Team:USTC/Project
From 2009.igem.org
(→Principle of Operation) |
(→Principle of Operation) |
||
| Line 179: | Line 179: | ||
|} | |} | ||
What if we mix them together: | What if we mix them together: | ||
| - | [[Image: | + | [[Image:mixhl.jpg|500px|center]] |
---- | ---- | ||
Revision as of 20:05, 21 October 2009
| Home | Team | Project | Modeling | Parts | Standard & Protocol | Software Tool | Human Practice | Notebook |
|---|
Team:USTC/Project
Contents
|
The Background
Evolution vs. Design
Evolution and design are two sides of the coin.
Until 150 years ago, people believed that we are designed and created by the God, a god, gods or other [http://en.wikipedia.org/wiki/Intelligent_designer intelligent designer]s. After [http://en.wikipedia.org/wiki/Charles_Darwin Charles Darwin]'s [http://en.wikipedia.org/wiki/On_the_Origin_of_Species On the Origin of Species] in 1859, more and more people began to realize that a god is not necessary. During the process of evolution, we can be created automatically, without any intelligence.
[http://en.wikipedia.org/wiki/Design Design] and [http://en.wikipedia.org/wiki/Engineering engineering] are processes with the need of knowledge. Before we can design a machine, we have to know everything about the possible parts. If any information is unknown, we have to learn it. If nobody knows it, we have to measure it by ourselves. If all the knowledge needed are too much, we have to collaborate with others. [http://en.wikipedia.org/wiki/Computer-aided_design Computer-aided design] have to be used to accomplish more complex tasks. [http://en.wikipedia.org/wiki/Space_Shuttle Space shuttle] and [http://en.wikipedia.org/wiki/Microprocessor microprocessor] are seen as the most complex systems engineered, but they are far too simple when compared with the [http://en.wikipedia.org/wiki/Complexity complexity] of biological systems created by evolution.
On the other side, [http://en.wikipedia.org/wiki/Evolution evolution] is a completely different process. Variation is random, selection is directional based on the fitness to the environment, that is all. However, all the amazing things on our planet is emerged from this simple process, no more thing is needed.
Why? All the complexity is solved by this simple process without any input of knowledge?
Yes. The answer is on the scale. The evolution process is so simple that it can be scaled up infinitely. The complex problem can be solved in the distributed evolution system. Any success in the distributed system can be amplified by the selection process. Therefore, although the variation is random, it will be powerful enough to search for the solutions, when the scale of the system is big enough.
In the design process, the variation is directional based on the knowledge, but the process is not scalable, because of its requirement of intelligence. As a result, it is much more difficult to solve complex problems by design.
Directed Evolution
Although Darwin's theory was published 150 years ago, people have used the power of evolution more than ten thousand years. [http://en.wikipedia.org/wiki/Domestication Domestication] of plants and animals is done by [http://en.wikipedia.org/wiki/Artificial_selection artificial selection], which is the process of intentional breeding for certain traits, therefore changing the direction of evolution.
[http://en.wikipedia.org/wiki/Directed_evolution Directed evolution] is a method using the similar principle to create new biomolecules with desired properties. The targets of directed evolution include enzymes, antibodies, [http://en.wikipedia.org/wiki/Aptamer aptamer]s, [http://en.wikipedia.org/wiki/Ribozyme ribozyme]s, biosynthetic pathways, and synthetic genetic circuits. More information can be found on [http://www.che.caltech.edu/groups/fha/home.html the Frances H. Arnold research group], [http://ellingtonlab.org the Ellington lab], and [http://genetics.mgh.harvard.edu/szostakweb/ the Szostak lab].
The directed evolution experiment contains several rounds of 3 steps: variation, selection, and amplification. Variation is the mutation or recombination of the information encoded in the DNA, usually by error-prone PCR and [http://en.wikipedia.org/wiki/DNA_shuffling DNA shuffling] respectively. Selection is the process of separating the variants with desired phenotypes from others, it can either refer to screening (isolate good variants) or selection (eliminate bad variants), either in vivo or in vitro. Amplification is the replication of the variants after selection, which recovers the population size for the new round of directed evolution.
New function of biomolecules or biological systems is difficult to be rationally designed, but directed evolution is successfully used in solving these problems.
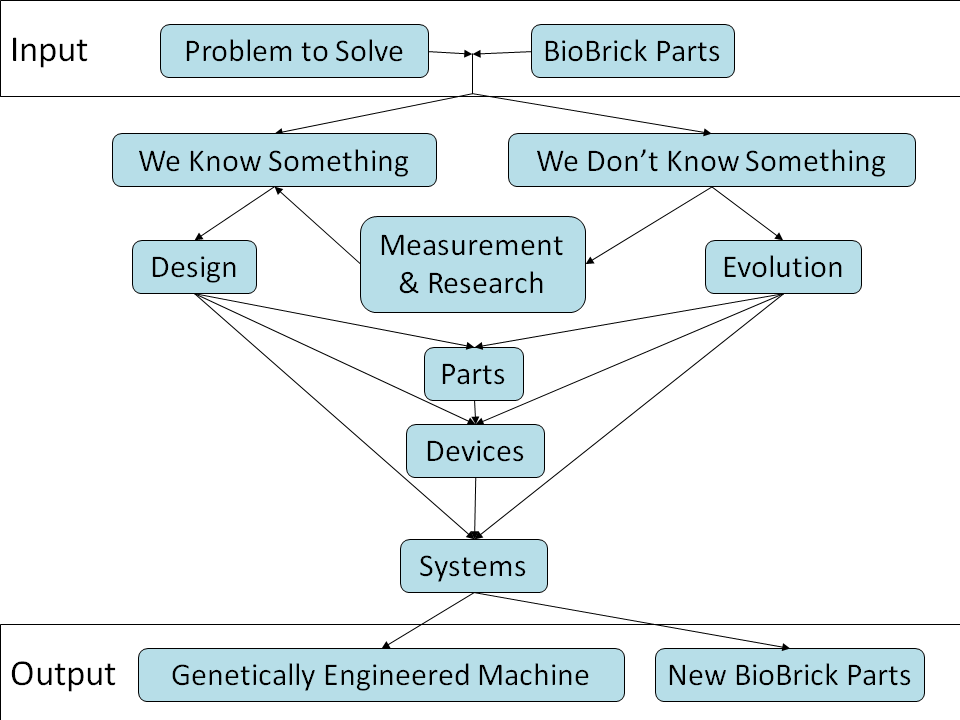
Design & Evolutionary Approaches in iGEM Projects
In the iGEM competition, teams specify, design, build, and test simple biological systems made from standard, interchangeable biological parts. Most projects engineer the systems by modeling and measurement, including this project itself. This engineering approach is proved to be feasible but difficult in the design of biological systems during this years.
Most BioBrick parts have never been characterized, because characterization and documentation of the parts usually takes a lot of time. Further more, the properties of BioBrick parts and devices are very sensitive to the conditions they work in, such as strain, plasmid, other parts used together, culture medium, growth rate, temperature, pH, shaking rate, light, and so on. It is very difficult to characterize all the parameters, so the parts often have to be remeasured in different projects before the behavior of the system can be predicted.
There are also many projects using evolutionary approaches. These approaches use selection or screening to find the needed part, eliminate the steps of measurement or constructing new parts with desired properties.
| Team Name / People | Competition / Lab | Project | Approach | Wiki Link |
|---|---|---|---|---|
| Davidson | iGEM 2006 | Solving the Pancake Problem with an E. coli Computer | in vivo Recombination; Selection | [http://parts2.mit.edu/wiki/index.php/Davidson_2006 Davidson] |
| Missouri_Western | iGEM 2006 | Solving the Pancake Problem with an E. coli Computer | in vivo Recombination; Selection | [http://parts2.mit.edu/wiki/index.php/Missouri_Western_State_University_2006 Missouri_Western] |
| Alberta | iGEM 2007 | Plan B: Building Butanol BioBricks | Mutagenic Compound; Selection | [http://parts.mit.edu/igem07/index.php/Alberta Alberta] |
| Boston_University | iGEM 2007 | Increasing the Current Output of Microbial Fuel Cells Through the Directed Evolution of Shewanella oneidensis | Error-Prone PCR; Screening | [http://parts.mit.edu/igem07/index.php/Boston_University Boston_University] |
| Davidson_Missouri_W | iGEM 2007 | Living Hardware to Solve the Hamiltonian Path Problem | in vivo Recombination; Screening | [http://parts.mit.edu/igem07/index.php/Davidson_Missouri_W Davidson_Missouri_W] |
| Harvard | iGEM 2007 | Cling-E. coli: Bacteria on target | Screening | [http://parts.mit.edu/igem07/index.php/Harvard Harvard] |
| USTC | iGEM 2007 | Extensible Logic Circuit in Bacteria | Error-Prone PCR; Screening | [http://parts.mit.edu/igem07/index.php/USTC USTC] |
| Calgary_Software | iGEM 2008 | EvoGEM: An Evolutionary Approach to Designing Life | Evolutionary Algorithm | Calgary_Software |
| ETH_Zurich | iGEM 2008 | Make yourself simpler, stupid! Or how engineering a self-minimizing cell leads to the Minimal Genome | Controlled expression of restriction enzymes and ligases in vivo; Selection | ETH_Zurich |
| IIT_Madras | iGEM 2008 | StressKit: A BioBrick library of Lac-repressed σ24, σ28, σ32 and σ38 promoters for Escherichia coli | Screening | IIT_Madras |
| Illinois | iGEM 2008 | Cell-based and in vitro antigenic sensors for medical diagnostics | Error-Prone PCR; Screening | Illinois |
| Newcastle_University | iGEM 2008 | A Computational Intelligence Approach to Developing a Diagnostic Biosensor: The Newcastle BugBusters Project | Evolutionary Algorithm | Newcastle_University |
| Peking_University | iGEM 2008 | A genetic circuit to direct evolution of proteins in vivo | in vivo Mutagenesis; Screening | Peking_University |
| Tokyo_Tech | iGEM 2008 | Coli.Touch – implementation of a pressure-responsive genetic circuit in E. Coli | Error-Prone PCR; Screening | Tokyo_Tech |
| Warsaw | iGEM 2008 | Bacterial device for creating and production of interactors for any given bait protein | in vivo Mutagenesis; Selection | Warsaw |
| Jason R. Kelly | Endy Lab | Screening plasmid | Screening | [http://openwetware.org/wiki/Endy:Screening_plasmid Endy:Screening_plasmid] |
Evolutionary Biology, Population Genetics & Evolutionary Algorithm
Problems to Be Solved
The Blueprint
The Goal
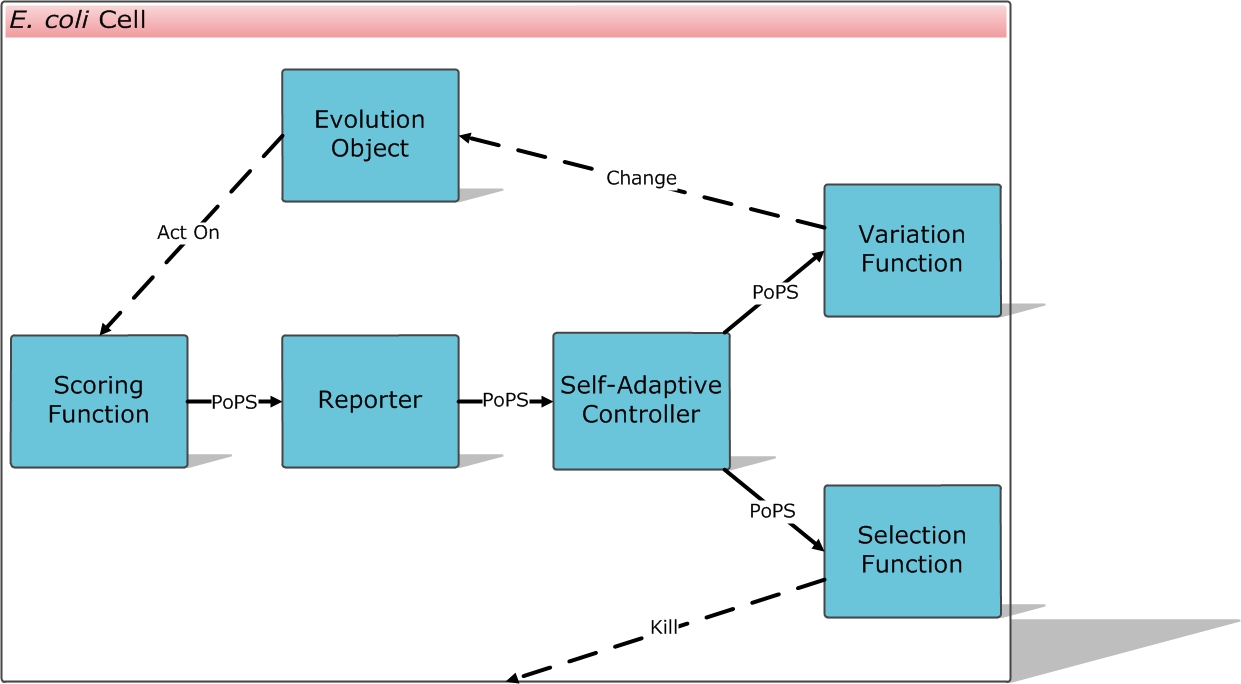
Modules & Flow Chart of the System
Scoring Function
Self-Adaptive Controller
Variation Function
Selection Function
Repoter
The Prototype
What to Do First?
Constitutive Promoter Family as Stimulus Signals
We choose to use constitutive promoters instead of the conditional operon impressions (represented by [http://en.wikipedia.org/wiki/Lac_operon IPTG-induced expression]) as the stimulus signals to test the system. The stimulus signals in a testing system are supposed to be definite and stable. However, the IPTG-induced signals are susceptible to many environmental factors. The process of inducible expression involves a series of dynamic actions in physical chemistry: the diffusion process of IPTG molecules, and the equilibruim between the attachment and disattachment of IPTG to the promoter. That way, the expression signals would fluctuate in a large scope in experiments and the mathematic analyses would be very complicate.
Comparatively, the stimulus signals based on a series of constitutive promoters of different levels are far stabler since the process are relatively direct.
- The signals produced by the constitutive promoters can maintain at the steady state during the measurement.
- Constitutive-promoter expressions can give out several different stimulus signals in one system without any disturbation among them. That is, several constitutive promoters can work independently in one system to produce double or triple stimulus signals. Instead, the IPTG-induced testing system can imput only one signal at a time corresponding to the concentration of the IPTG.
- Two or more systems with different stimulus signals can grow in the same nurture if the signals are produced by the constitutive promoters. For example, in our project, several kinds of E.coli with different imputs signals can grow in the same culture medium preparing for the screen.
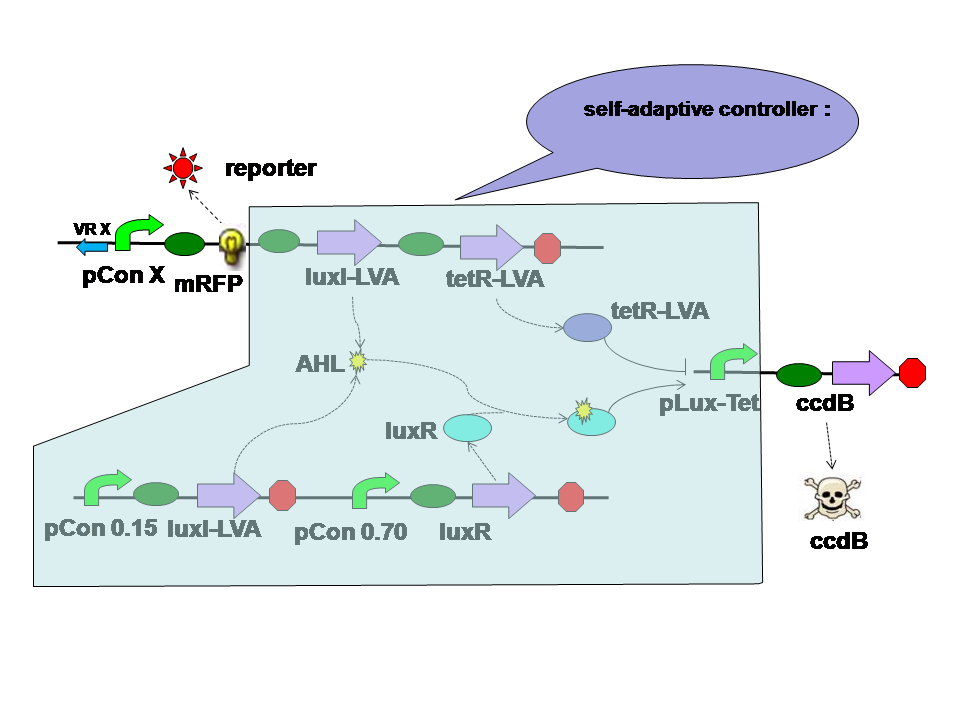
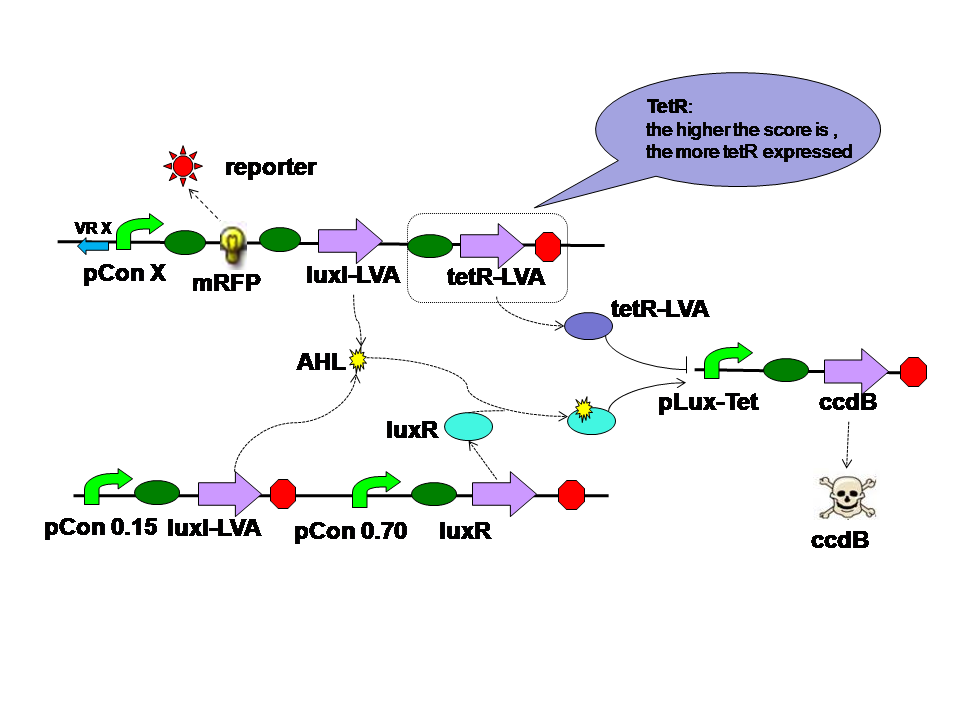
Design of the Self-Adaptive Controller
|
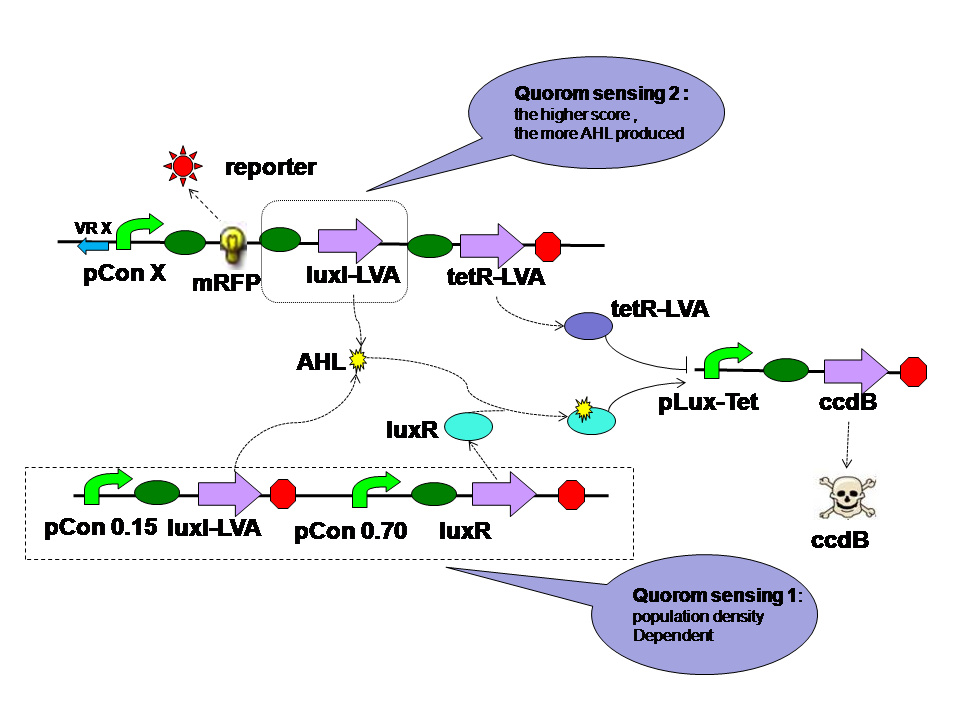
The self-adaptive controller consists of two quorom sensing parts one tetR parts and a hybrid promoter. |
|
There are two quorom sensing parts in the self adaptive controller. LuxR is expressed by a constitutive promoter and interacts with AHL produced by both LuxI in two parts. The complex then activates hybrid promoter. LuxI in different parts perform different functions. LuxI's expression in Quorom sensing 1 depends on the population density and take effect in evolution in the aspect of cell density. LuxI in part 2, on the other hand, is expressed correspondent to the evolution score directly. Thus it takes part in the evolution process directly. |
|
TetR is expressed correspondent to the evolution score, just like the LuxI from quorom sensing part 2. However tetR represses the hybrid promoter. High score means high tetR level. High tetR level means low death rate. |
Principle of Operation
- Properties of selection system
The selection system consists of hybrid promoter and ccdB. AHL-LuxR complex and tetR input perform contrast effect. We perform several experiments to identify how the hybrid promoter work. The property is as shown below:
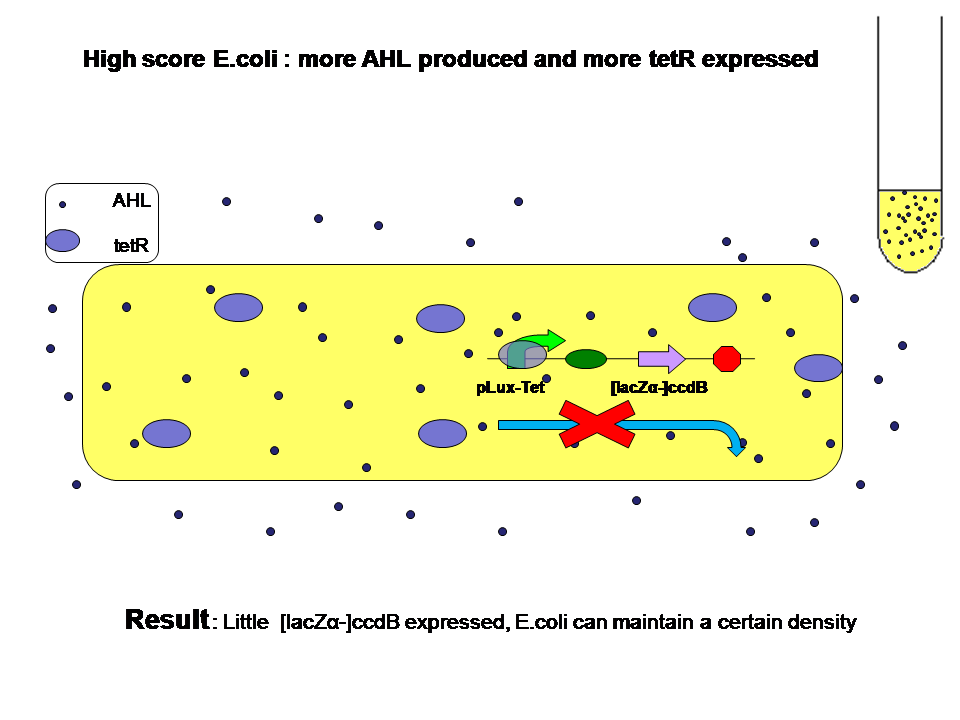
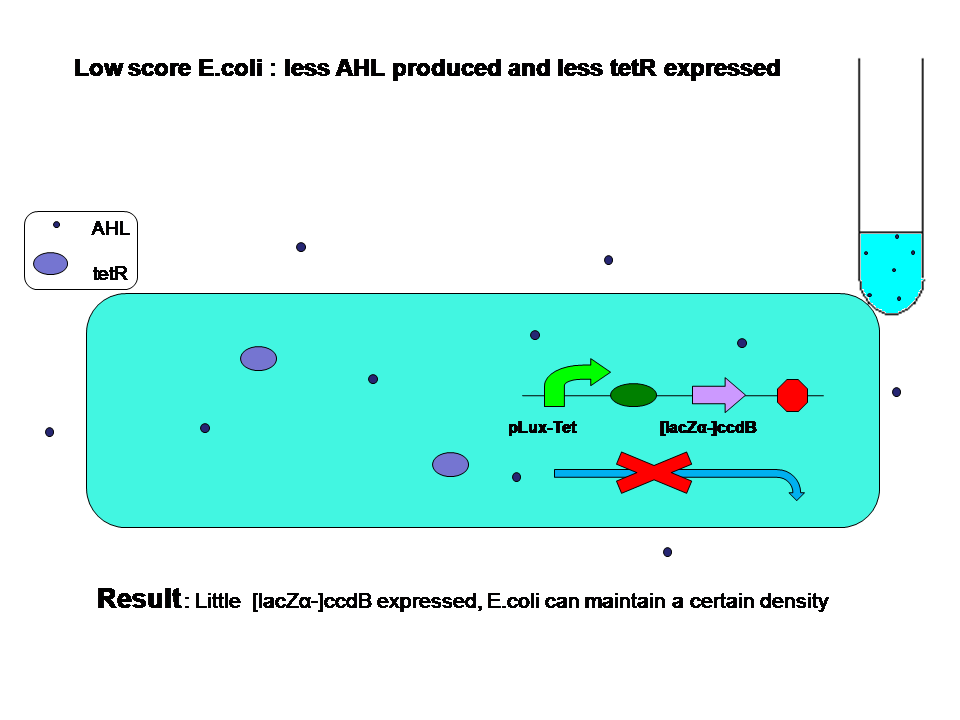
- How different "score" results in evolution
When high score and low score E.coli are grown seperately:
What if we mix them together:
Vector & Chassis
PSB1A3: high copy number
Top10
MD
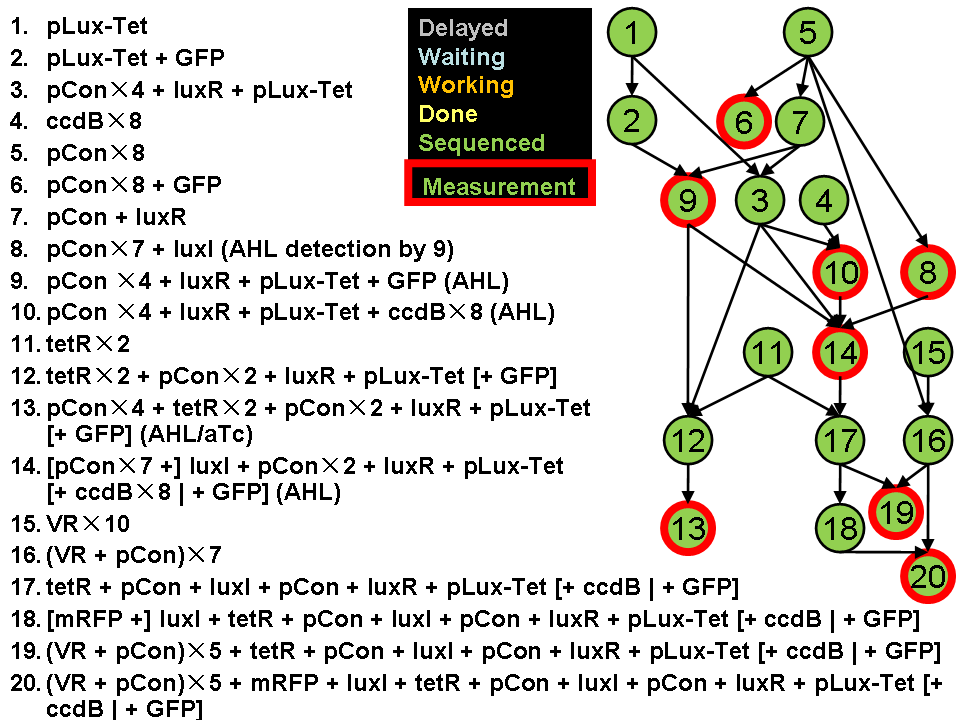
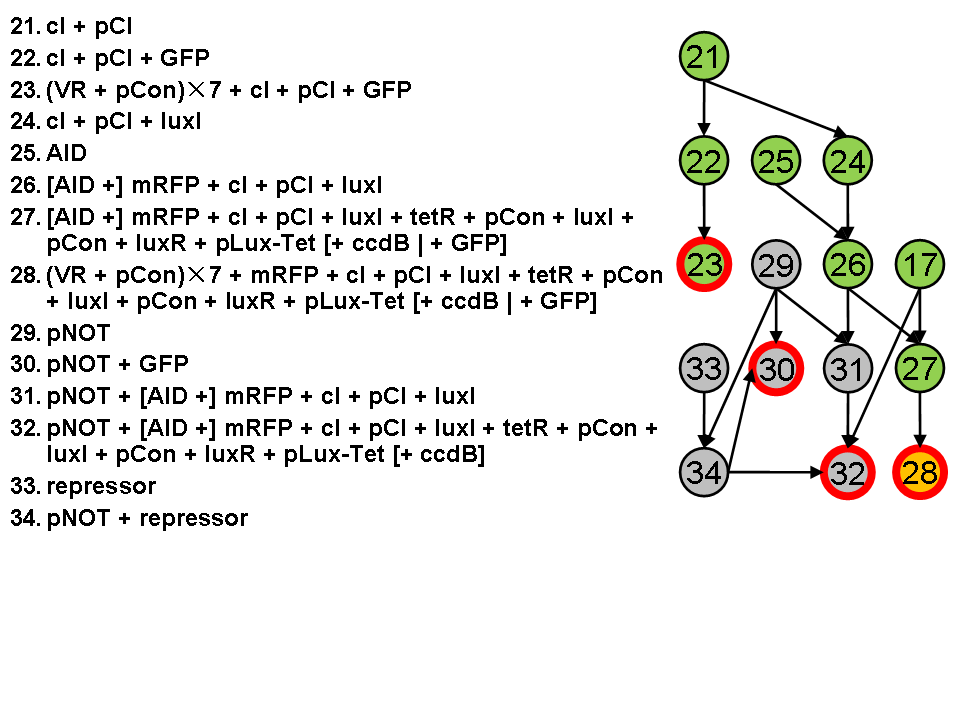
Assembly Road Map
In order to keep the whole process in perspective, we designed maps to direct our work in wet lab.
 "
"