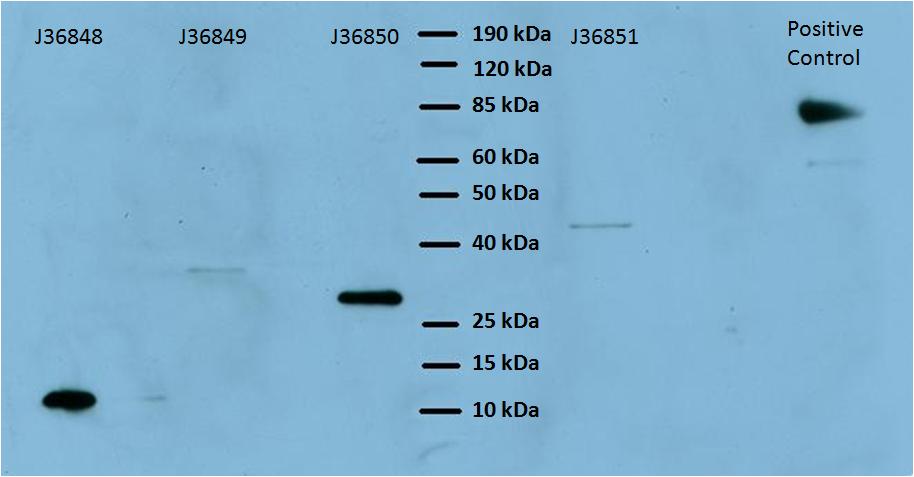
The expected sizes of each of these proteins are shown in the table below:
| Bio Brick | Length (in Da) |
|---|---|
| J36848 | 21478.7 |
| J36849 | 34600.9 |
| J36850 | 31215.4 |
| J36851 | 44972.3 |
Based on the table above, we were able to verify expression of the Harvard 2006 iGEM surface display parts. The varying intensity of the bands does not indicate the strength of expression because the protein amount was not normalized before it was inserted into the gel. Next we wanted to determine whether functional, biotin-binding streptavidin was displayed on the cell surface in cells expressing these parts. To test this, we used two different methods: visualization by fluorescence microscopy and by flow cytometer.
Streptavidin-Biotin Binding Is Not Visualized By Fluorescence Microscopy
The goal of this experiment was to confirm the display of streptavidin on the surface of the cell. To do this, we incubated cells expressing the Harvard 2006 iGEM streptavidin surface display parts with a biotinylated fluorophore9. If cells are expressing functional streptavidin on the surface of the cell, they should bind the biotinylated fluorophore, and this binding should be detectable in our florescence microscope as a halo of fluorescence surrounding each individual cell. As a positive control for streptavidin-biotin binding we incubated the biotinylated fluorophore with streptavidin-coated beads that were roughly the same size (with respect to volume) as our cells (except spherical) (see Notebook page for protocols).
Data
Streptavidin-Biotin Binding Is Not Visualized By Flow Cytometry
The goal of this experiment was to visualize biotin binding to streptavidin over a whole population of cells. Whereas the Microscope experiment looked at a few localized cells, we set the cytometer to look 50,000 streptavidin-expressing cells after incubation (and washing) with the biotinylated fluorophore and read the resulting fluorescence (see Notebook page for protocols). Like in the microscopy assay, we used streptavidin-coated beads as a positive control. The cytometer reads the fluorescence of each individual particle (cell or bead) passing through it, which allows for accurate readings, and monitoring of a much larger sample size than we visualized under the microscope. The results of this experiment are described below.
Data
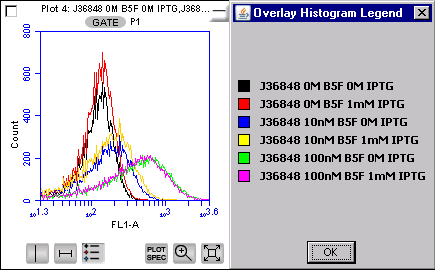
BBa_J36848 This image shows both the induced and uninduced cells for part 48 in varying levels of fluorophore (0nM to 100nM). The cells were induced at 1mM IPTG. This data shows that there is no appreciable difference between the induced and uninduced cells at any given level of fluorophore. All curves appear to have the same amount of fluorescence. We found similar results using a microscopy assay. |
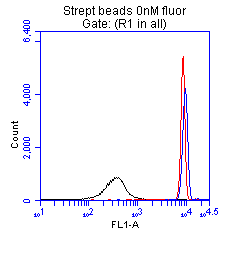
+ Control We used the streptavidin coated beads11 to show us what the magnitude of fluorescence increase we should see with increased fluorophore levels. We were able to see that as the level of fluorophore was increased we could see increased retention between the beads and the fluorophore. The black is beads with no fluorophore, the red is with 10 nM, and the purple is 100 nM. These showed a clear difference between the beads without fluorophore, and the beads with fluorophore. The cells shown at right, matched the readings of the beads when they had no fluorophore added. |
In the flow cytometer, our positive control, streptavidin-coated beads, showed a clear distinction between beads pre-incubated with biotinylated fluorophore and the beads without fluorophore, indicating that this assay is capable of visualizing streptavidin binding to the biotinylated fluorophore. Like in the microscopy assay, we did not observe appreciable binding occurring with any of the ompA-streptavidin parts to biotin at the population level. This data only shows part number 48. However this data is applicable to all of the parts we observed. All streptavidin-expressing cells demonstrated low levels of fluorescence comparable to those of beads not pre-incubated with fluorophore.
Conclusion
In conclusion, we demonstrated expression of the Harvard 2006 streptavidin surface display parts via Western Blot. We were unable, however, to visualize binding between a biotinylated fluorophore and the cells expressing these proteins. This indicates that the streptavidin display system is likely not binding biotin correctly. We hypothesize that streptavidin might not having enough room on the cell surface to form tetramers (which is its native state), and thus may not be binding to biotin with high efficiency. In order to generate a construct that demonstrates tight binding to to biotin, we propose to design a generalized cell surface display system, and to computationally design a biotin-binding streptavidin monomer using software developed at the University of Washington.
These solutions are described in the future directions section.
Citations
- The Nano-tag, a streptavidin-binding peptide for the purification and detection of recombinant proteins [http://www.ncbi.nlm.nih.gov/pubmed/14680960 doi:10.1016/j.physletb.2003.10.071]
- Structural origins of high-affinity biotin binding to streptavidin. PC Weber, DH Ohlendorf, JJ Wendoloski, and FR Salemme (6 January 1989). Science 243 (4887), 85. [http://www.ncbi.nlm.nih.gov/pubmed/2911722 DOI: 10.1126/science.2911722]
- Display of green fluorescent protein on Escherichia coli cell surface. Shi H, Wen Su W. Department of Biological & Agricultural Engineering, University of Missouri-Columbia, 65211, Columbia, MO, USA [http://www.ncbi.nlm.nih.gov/pubmed/11118595 Enzyme Microb Technol. 2001 Jan 2;28(1):25-34]
- Cell surface display of organophosphorus hydrolase using ice nucleation protein. Shimazu M, Mulchandani A, Chen W.Department of Chemical and Environmental Engineering, and Environmental Toxicology Program, University of California, Riverside, CA 92521, USA. [http://www.ncbi.nlm.nih.gov/pubmed/11170483 Biotechnol Prog. 2001 Jan-Feb;17(1):76-80]
- Anchorage of cyclodextrin glucanotransferase on the outer membrane of Escherichia coli. Wan HM, Chang BY, Lin SC. Department of Chemical Engineering, National Chung Hsing University, Taichung, 402, Taiwan. [http://www.ncbi.nlm.nih.gov/pubmed/12115409 Biotechnol Bioeng. 2002 Aug 20;79(4):457-64]
- Cell surface display of functional macromolecule fusions on Escherichia coli for development of an autofluorescent whole-cell biocatalyst. Yang C, Zhao Q, Liu Z, Li Q, Qiao C, Mulchandani A, Chen W. State Key Laboratory of Integrated Management of Pest Insects and Rodents, Institute of Zoology, Chinese Academy of Sciences, Beijing 100101, China. [http://www.ncbi.nlm.nih.gov/pubmed/18767673 Environ Sci Technol. 2008 Aug 15;42(16):6105-10]
- Cell surface streptavidin. Tsai P. [http://dx.doi.org/10.1049/iet-stb:20070002 IET Synth Biol. 2007;1(1.2):32]
- [http://www.kpl.com/catalog/productdetail.cfm?catalog_ID=17&Category_ID=448&Product_ID=1085 HisDetector Nickel-HRP]
- Biotin-5-fluorescein, [http://www.sigmaaldrich.com/catalog/ProductDetail.do?lang=en&N4=53608|SIGMA&N5=SEARCH_CONCAT_PNO|BRAND_KEY&F=SPEC Sigma Aldrich]
- [http://rsbweb.nih.gov/ij/docs/index.html ImageJ]
- SVP-15-5 streptavidin coated polstyerene spheres (1.5-1.9 µm), [http://www.spherotech.com/coa_pol_par.htm Spherotech]
 "
"