Team:Virginia Commonwealth/Design
From 2009.igem.org
(→UP-element design) |
(→UP-element design) |
||
| Line 13: | Line 13: | ||
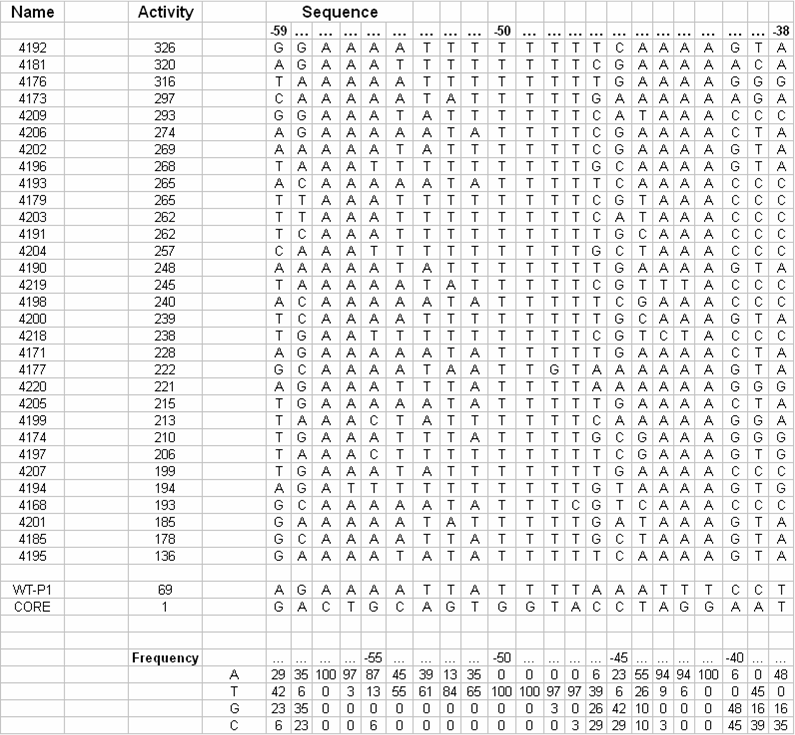
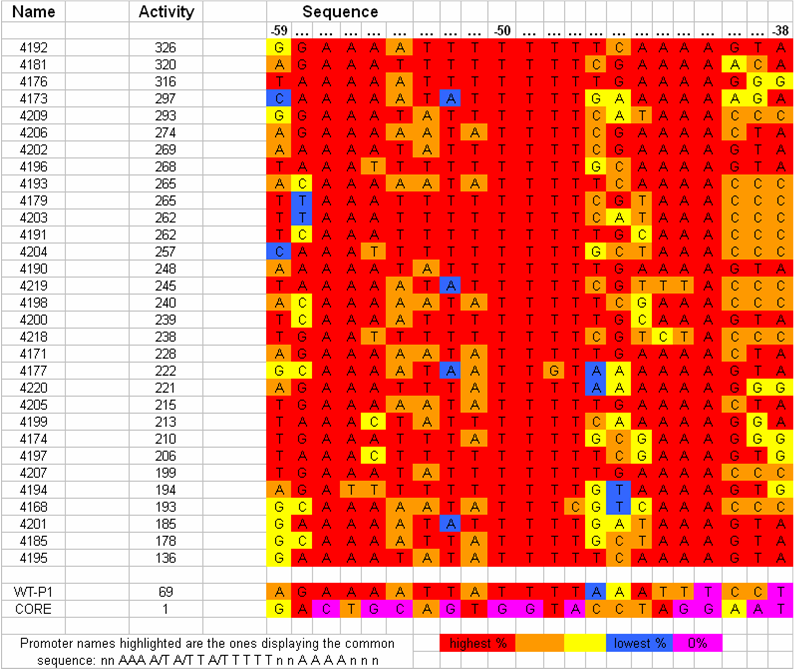
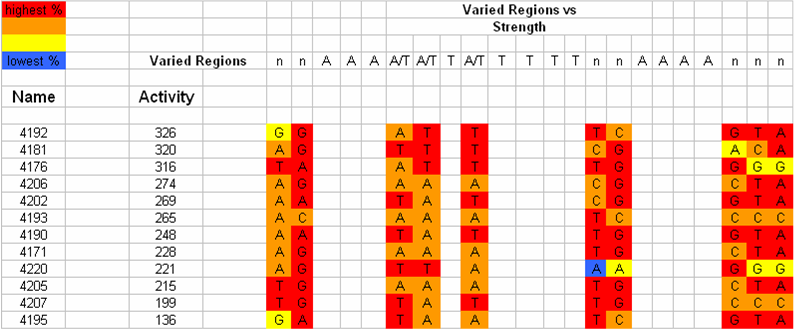
[[Image:VCU 2009 UP-element tabulated nucleotide frequency.png|thumb|center|650px|Tabulation of UP-element sequences in order of observed strengths and correlation with nucleotide frequency]] | [[Image:VCU 2009 UP-element tabulated nucleotide frequency.png|thumb|center|650px|Tabulation of UP-element sequences in order of observed strengths and correlation with nucleotide frequency]] | ||
| - | Data showed direct correlation between the strength of naturally occurring UP elements and the frequency of each nucleotide at each respective position. Thirty one UP element sequences were analyzed for nucleotide frequency and the relative activity was tabulated for each sequence (Estrem, ''et. al'') From this a consensus UP element sequence was identified which was composed of the most frequently occurring nucleotide at each position. This consensus sequence is theoretically the strongest UP element sequence. | + | Data showed direct correlation between the strength of naturally occurring UP elements and the frequency of each nucleotide at each respective position. Thirty one UP element sequences were analyzed for nucleotide frequency and the relative activity was tabulated for each sequence (Estrem, ''et. al''). From this a consensus UP element sequence was identified which was composed of the most frequently occurring nucleotide at each position. This consensus sequence is theoretically the strongest UP element sequence. |
Revision as of 18:10, 21 October 2009
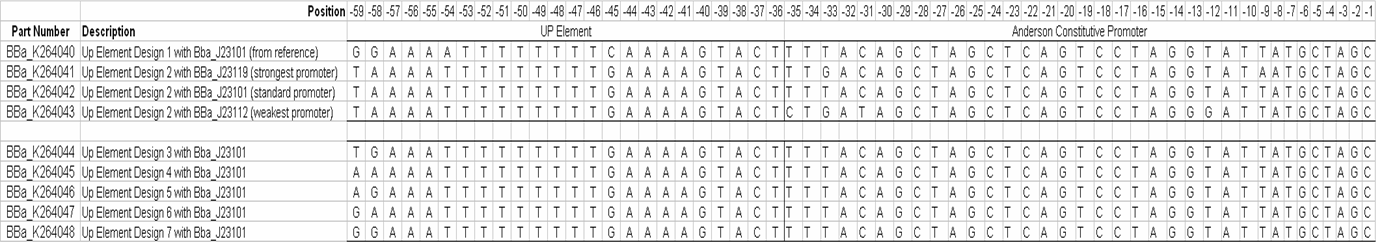
UP-element designUP-elements are known to significantly increase the RNA polymerase-recruiting power of promoters by interacting with the alpha subunit of RNA polymerase. Work has been done to identify a consensus sequence for this transcriptional enhancer based on 31 natural E. coli promoters (Estrem ST, Gaal T, Ross W, Gourse RL, Identification of an UP element consensus sequence for bacterial promoters, PNAS, (1998), 95, 9761-9766.).
We are working on creating this UP element as a modular BioBrick part and characterizing its activity using the promoter characterization method that we develop. We will also manipulate this element using a bottom-up design approach to get UP elements of varying degree's of strength. Thus increasing the level of control we have over gene expression. Data showed direct correlation between the strength of naturally occurring UP elements and the frequency of each nucleotide at each respective position. Thirty one UP element sequences were analyzed for nucleotide frequency and the relative activity was tabulated for each sequence (Estrem, et. al). From this a consensus UP element sequence was identified which was composed of the most frequently occurring nucleotide at each position. This consensus sequence is theoretically the strongest UP element sequence.
Design strategiesPromoter design will be approached from the bottom up. First, a consensus promoter sequence will be studied and accepted by researching the work done by others on various promoters. This consensus will be our starting point as we identify which nucleotide and nucleotide sequences are most important to RNAP binding affinity. We will use direct synthesis to construct promoter sequences that we will test. Future work
| |
 "
"