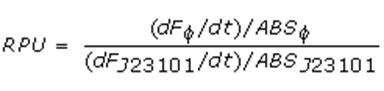Team:UNIPV-Pavia/Parts Characterization
From 2009.igem.org
(→Characterization) |
|||
| Line 692: | Line 692: | ||
<td>BBa_K173000</td> | <td>BBa_K173000</td> | ||
<td>36.32</td> | <td>36.32</td> | ||
| - | <td> | + | <td> 2.044428 [1.996125 ; 2.075898] |
| + | </td> | ||
</tr> | </tr> | ||
| Line 698: | Line 699: | ||
<td>BBa_K173001</td> | <td>BBa_K173001</td> | ||
<td>36.34</td> | <td>36.34</td> | ||
| - | <td> | + | <td>1 (reference standard)</td> |
</tr> | </tr> | ||
| Line 704: | Line 705: | ||
<td>BBa_K173002</td> | <td>BBa_K173002</td> | ||
<td>35.44</td> | <td>35.44</td> | ||
| - | <td> | + | <td> 0.692686 [0.643722 ; 0.734648] |
| + | </td> | ||
</tr> | </tr> | ||
Revision as of 09:31, 21 October 2009

|
|
|
Parts Characterization |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Here we describe the characterization results of 4 parts of our own design, 2 existing parts re-built because they were inconsistent and 7 existing parts taken from the Registry. When not reported differently, all the experiments have been performed according to Growth conditions and Data analysis sections.
Re-built existing parts (BBa_our part code/BBa_existing part code): Existing parts from the Registry:
Existing parts: sequence debugging
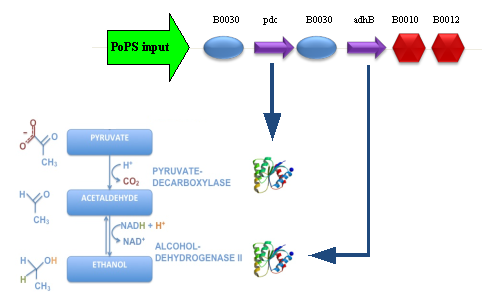
Our new partsBBa_K173003 - ethanol producing deviceDescriptionThis device takes PoPS as input and produces pyruvate decarboxylase (pdc) and alcohol dehydrogenase II (adhB) enzymes. Pyruvate decarboxylase (pdc, ) catalyses the decarboxylation of pyruvic acid to acetaldehyde and carbon dioxide, while alcohol dehydrogenase II (adhB, ) catalyses the acetaldehyde reduction to ethanol. The latter enzyme, as reported in literature, is also able to work in the opposite direction, oxidizing ethanol to acetaldehyde. The two enzymes come from Zymomonas mobilis ethanologenic bacterium and constitute an essential step of alcoholic fermentation. So, this device contains the minimum set of genes that are required to engineer a heterologous fermentation pathway. The coding sequences of pdc and adhB genes have been optimized for Escherichia coli codon usage. CharacterizationQualitative phenotype characterization PLATES RESULTS This device has been cloned downstream of 4 different promoters, one of which in two different vectors. These assemblies were transformed in TOP10 and gave the following phenotypes on LB agar plates and also on LB agar plates + 2% glucose (all the plates have been incubated at 37°C for about 11 hours, then some colonies were picked with a sterile tip and plates were incubated for additional 10 hours):
These results demonstrate that a constitutive expression of pdc and adhB gives a high metabolic burden to E. coli, in fact the only normal phenotype on plates was with non-induced device. This assembled part has been submitted to the Registry as .
Grown cultures of and after the FIRST INOCULUM showed different phenotypes (qualitative analysis), as a function of the used protocol.
PROTOCOL#3 RESULTS (i.e. shaken anaerobic falcon tubes, 2% glucose):
These results confirm the metabolic burden given by this BioBrick when gene expression is triggered: induced strains bearing a high copy plasmid containing do not survive, while low copy ones can survive even at high induction (1 uM). We decided to exploit the lux promoter leakage activity for in pSB1AK3 (high copy), while we decided to induce in pSB4C5 (low copy) the with 1 uM of 3OC6-HSL during our fermentation experiments. The qualitative phenotypes of the grown cultures of and after fermentation in 10% glucose showed that pellets were higher in (in both pSB1AK3 uninduced and pSB4C5 induced) than in and in the negative control .
Quantitative characterization pH pH has been measured in grown cultures after the FIRST INOCULUM (in 2% glucose after 24 hours) and after the SECOND INOCULUM (in 10% glucose after 24 or 48 hours). The measurements performed are the following: PROTOCOL#1: pH at the end of fermentation (after the SECOND INOCULUM). PROTOCOL#2: pH after the FIRST and the SECOND (end of fermentation) INOCULUM PROTOCOL#3: pH after the FIRST and the SECOND (end of fermentation) INOCULUM The shown results suggest that in strains bearing expressed pdc and adhB the fermentation does not give high levels of organic acids as in negative control strains which do not have pdc and adhB. It is surprising that the pH of in pSB1AK3 (pdc and adhB without any promoter) is slightly higher than in the negative controls. This could be due to a weak spurious transcription of pdc and adhB, amplified by the high copy number plasmid. This may re-direct part of pyruvate metabolism to ethanol and not to organic acids. Of course this phenomenon should be further investigated. Growth curves and its suitable negative controls growth was tested in our microplate reader: 200 ul aliquots (3 or 4 replicates) were taken from a just inoculated 30 ml culture (LB + 10% glucose) of the SECOND INOCULUM of PROTOCOL#3 and let grow in the microplate reader according to the automatic protocol described in Microplate reader experiments. The microplate was "sealed" with parafilm in order to create an anaerobic condition. This test showed that in pSB1AK3 is able to reach much higher OD600 than the other cultures, reaching ? values. in pSB4C5 both induced and uninduced had comparable growth curves and reached OD600 of about ?. Negative controls ( in pSB1A2 and in pSB1AK3) had comparable growth curves and reached low OD (about ?). It is surprising that in pSB1AK3 reached a higher OD (?) than the negative controls. The shown results are in complete accordance with the pH results, confirming that strains bearing in pSB1AK3 and in pSB4C5 produce less organic acids than the negative controls, hopefully thanks to pdc and adhB and consequently grow better.
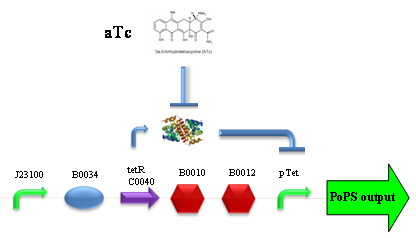
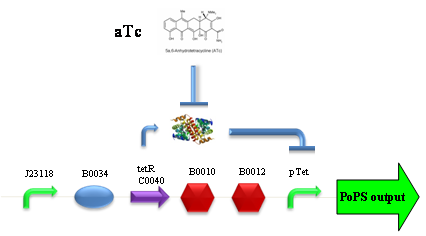
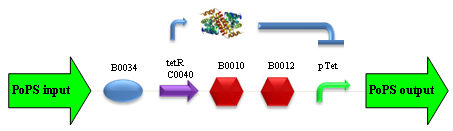
PROTOCOL#1: ethanol at the end of fermentation. PROTOCOL#2: ethanol at the end of fermentation. PROTOCOL#2: ethanol at the end of fermentation. PROTOCOL#3: ethanol at the end of fermentation. ConclusionsBBa_K173007 - aTc inducible device with J23100 promoterDescriptionThis is an aTc sensing device. promoter drives the constitutive production of tetR repressor (), which inhibits tetR promoter () activity. When aTc is added to the medium, it binds tetR and inhibits it. So, the PoPS output is a function of the aTc concentration. A tight regulation is expected for this inducible system because BBa_J23100 is a strong promoter and so tetR repressor should be produced at extremely high levels. The data below are referred to , which is the measurement system of . Characterization
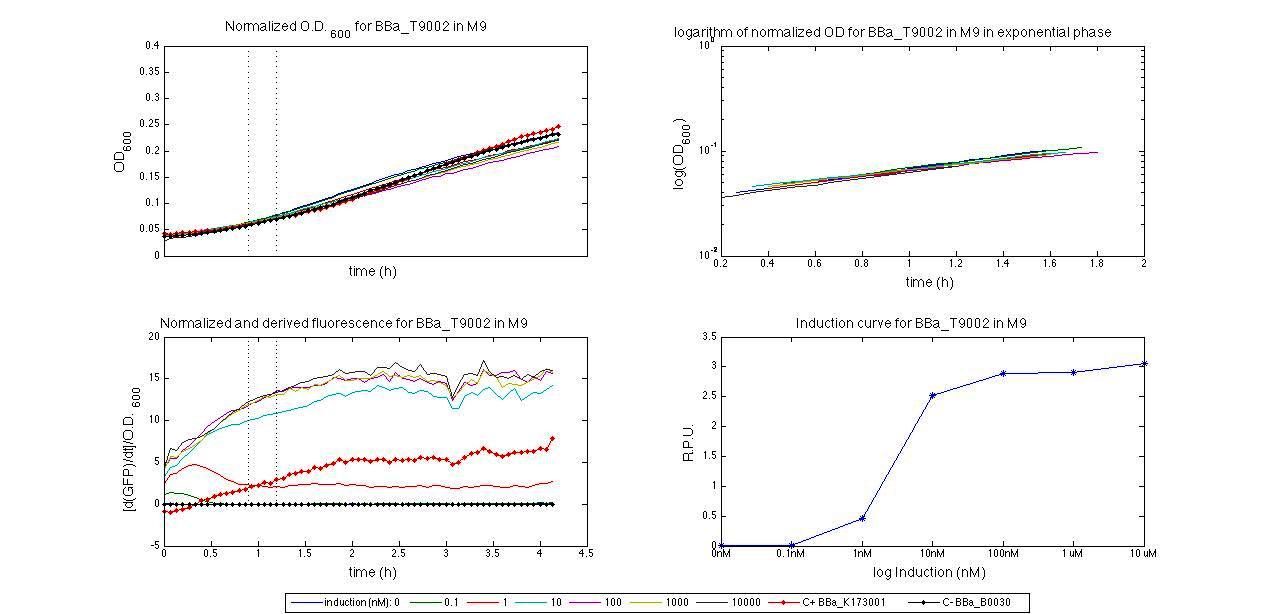
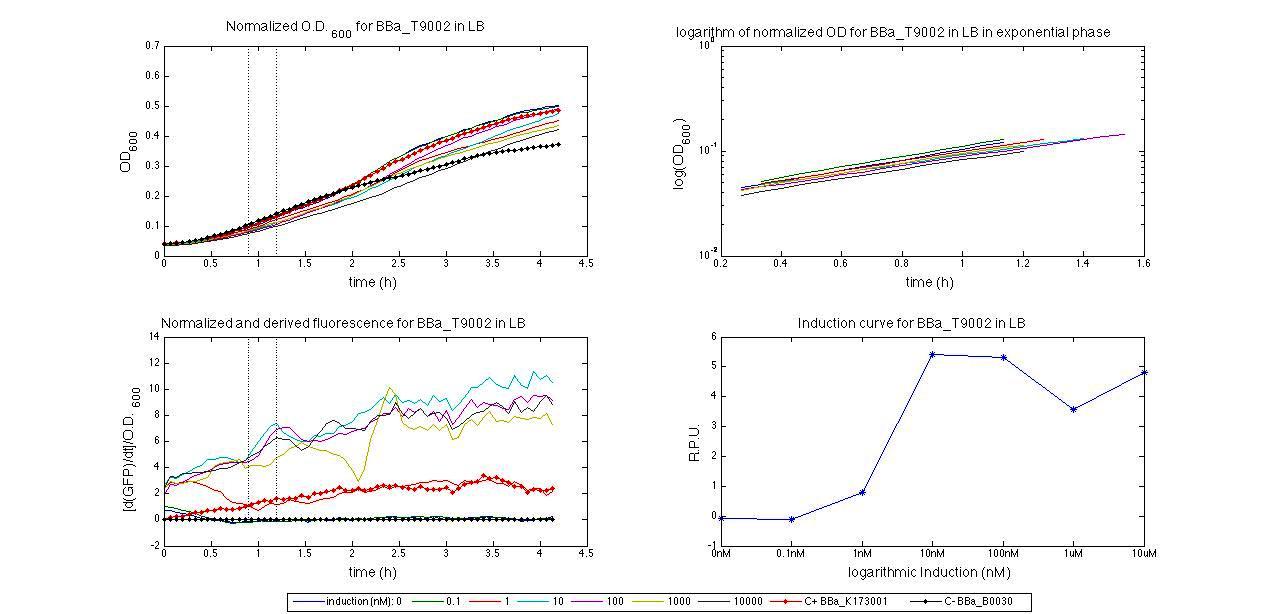
ConclusionsWe demonstrated that this part works as expected, sensing the aTc concentration provided in the culture medium. The transfer function of this sensor has been characterized in standard units (RPUs) in two different growth media (LB and M9 supplemented with glycerol), as well as the metabolic burden (in terms of doubling time) which affects E. coli bearing this part. On the other hand, we did not expect to have a higher GFP synthesis rate per cell after the exponential growth phase than in the exponential phase itself (as reported in the 3rd plot). This parts shows to have a very low leakage rate (about 0.025 RPU) but also a very low induction for high concentrations of aTc. So it can be used for tight regulation, in those systems who need very low leakage rates (no gene expressed in absence of inductor) and a response not important. BBa_K173011 - aTc inducible device with J23118 promoterDescriptionThis is an aTc sensing device. promoter drives the constitutive production of tetR repressor (), which inhibits tetR promoter () activity. When aTc is added to the medium, it binds tetR and inhibits it. So, the PoPS output is a function of the aTc concentration. A less tight regulation is expected for this inducible system than in because BBa_J23118 promoter is weaker than BBa_J23100 and so tetR repressor shold be produced at lower levels than in the other sensor. The data below are referred to , which is the measurement system of . Characterization
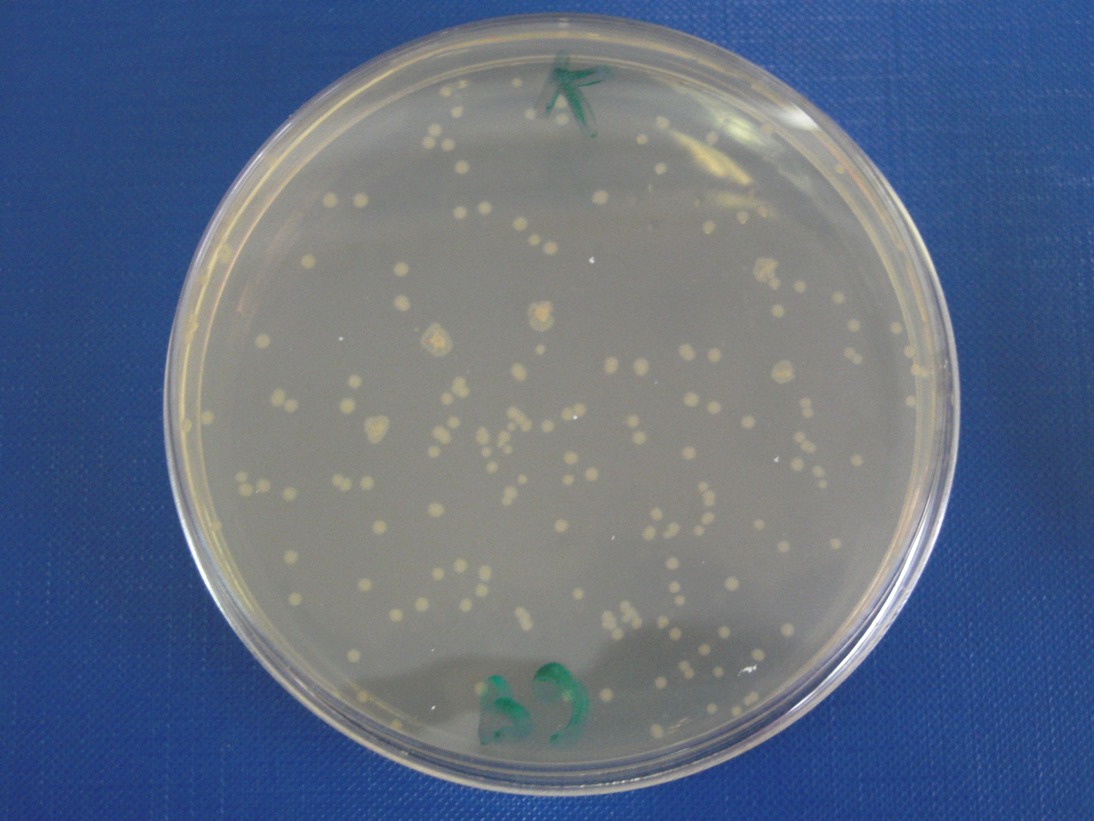
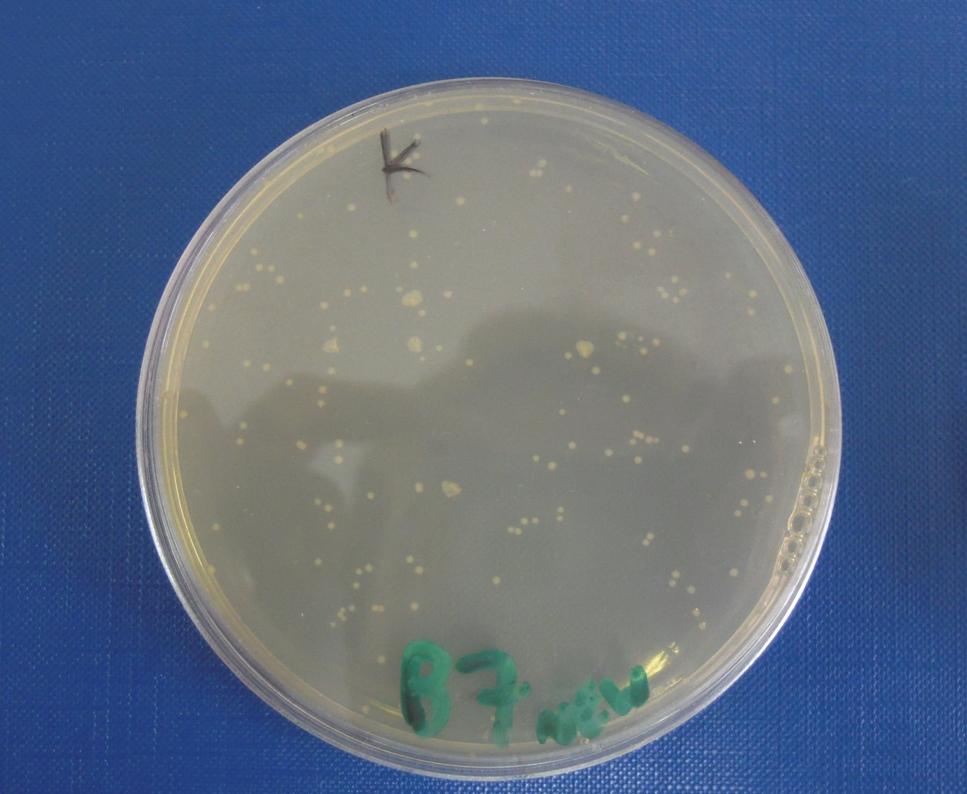
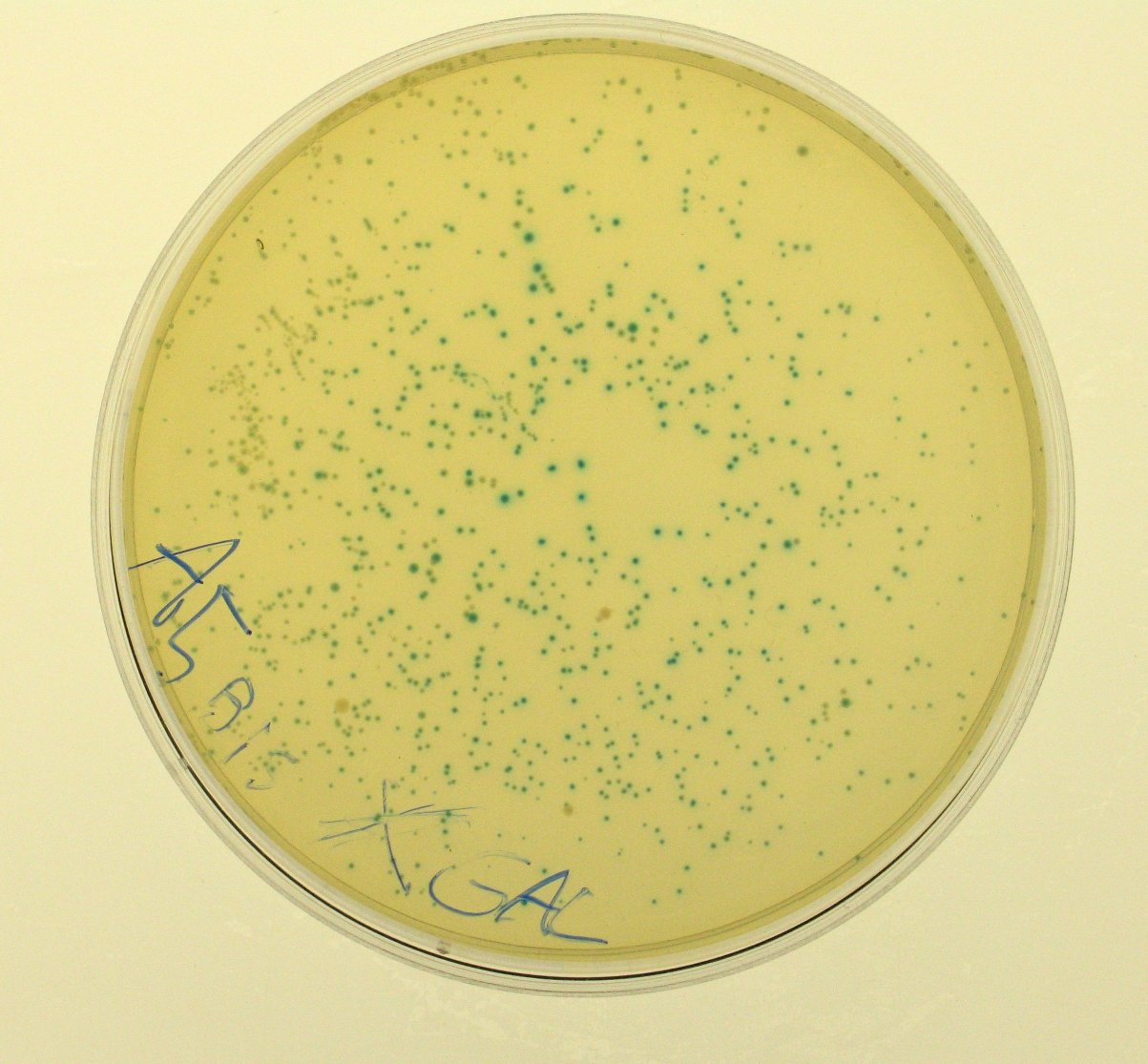
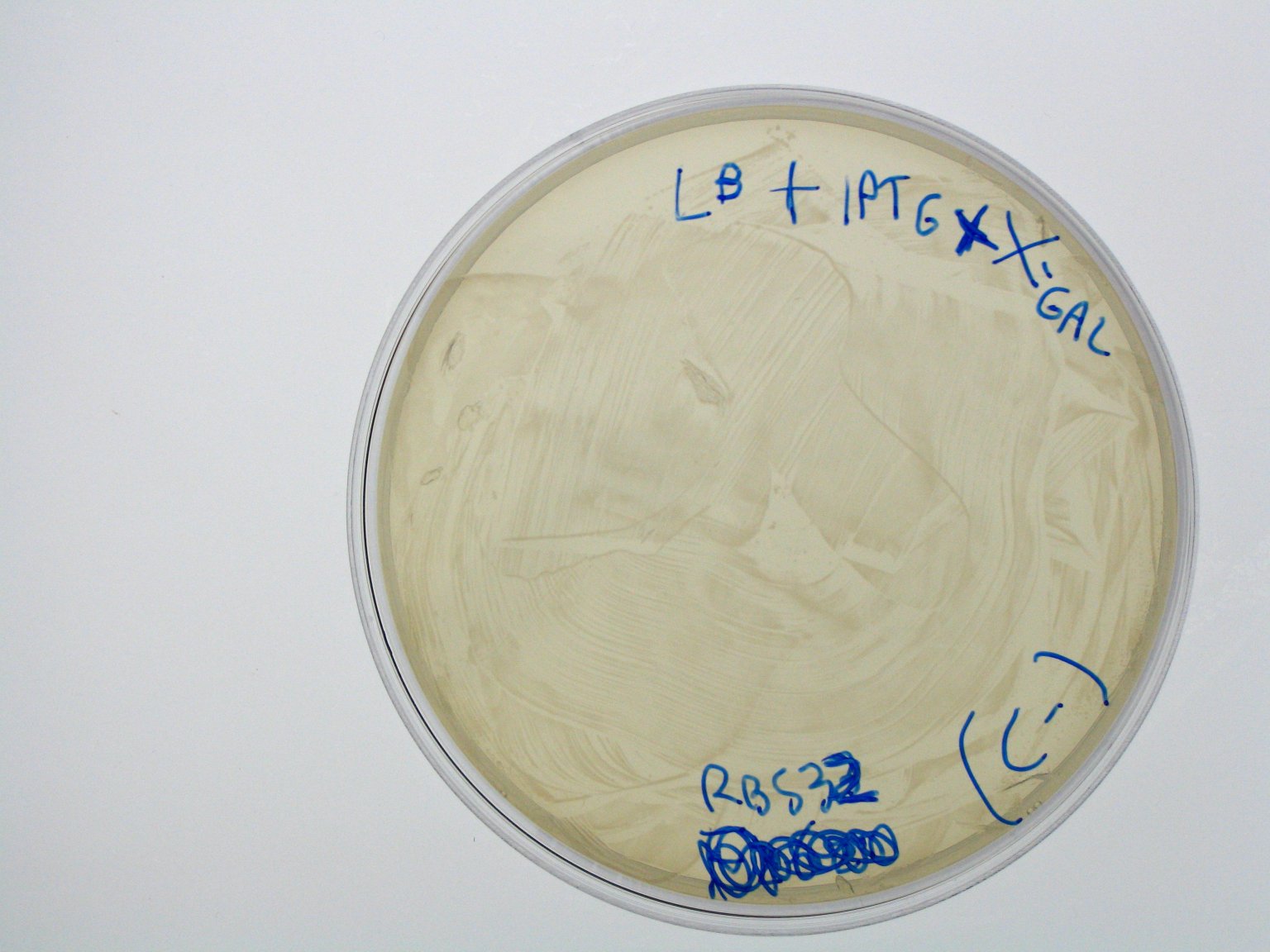
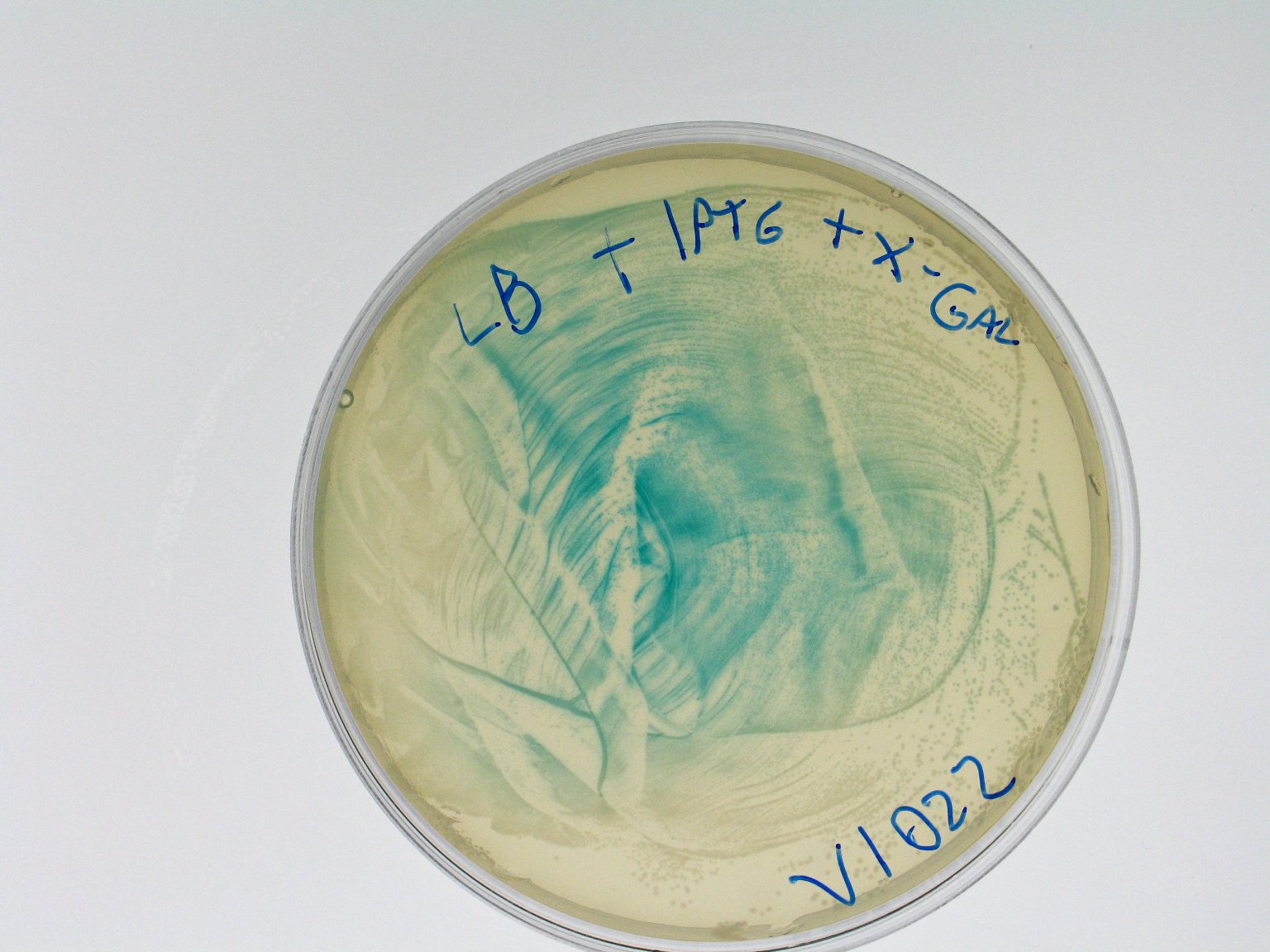
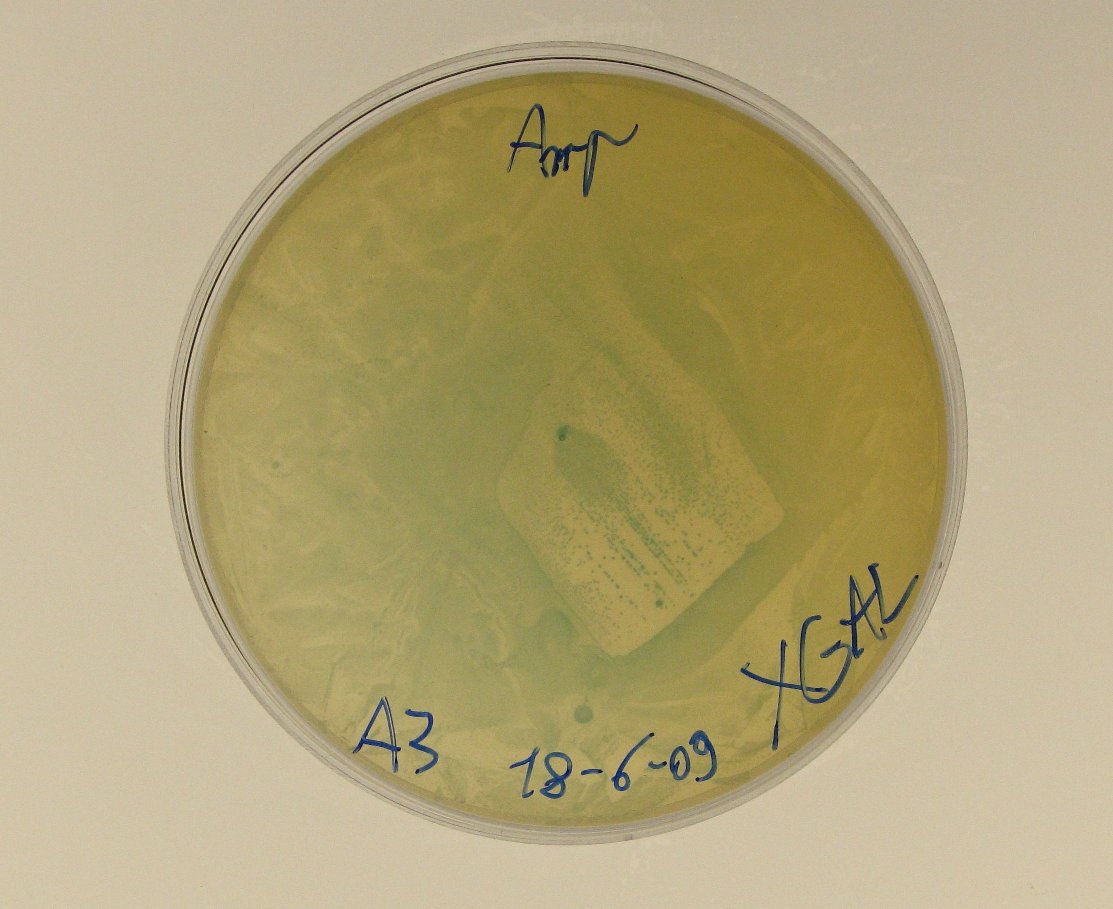
ConclusionsWe demonstrated that this part works as expected because GFP is produced as an increasing function of the aTc concentration provided in the culture medium. The transfer function of this sensor has been characterized in standard units (RPUs) in two different growth media (LB and M9 supplemented with glycerol), as well as the metabolic burden (in terms of doubling time) which affects E. coli bearing this part. On the other hand, as for BBa_K173007, we did not expect to have a higher GFP synthesis rate per cell after the exponential growth phase than in the exponential phase itself (as reported in the 3rd plot). This part shows a high level of leakage, of about 1.5 RPU but also high levels of induction for high concentrations of aTc. For this reason, it can be used in those cases where an important response in term of gene expression is required in presence of inductor, but is not suitabel if the absence of expression has to be absten when inductor is absent. BBa_K173010 - lactose/IPTG inducible device with J23118 promoterDescriptionThis should work as a lactose/IPTG sensor. promoter drives the constitutive production of lacI repressor (), which inhibits lac promoter () activity. When lactose or IPTG is added to the medium, it binds lacI and inhibits it. So, the PoPS output is a function of lactose/IPTG concentration. Thanks to the hybrid lac promoter (), designed taking the Plambda promoter () and substituting its cI () binding sites with two lacI binding sites, the behaviour of this device is not a function of glucose concentration because the wild type CAP binding sites are not present in this artificial lac promoter. The test performed to characterize this device has been done with , which is the measurement system of itself. CharacterizationConclusionsWe did not perform any standard measurement on this device because preliminary tests showed that GFP levels of induced and non induced cultures were the same and were equals to negative control. Unfortunately, we did not check if the sequence of , so we do not know if it was actually correct. Another possibility is that lacI is produced at so high levels that 2 mM of IPTG is not sufficient to induce the lac promoter. Further tests should be done for this system. Re-built existing partsBBa_K173004/BBa_I732019 - beta-galactosidase protein generatorDescriptionThis is a beta-galactosidase protein generator with strong RBS. We built up . It is a twin of , which was classified as "inconsistent" by iGEM HQ in 2008 and so we decided to improve this part submitting a new consistent DNA to the Registry. This part takes PoPS as input to express lacZ gene (), encoding for beta-galactosidase enzyme. This enzyme can be used to cleave lactose molecule to glucose and galactose, but can also be used as a reporter protein for colorimetric assays (together with X-Gal or ONPG as a substrate). X-gal is cleaved by β-galactosidase yielding galactose and 5-bromo-4-chloro-3-hydroxyindole. The latter is then oxidized into 5,5'-dibromo-4,4'-dichloro-indigo, an insoluble blue product. Characterization50 ul of , , (positive control) and were plated on LB + Amp + X-Gal + IPTG plates (except for for which non selective LB was used). Plated bacteria were incubated at 37°C for about 16 hours and then a picture of the plates was taken. To prepare these plates, 20 ul of X-Gal 40 mg/ml in DMF and 20 ul of ready made IPTG were diluted in 60 ul of SOC medium and spread on a LB agar plate. ConclusionsWe improved the existing part building up and testing its activity. This new part has been submitted to the Registry, as well as its physical DNA, allowing future users to assemble this beta-gal protein generator in their own project. This part has shown to work as expected when a PoPS input is given, being able to cleave X-Gal on LB agar plates. In our case, we have tested this part with upstream, which provides a promoter strength of ? RPU in LB medium. This assembled part can be considered as a measurement system for . It has been tested in pSB1AK3 vector. Surprisingly, as reported in the 4th picture, in the plate with the promoterless protein generator blue colour can be seen. It may be due to i) spurious transcription of the protein generator in the high copy number plasmid pSB1AK3 or ii) to the recombination occurred between plasmidic lacZ and genomic lacZdeltaM15, in which the working lacZ was integrated in E. coli genome under the control of lac promoter. This phenomenon has still to be studied. Caltech iGEM 2008 team reported this phenomenon in a similar protein generator, in which beta-gal assay was performed. Anyway, further comparative tests should be done in order to see if lactose cleavage can be performed faster than in wild type E. coli, after the choice of a suitable promoter which controls this protein generator. BBa_K173005/BBa_Q04400 - tetR QPIDescriptionCharacterizationThis part has been charachterized only in M9 supplemented medium. In LB only a measure in absence of inductor has been performed.
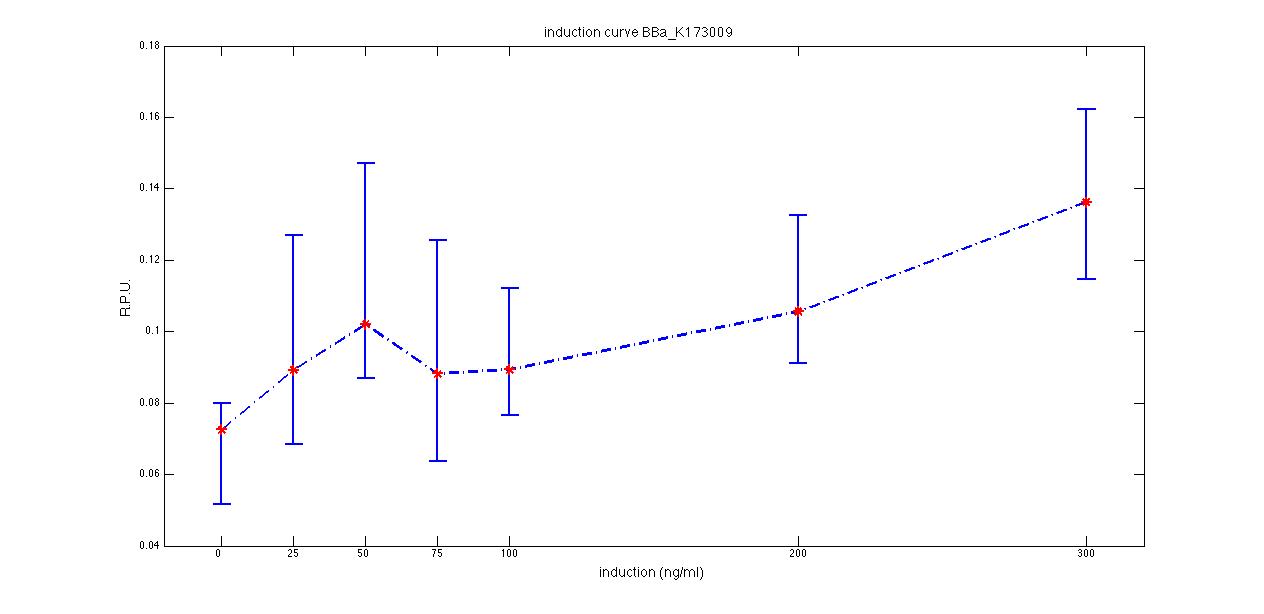
ConclusionsExisting parts from the RegistryBBa_J23100, BBa_J23101, BBa_J23118 - constitutive promoter family membersDescriptionThese three promoters are from the Anderson Promoter Collection, which is a library of constitutive sigma70 bacterial promoters. The strength of each promoter of the library has already been estimated in saturation growth phase cultures in LB, but here we provide the characterization of BBa_J23100 and BBa_J23118 in standard units (RPUs) in LB medium, in order to add experience and data for these BioBricks. BBa_J23101 is the reference standard promoter, so it has RPU=1 for definition. The data shown below are referred to , and that are the measurement parts of respectively , and . Characterization
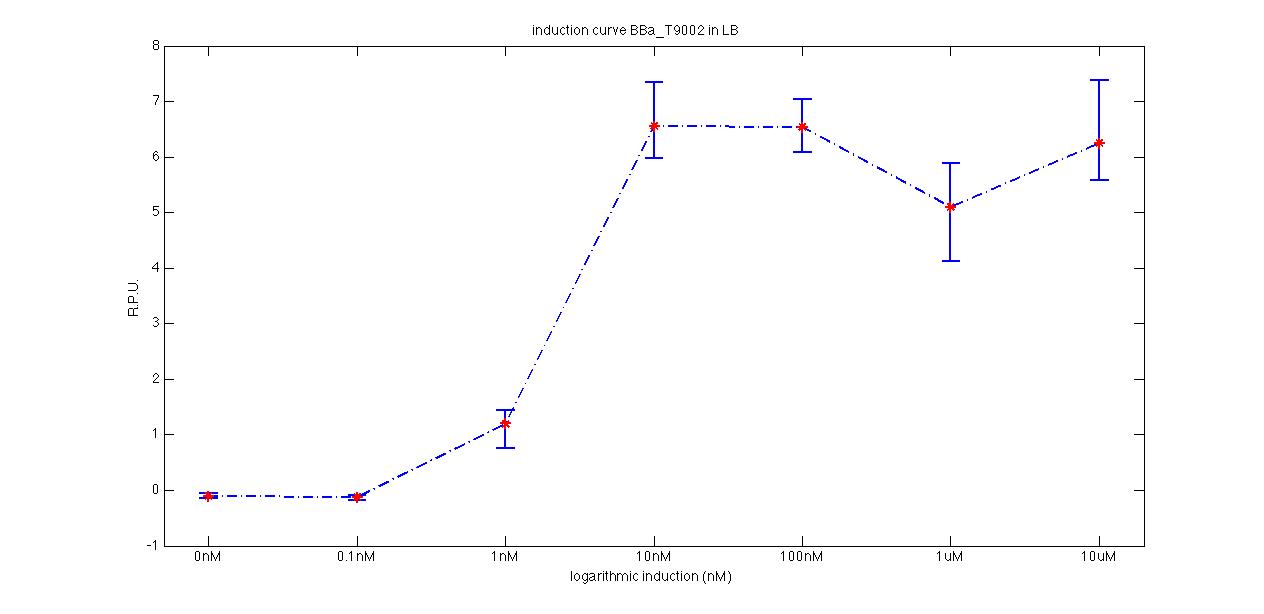
ConclusionsRPU estimation of these promoters was not present in the Registry and even the doubling time of these parts was not documented. We added these data in the pages of , and characterized parts, hoping that they can be useful for promoter comparison in standard units. If we consider the promoter ranking, provided in saturation phase in the [http://partsregistry.org/Promoters/Catalog/Anderson Anderson Promoter Collection Registry page], the estimated strength in RPU of BBa_J23100 and BBa_J23118 are in accordance with these values: INSERIRE I VALORI E COMMENTARE Note: plasmid is equals to pSB1A2 with a RFP expression system downstream of the cloning site. BBa_F2620 - 3OC6HSL receiver deviceDescriptionThis device gives PoPS as output and can be induced with 3OC6-HSL autoinducer molecule: it binds luxR protein (encoded by ), which is constitutively expressed by tetR promoter (). LuxR-HSL complex can work as a transcriptional activator for lux promoter (). Several studies have been performed on this BioBrick. Here we provide the experimental characterization we performed during this summer. The tests have been performed through measurement system, which has a GFP protein generator downstream. Characterization
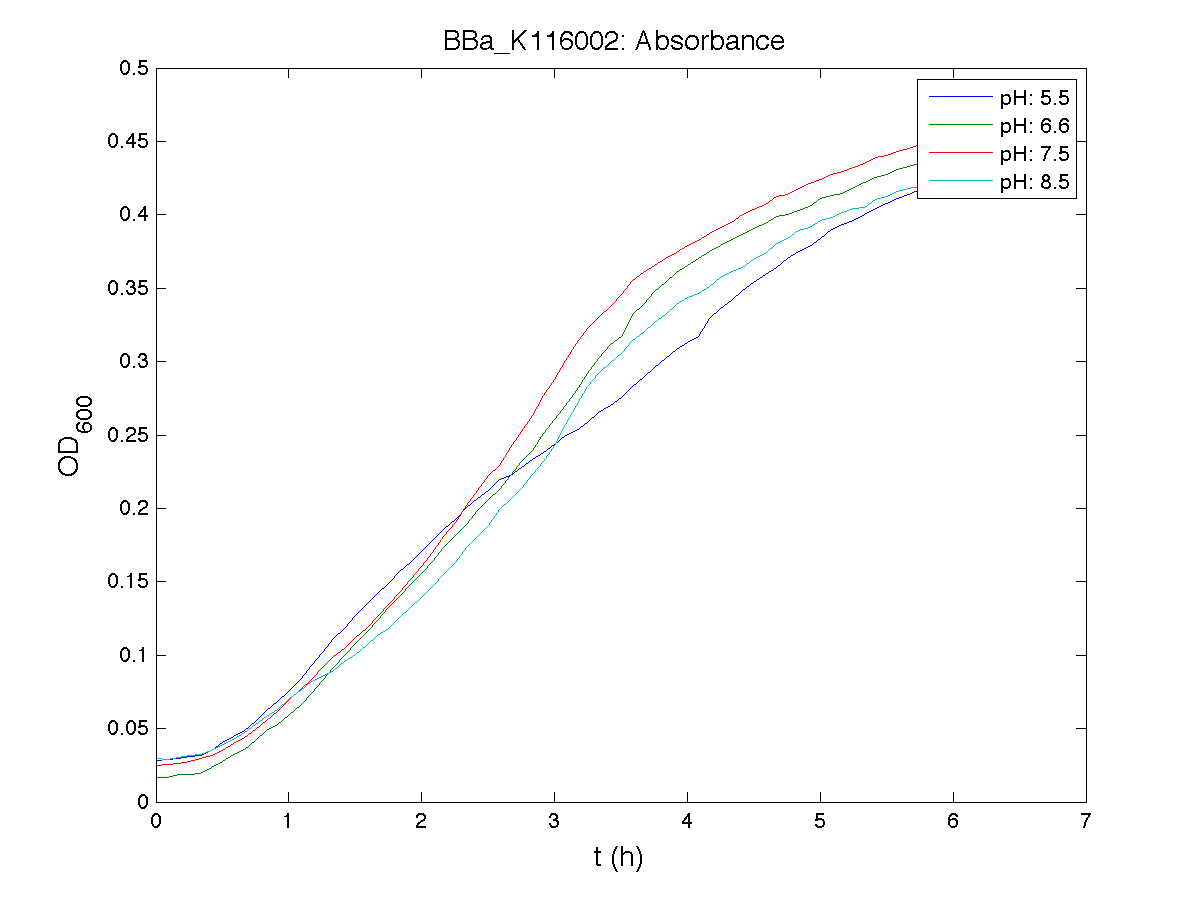
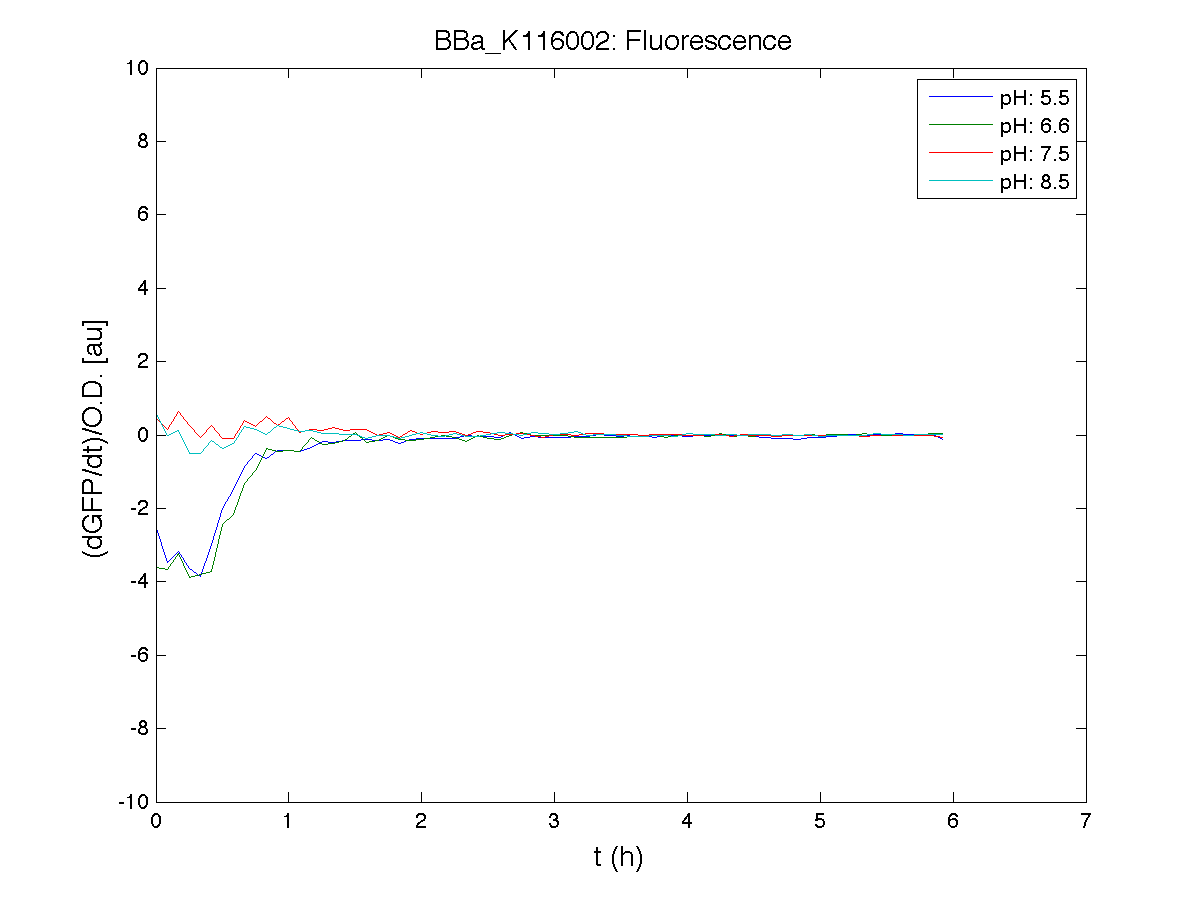
ConclusionsThe induction curve of the receiver device, reported in page (M9 supplemented medium), was represented in PoPS units, while ours is reported in RPUs and has been obtained through a very similar protocol (see Growth conditions section). Anyway, the experiments we performed in M9 supplemented medium confirmed the induction curve shape of this device, with a switch point between ? and ?. We also estimated this transfer function of this device in LB medium, for which no data were reported in the Registry. BBa_K116001 - nhaA promoterfrom iGEM 2008 NYMU-Taipei We received this BioBrick from iGEM in September but the bacterial strain that contained the plasmid wasn't declared. So we decided to sequence it as check and transform it into E.coli TOP10. We wanted to perform some experiments to better understand how it works and if can be successfully used. We performed several experiments with different LB medium and we got almost the same results. We used:
Here we show just two experiments to explicate our work. You can download the complete list from this link. Experiment Na+ 0MMotivationIn our opinion the working principle of the antiporter Na+/H+ channel described in [Rachel Karpel et al., Etana Padan et al., N. Dover et al.] makes the nhaA promoter a Na+ sensor and only under certain conditions (presence of Na+) a pH sensor. Methods
ResultsCommentsAs expected didn't produce any GFP. So we can consider it a Na+ sensor and only secondarily a pH sensor. Experiment Na+ 250mMMotivationWe’ll try again to make E.coli producing GFP at the variation of pH. Methods
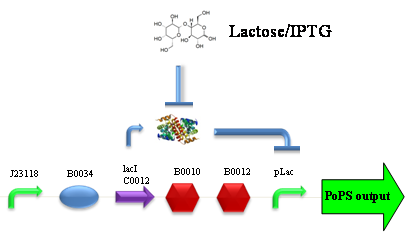
ResultsCommentsWe didn't expect this. After looking better for a motivation in some articles ([Rachel Karpel et al.]) we think this could be because of the E.coli strain: we use TOP10 while a special strain (delta-pump) without some membrane proteins that regulate E.coli homeostasis is used in other experiments. Final considerationsIn our opinion this sensor (primarily sodium sensor and secondarily pH sensor) needs very particular conditions to work (first of all a specific bacterial strain) we couldn’t reproduce, so we consider it almost unusable. BBa_K112808 - Enterobacteria phage T4 Lysis DeviceDescriptionCharacterizationClonclusionsBBa_R0011 - Plac hybrid promoterDescriptionThe hybrid lac promoter (BBa_R0011) has been designed taking the Plambda promoter (BBa_R0051) and substituting its cI (BBa_C0051) binding sites with two lacI binding sites. This promoter can be repressed by lacI (BBa_C0012), which can be repressed by lactose or IPTG, providing a lactose/IPTG inducible system. Differently from wild type lac promoter, this part does not have any CAP binding sites, so its behaviour is glucose-independent. Even if lacI is not expressed in this BioBrick, strains bearing a genomic copy of lacI can repress this promoter, which acts as a glucose-independent lactose/IPTG sensor. In the other strains BBa_R0011 acts as a constitutive promoter. Here we provide the characterization of this promoter in E. coli TOP10, which has a lacI genomic copy, constitutively expressed in a weak manner. The data below are referred to , which is the measurement system of . Characterization
ConclusionsExisting parts: sequence debuggingDuring this summer, we sequenced and carefully analyzed sequencing results of some existing BioBrick parts. Some of them were not present in DNA Distribution and we received them from iGEM HQ. All these comments are also reported in the Experience pages of the relative BioBrick parts in the Registry web site.
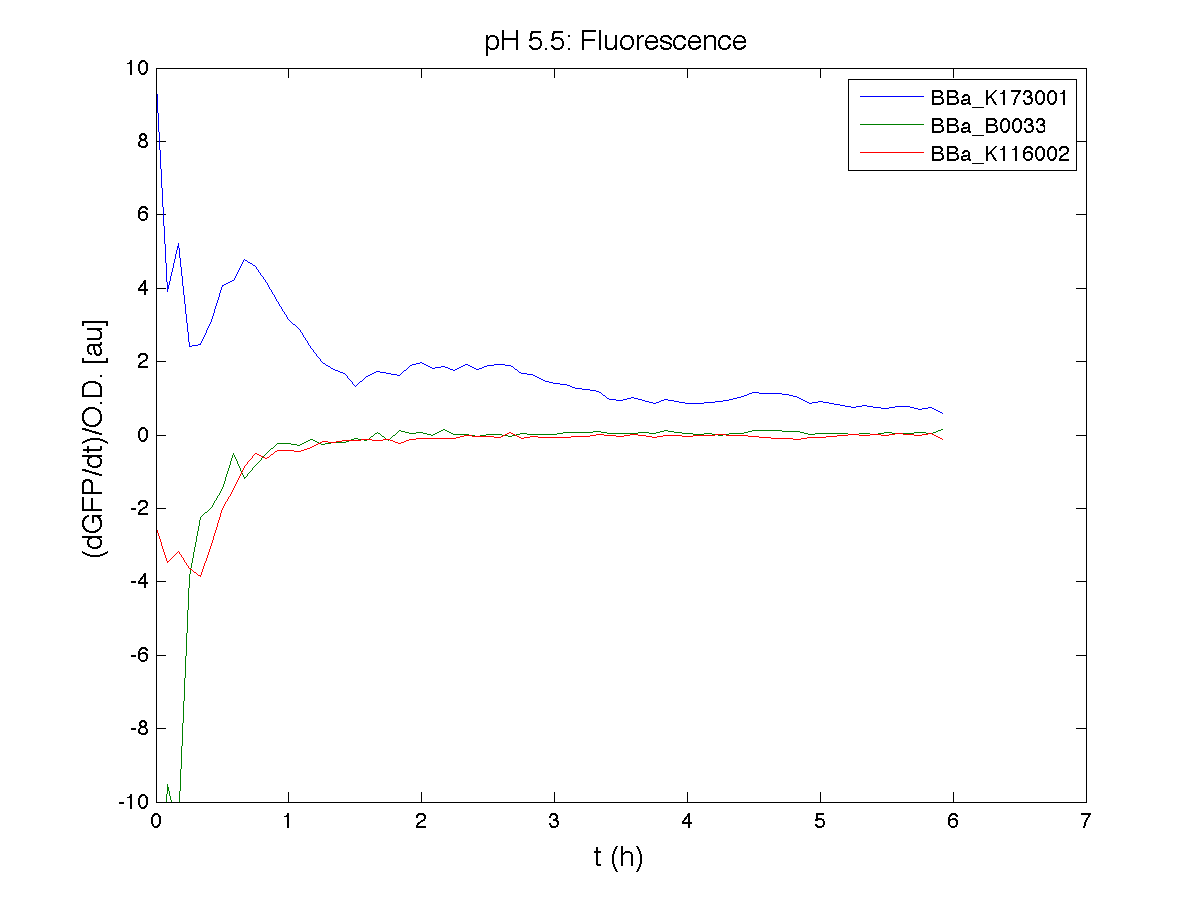
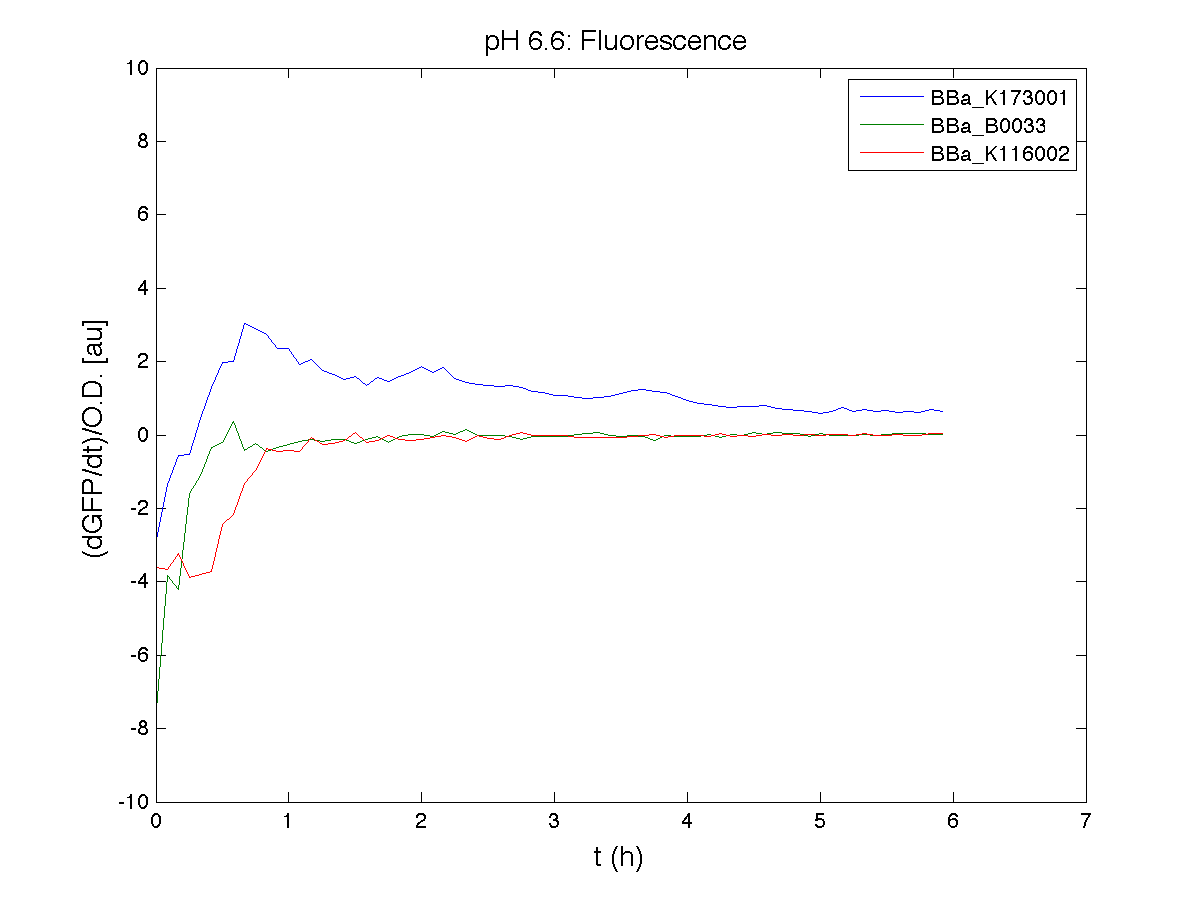
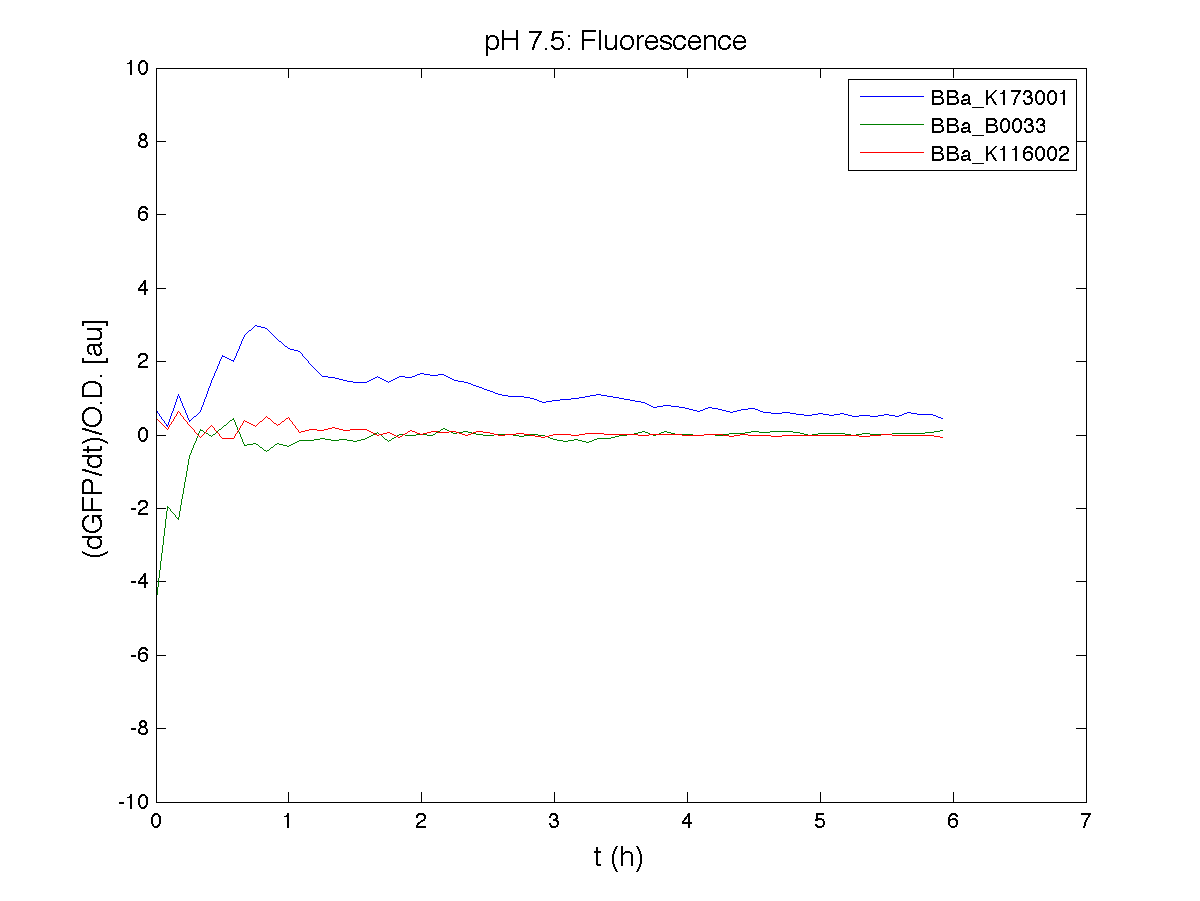
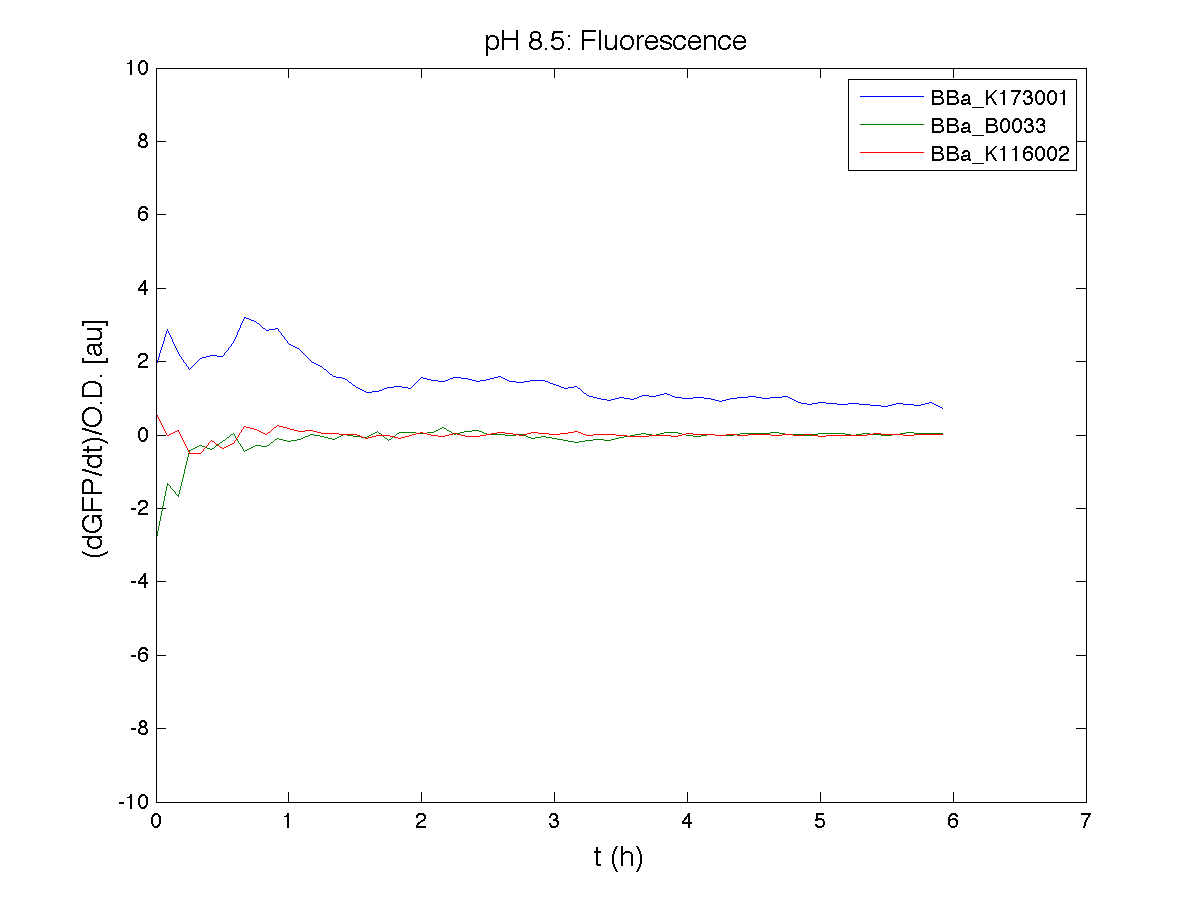
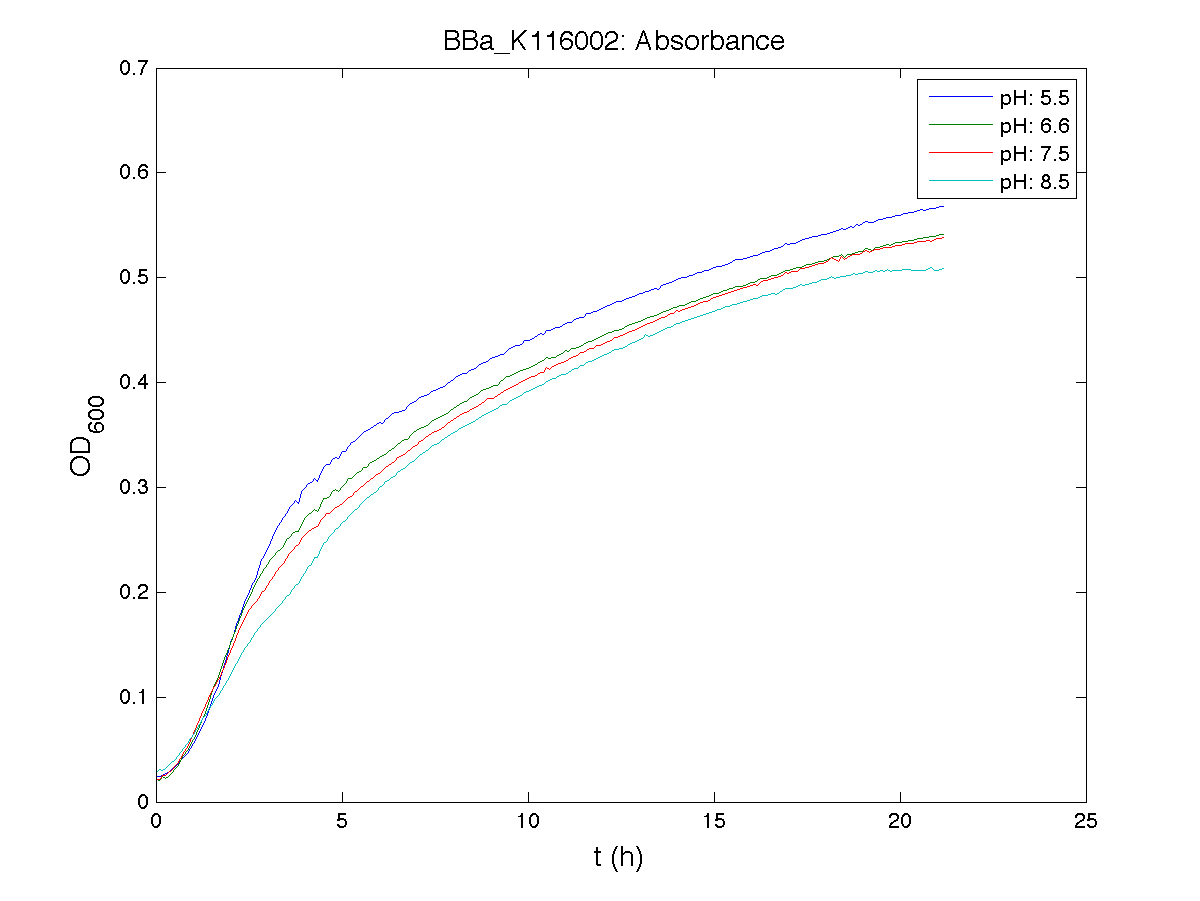
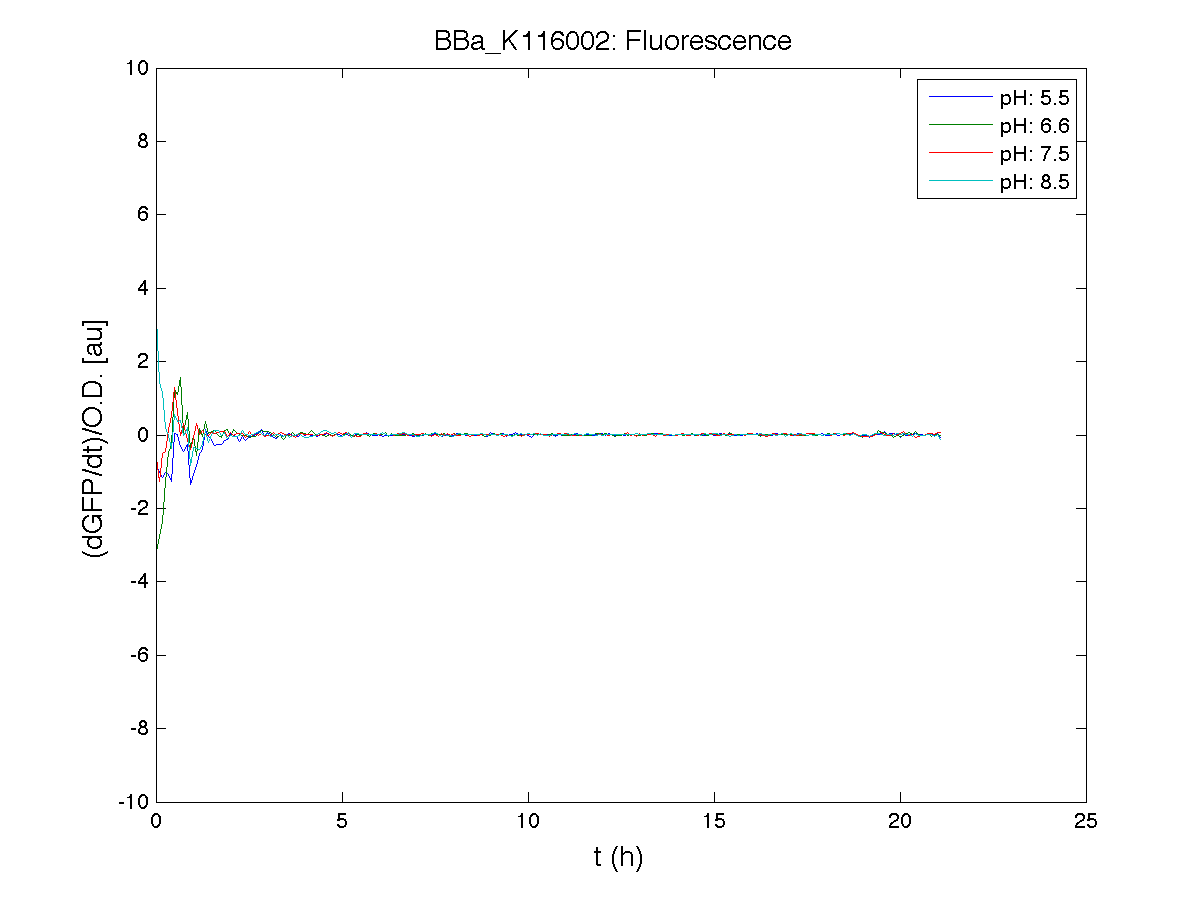
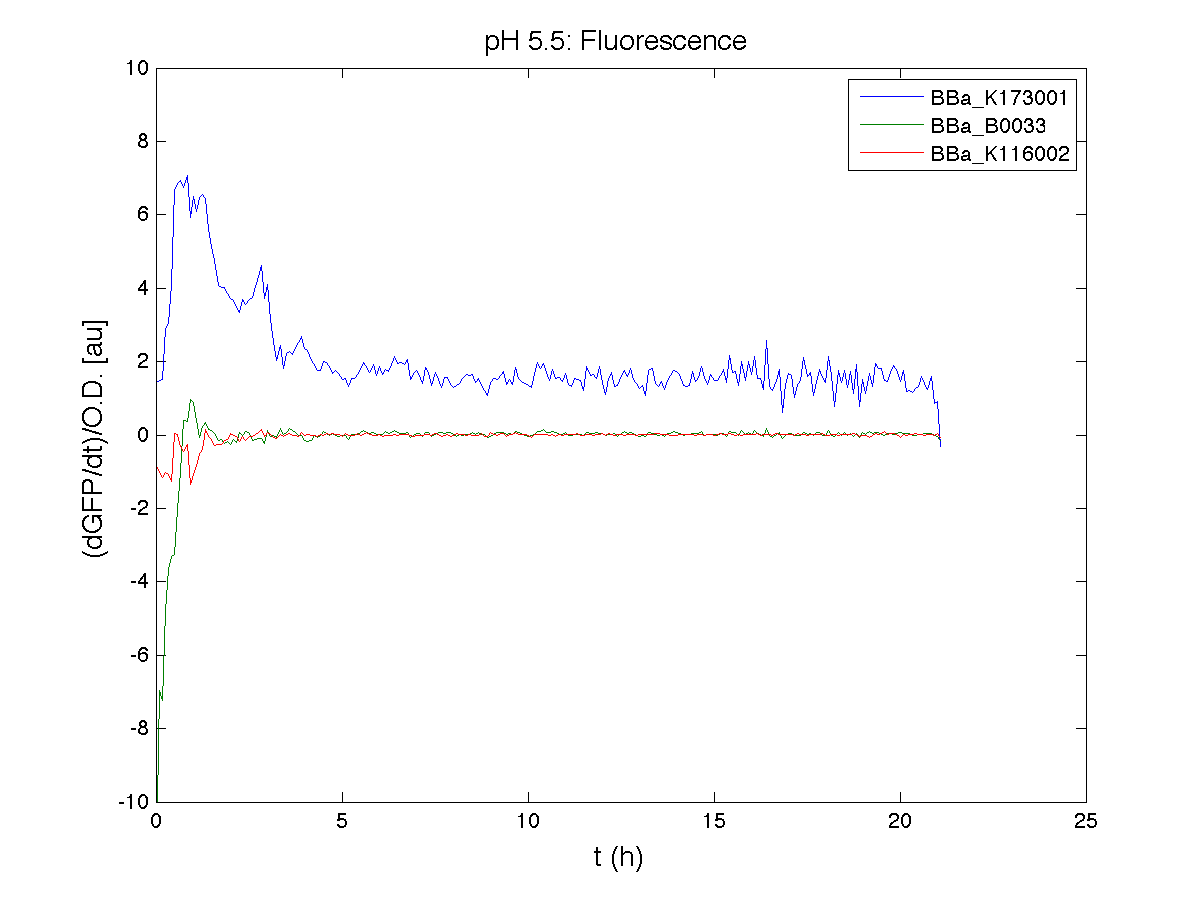
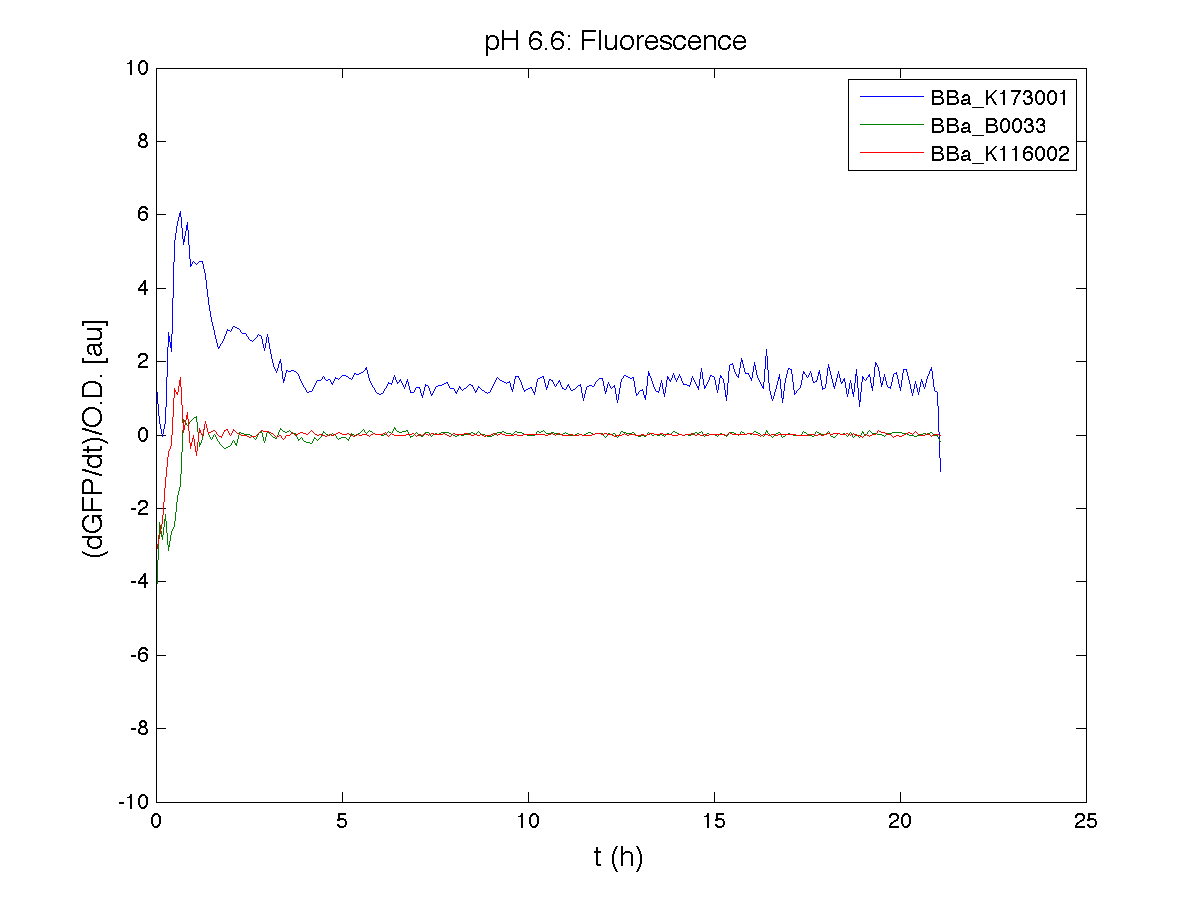
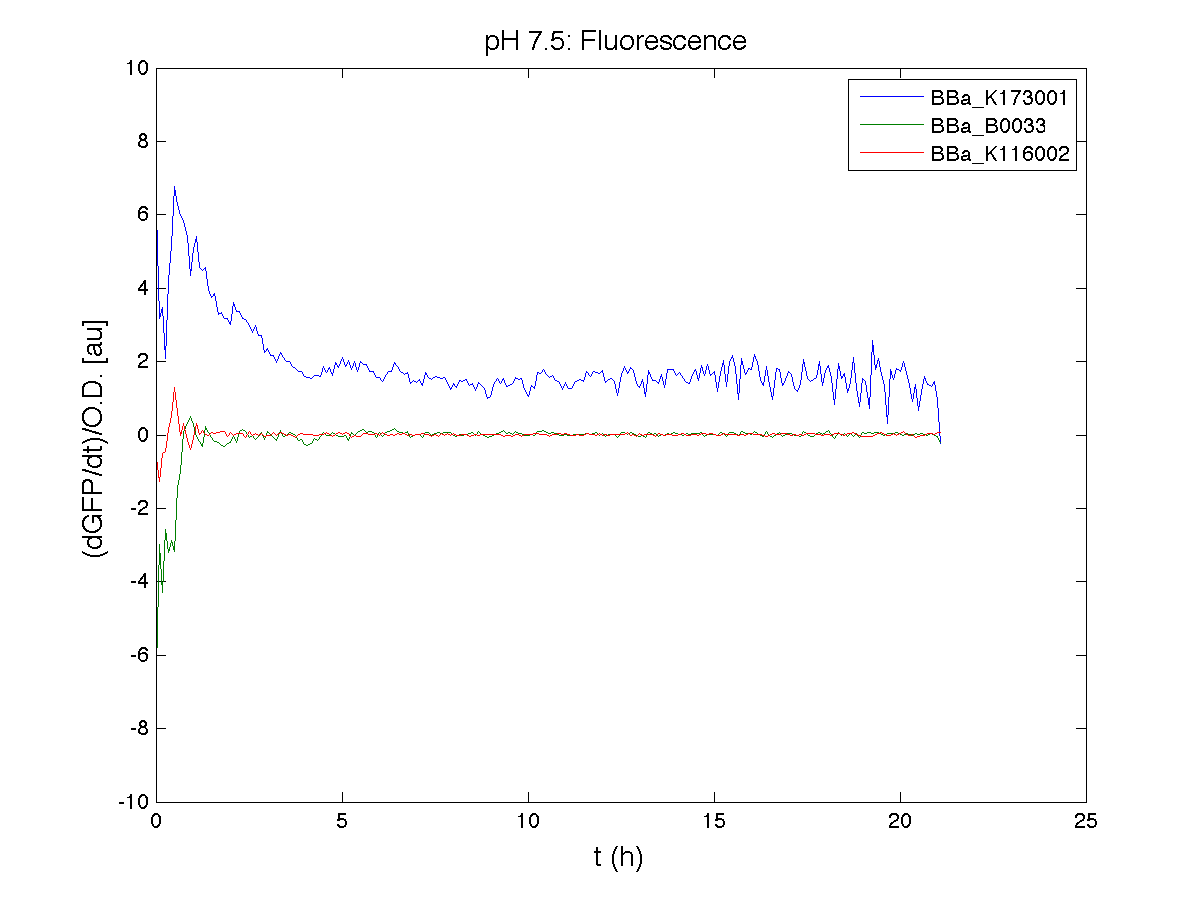
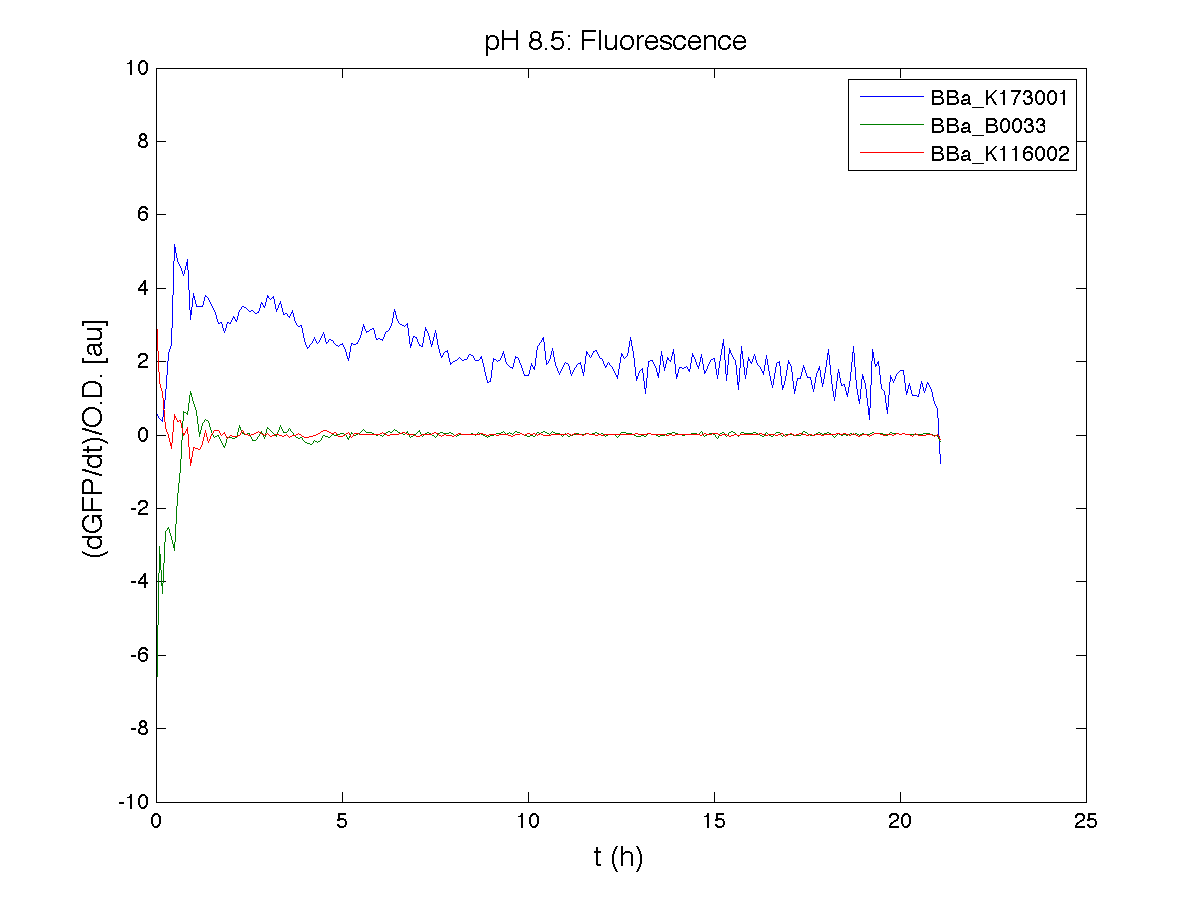
Because BBa_K116002 is the measurement system of this BioBrick, we used it to characterize the activity of nhaA promoter (see Application section above).
However, the part should work properly because the regulatory parts and the amino acid sequence of the genes are correct. This has also been confirmed by functional tests (see Applications section).
Growth conditionsMicroplate reader experiments
Fermentation experimentsPROTOCOL#1
PROTOCOL#2
PROTOCOL#3
Data analysisGrowth curvesThe presented growth curves have all been processed as OD600_culture-OD600_broth for each time sample. OD600_broth is the medium in the same conditions as in the culture (e.g. induced with the same inducer concentration as in the culture). Doubling timeThe natural logarithm of the growth curves (processed according to the above section) was computed and the linear phase (corresponding to the bacterial exponential growth phase) was isolated by visual inspection. Then the linear regression was performed in order to estimate the slope of the line m. Finally the doubling time was estimated as d=ln(2)/m [minutes]. In the case of multiple growth curves for a strain, the mean value of the processed curves was computed for each time sample and then this procedure was performed. Relative Promoter Units (RPUs)The RPUs are standard units proposed by Kelly J. et al., 2008, in which the transcriptional strength of a promoter can be measured using a reference standard, just like the ground in electric circuits. RPUs have been computed as: in which:
RPU measurement has the following advantages:
The hypotheses on which RPU theory is based can be found in Kelly J. et al., 2008, as well as all the mathematical steps. From our point of view, the main hypotheses to satisfy are the following:
Inducible systemsEvery experiment is performed on the following cultures:
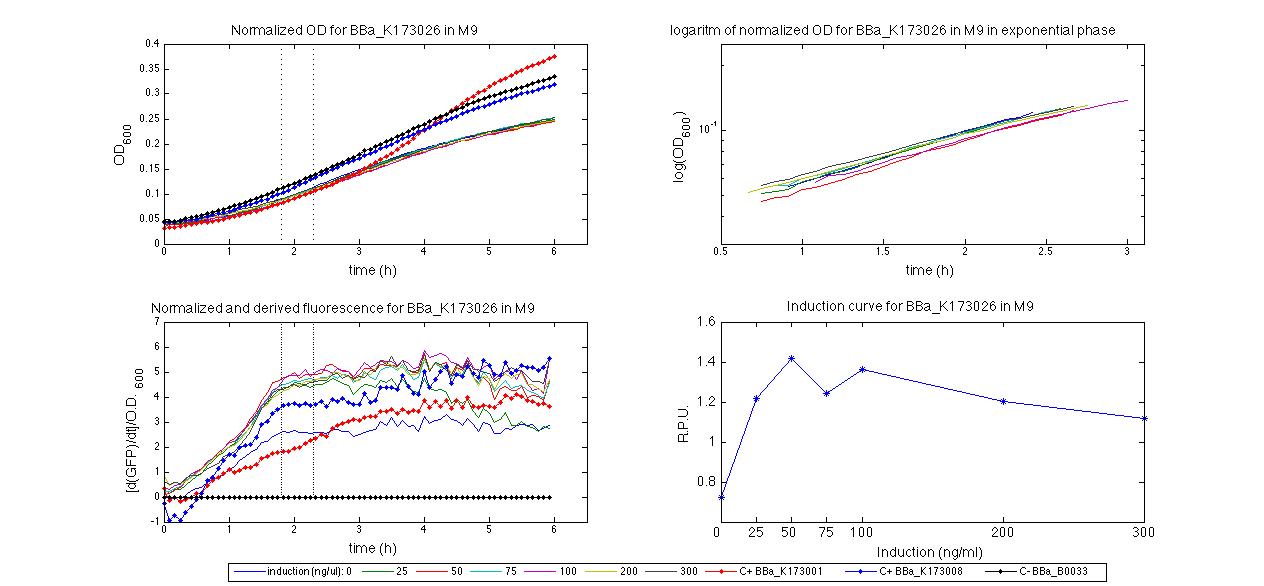
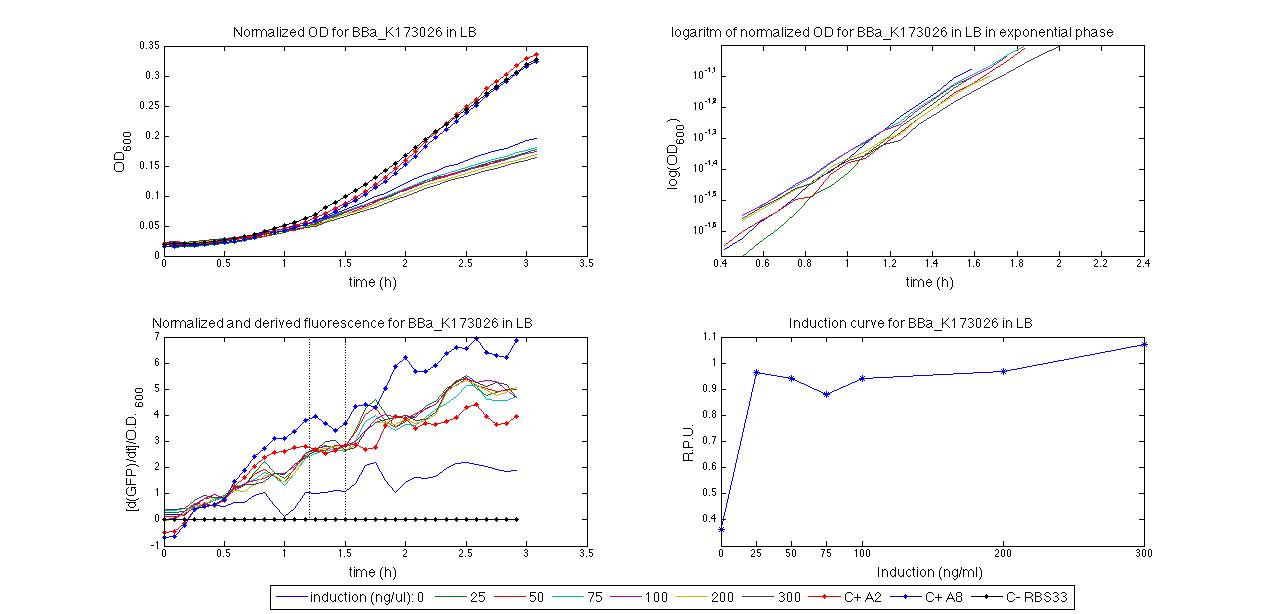
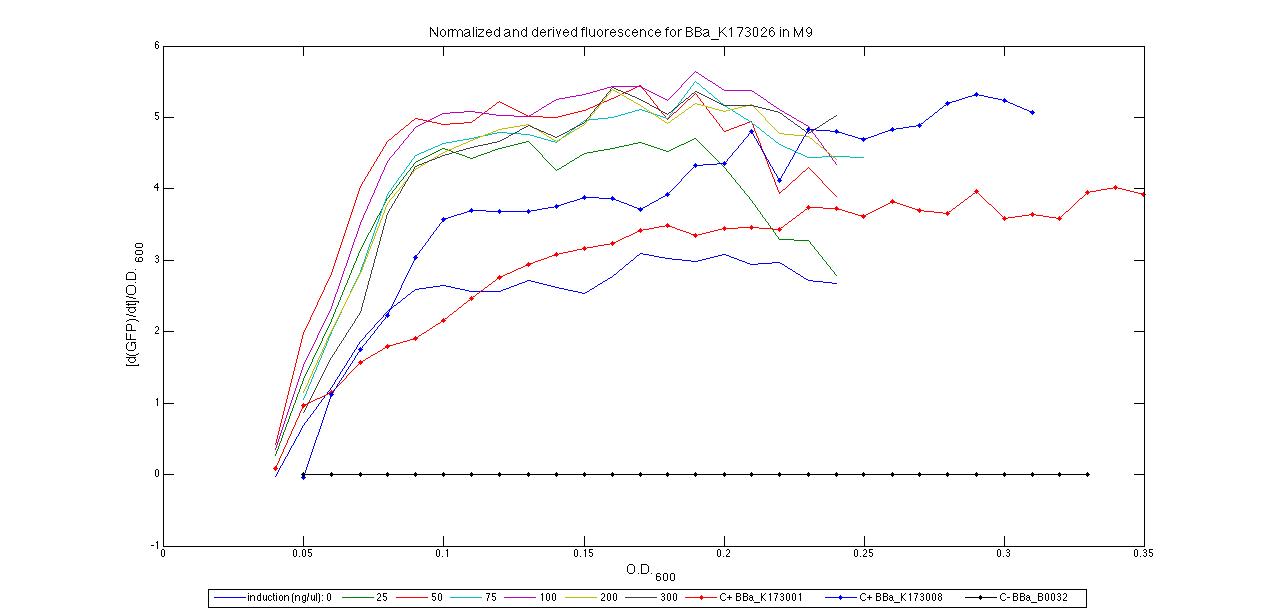
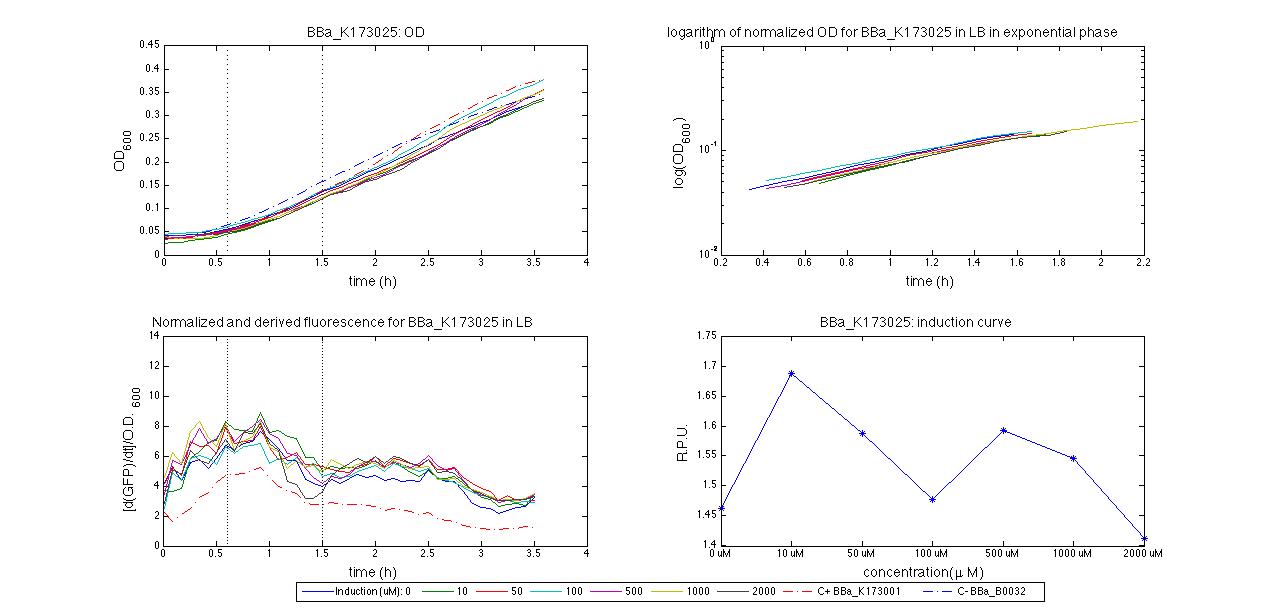
For inducible systems several plots are reported. The first plot is a panel containing 4 subplots, numerated this way:
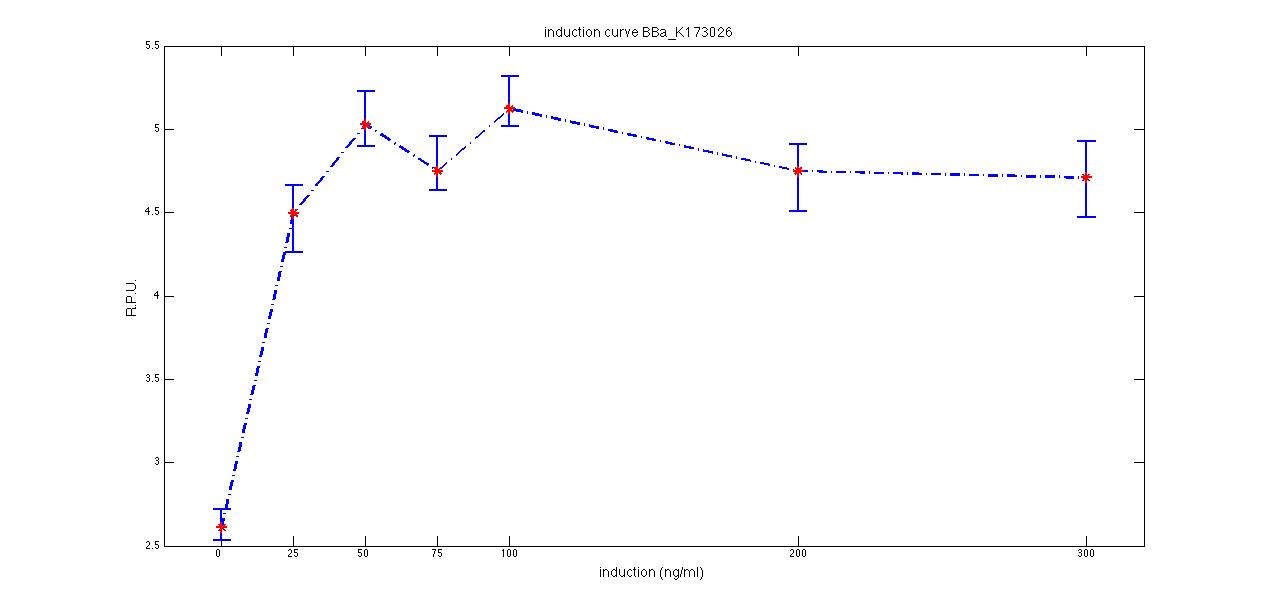
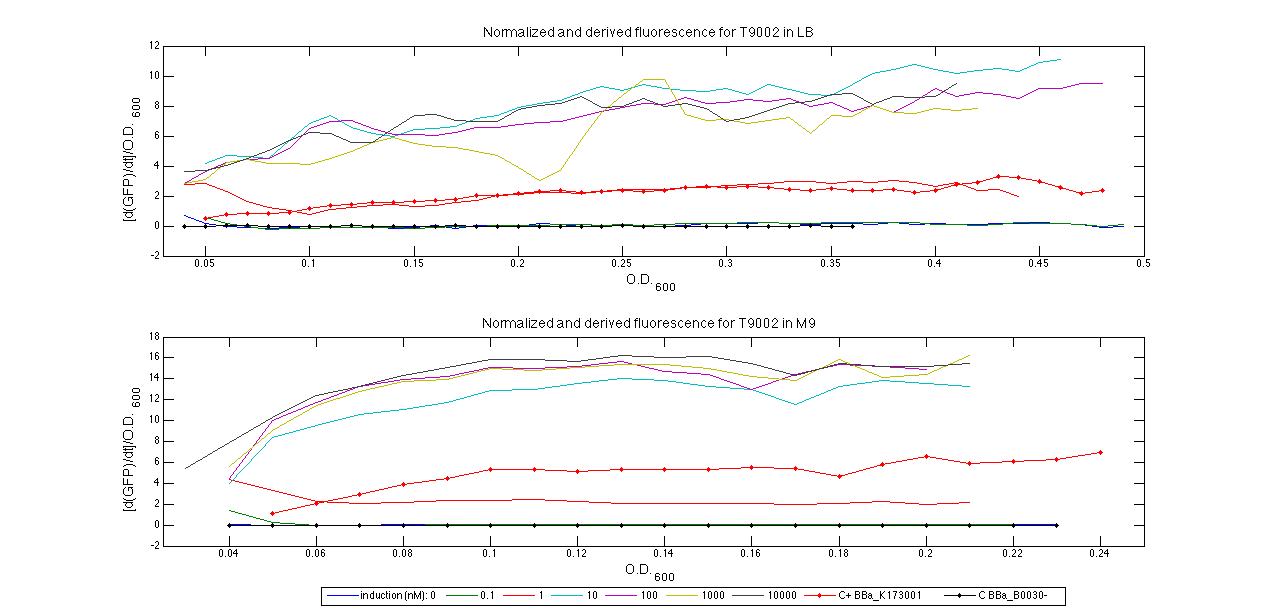
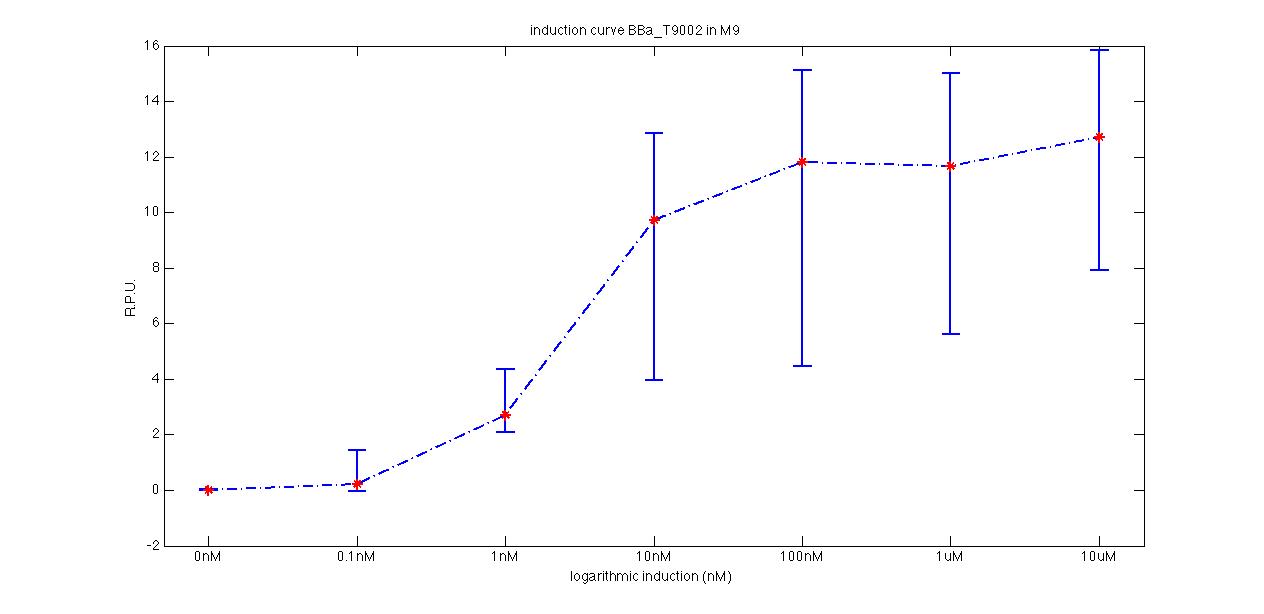
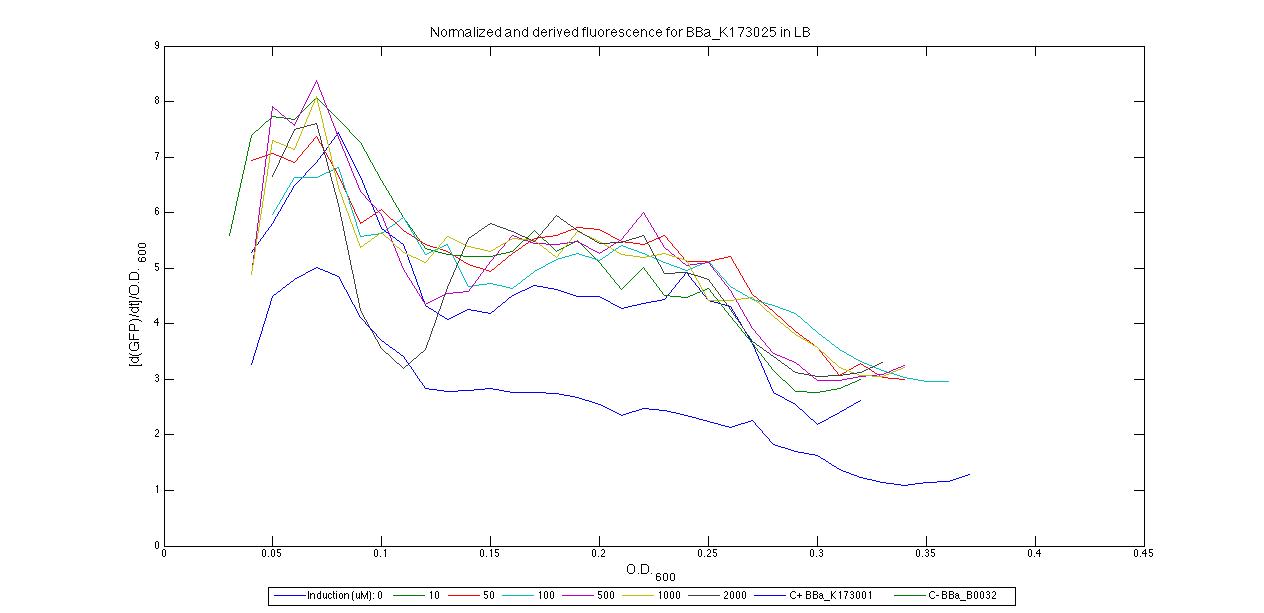
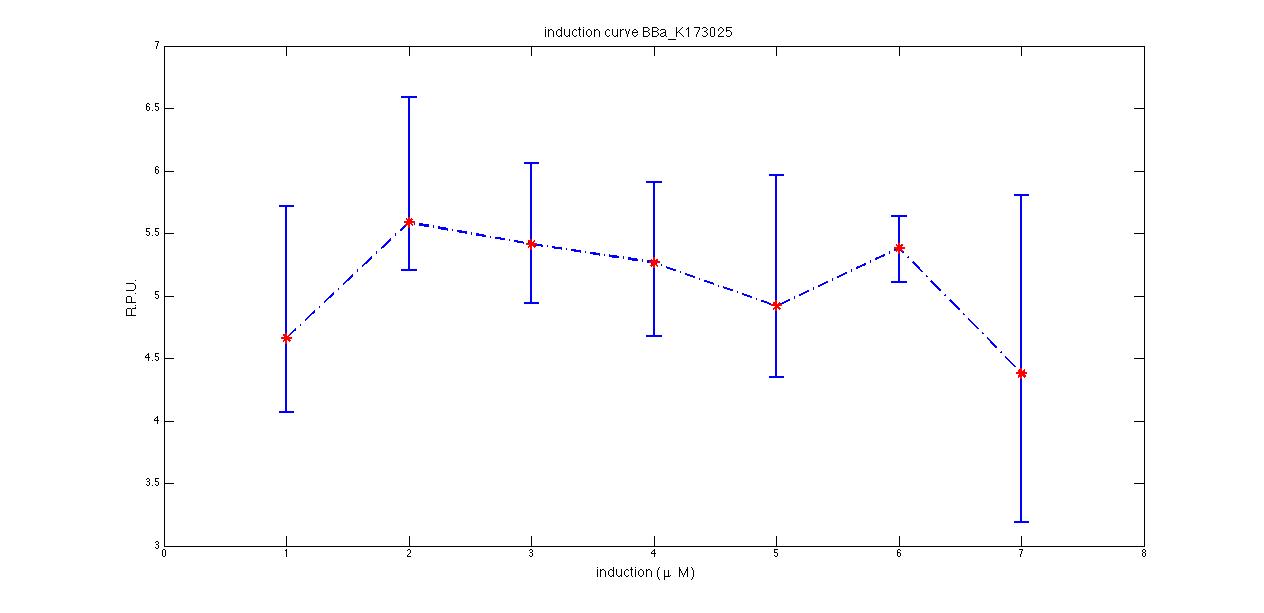
Plot (1) contains growth curves of the cultures, after blank value has been removed. Every curve is calculated averaging on three replicates of the same culture and subtracting the blank for each time sample. Blank is calculated averaging the replicates of blank wells. Plot (2) shows the logarithm of absorbance in exponential phase of bacterial growth, determined by a visual inspection of log-plots. These values are used to evaluate doubling time and R.P.U.. Plot (3) contains (dGFP/dt)/OD, the value named Scell in Canton procedure for RPU evaluation. The last plot (4) contains the induction curve, showing the R.P.U. value for every inductor concentraion. In these plots are reported black veritcal lines that define the range of values used to evaluate RPU. It is important to underline, as explained in next paragraph, that RPU are calculated on cultures at the same OD level, not at the same time. The second graphic shows Scell VS O.D.. This plot allows the conparison of Scell values between different cultures, that are supposed to reach the same level of growth not at the same time, but at the same O.D. value. The third graphic shows the induction curve. The RPU value is calculated on Scell values corresponding to OD values in exponential phase (tpitcally, from 0.05 to 0.16). The curve is obtained averaging in time Scell values corresponding to exponential phase. Error bars rapresent the minimum and maximum value of R.P.U. belonging to the range of O.D. in exponential phase. In RPU evaluation the hypothesis of steady state has to be validated. This hypothesis corresponds to a constant behavior of Scell in time. In exponential phase in several cases it is possible to observe that this variable isn't constant, but grows after exponential phase is over. This behaviour is totally unexpected and can't be justifyed by any biological argument. It is also important to undelrine that the reported methodology has shown how variable R.P.U. value can be. This parameter, in fact, is very sensitive to the respondind OD value, as shown from induction curves, where error bars are sometimes wide among the curve. So it is foundamental to define a standardized methodology for RPU evaluation, not sensitive to OD or time choose. Materials
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"