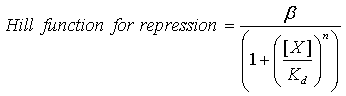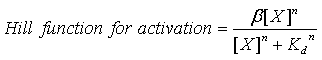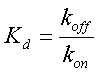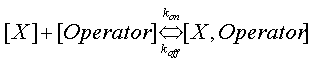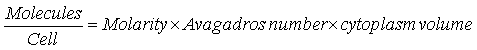Team:Aberdeen Scotland/parameters/invest 1
From 2009.igem.org
(→Discussion) |
(→Dissociation Constants) |
||
| Line 6: | Line 6: | ||
=== Introduction === | === Introduction === | ||
| - | Our model uses | + | Our model to describe the transcriptionl dynamics of the constructs uses Hill kinetics; we have three repression Hill functions of the form: |
[[Image:Dissociation_Constants_Eq_1.gif|center]] | [[Image:Dissociation_Constants_Eq_1.gif|center]] | ||
| - | + | Moreovre, the model has one activation Hill function of the following form: | |
[[Image:Dissociation_Constants_Eq_2.gif|center]] | [[Image:Dissociation_Constants_Eq_2.gif|center]] | ||
| - | + | and one repression / induction Hill function of the following form | |
[[Image:Dissociation_Constants_Eq_3.gif|center]] | [[Image:Dissociation_Constants_Eq_3.gif|center]] | ||
| - | + | where β is the maximal transcription rate, [X] is the concentration of protein X and K<sub>d</sub> is the dissociation constant for molecule X to the promoter in question. Similarly, [S] is the concentration of the inducer S and K<sub>s</sub> is the dissociation constant for the inducer to the repressor X. The dissciation cnstant K<sub>d</sub> is defined as follows: | |
[[Image:Dissociation_Constants_Eq_4.gif|center]] | [[Image:Dissociation_Constants_Eq_4.gif|center]] | ||
| - | + | where k<sub>off</sub> and k<sub>on</sub> are the on and off rates in the following reaction | |
[[Image:Dissociation_Constants_Eq_5.gif|center]] | [[Image:Dissociation_Constants_Eq_5.gif|center]] | ||
| - | K<sub>d</sub> has | + | The dissociation constant K<sub>d</sub> has more biologically meaningful definition; it is the concentration of X at which the promoter will be free 50% of the time. |
{{:Team:Aberdeen_Scotland/break}} | {{:Team:Aberdeen_Scotland/break}} | ||
| Line 32: | Line 32: | ||
=== Discontinuity === | === Discontinuity === | ||
| - | The units of K<sub>d</sub> are usually given in M, the molarity, or moles-per-litre. Our model works with the exact number of molecules so we | + | The units of K<sub>d</sub> are usually given in M, the molarity, or moles-per-litre. Our model works with the exact number of molecules, so that we convert the K<sub>d</sub> values into molecules-per-cell. This is achieved as follows: |
[[Image:Dissociation_Constants_Eq_6.gif|center]] | [[Image:Dissociation_Constants_Eq_6.gif|center]] | ||
| Line 40: | Line 40: | ||
This conversion constant of Avogadro’s number multiplied by the cytoplasm volume is ~ 402000000 (402 million). | This conversion constant of Avogadro’s number multiplied by the cytoplasm volume is ~ 402000000 (402 million). | ||
| - | + | Here, we face a major problem, since most dissociation constants found in the literature equate to a value of molecules per cell that is less than 1. Clearly, in a cell with 10 plasmids and therefore 10 operators, 1 molecule could not repress all of them! | |
| - | Below is a table of conflicting information we found. This is an extract from the ETHZ Wiki [6] with the | + | Below is a table of conflicting information we found. This is an extract from the ETHZ Wiki [6] , where a new colum with the values of K<sub>d</sub> in units of molecule-per-cell has been added. |
| Line 95: | Line 95: | ||
| - | And here are | + | And here are other parameters that we found in the literature: |
Revision as of 12:48, 17 August 2009
University of Aberdeen - Pico Plumber
Contents |
Dissociation Constants
Introduction
Our model to describe the transcriptionl dynamics of the constructs uses Hill kinetics; we have three repression Hill functions of the form:
Moreovre, the model has one activation Hill function of the following form:
and one repression / induction Hill function of the following form
where β is the maximal transcription rate, [X] is the concentration of protein X and Kd is the dissociation constant for molecule X to the promoter in question. Similarly, [S] is the concentration of the inducer S and Ks is the dissociation constant for the inducer to the repressor X. The dissciation cnstant Kd is defined as follows:
where koff and kon are the on and off rates in the following reaction
The dissociation constant Kd has more biologically meaningful definition; it is the concentration of X at which the promoter will be free 50% of the time.
Discontinuity
The units of Kd are usually given in M, the molarity, or moles-per-litre. Our model works with the exact number of molecules, so that we convert the Kd values into molecules-per-cell. This is achieved as follows:
Where the volume of the cytoplasm of the cell is 6.7×10-16 litres
This conversion constant of Avogadro’s number multiplied by the cytoplasm volume is ~ 402000000 (402 million).
Here, we face a major problem, since most dissociation constants found in the literature equate to a value of molecules per cell that is less than 1. Clearly, in a cell with 10 plasmids and therefore 10 operators, 1 molecule could not repress all of them!
Below is a table of conflicting information we found. This is an extract from the ETHZ Wiki [6] , where a new colum with the values of Kd in units of molecule-per-cell has been added.
| Parameter | Value | Molecules per cell | Description |
| KLacI | 0.1 - 1 [pM] OR 800 [nM] | 0.00004-0.0004 molecules OR 322 molecules | LacI repressor dissociation constant |
| KIPTG | 1.3 [µM] | 522 molecules | IPTG-LacI repressor dissociation constant |
| KtetR | 179 [pM] | 0.07 molecules | TetR repressor dissociation constant |
| KcI | 8 [pM] OR 50 [nM] | 0.003 molecules OR 20 molecules | cI repressor dissociation constant |
| KHSL | 0.09 - 1 [µM] | 402 molecules | HSL-LuxR activator dissociation constant |
And here are other parameters that we found in the literature:
| Parameter | Value | Molecules per cell | Description | Reference |
| KLacI | ~1*10 -12 M OR ~1.8*10-12 M | 0.0004 molecules OR 0.00072 molecules | Dissociation constant for LacI to LacO DNA site | [1][2] |
| KIPTG | 1*10-6 M | 402 molecules | Dissociation constant for IPTG to LacI | [3] |
| KtetR | (5.6 ± 2) × 10-9 M OR 1.53*10-8 M | 2.25 molecules OR 6.1506 molecules | Dissociation constant for TetR to TetO | [4][5] |
| KcI | 50 * 10-9 M | 20 molecules | Dissocitation constant for cI to DNA site | [6] |
Discussion
Upon further investigation we have concluded that the majority of Kd values found in papers were unrealistically low for the following reasons:
1. Most Kd values are measured in-vitro, which yields a low measurement since the conditions of the reaction - most notably the salt concentration and pressure - are completely different than in an E.coli cell. The salt concentration affects the reaction significantly since it lowers the electrostatic affinity of the protein to the operon. We know from Thermodynamics that pressure and temperature will change reaction kinetics and hence the in vitro experiments will have different reaction rates and hence different Kd values than would be found in the cell.
2. We have found measurements of Kd values which have been done in conditions which try to replicate in vivo conditions. These Kd values are better, but also infeasibly low, since they do not take into account non-specific DNA binding and cell pressure.
3. In our model, we describe concentrations in terms of molecules-per-cell, instead of moles-per-litre. Upon converting the Kd values from moles-per-lire to molecules-per-cell we found that a great deal the Kd values we found were less than 1 molecule per cell. This implies that less than one protein (LacI, TetR etc.) is required to half the overall production. This is physically irrepresentative for a number of reasons, including the fact that the probability that one protein molecule will collide with a single operon in the cell at the correct angle is close to zero.
We consulted with Prof. Peter McGlynn of the Institute of Medical Sciences in Aberdeen, who agreed with our analysis that the Kd values were infeasibly low and introduced the idea of non-specific DNA binding to us. He showed us a PhD thesis from one of his students, Dr. Bryony Payne, from 2006. In this thesis a far more direct and accurate measurement of the number of LacI tetermers present in the cell was made. It stated that on average 340 tetramers of LacI were present in an e. coli cell. Since under normal conditions (when glucose is avalible and no lactose is present) the LacO operon is repressed. We can assume that 340 tetermers will fully repress the lacO operon. From this value, we estimated the Kd value for the LacO operon to be 700 molecules-per-cell.
Our new estimations for Kd
Starting our estimation from Dr. Payne's 2006 paper; we are given that 340 LacI tetramers completely repress a promoter. Hence roughly 120 tetramers will give half repression. Assuming the tetramers are stable this gives a value of KLacI - Kd for LacI to LacO - of 4×170 or KLacI ~ 700.
For the LacI-IPTG complex formation, we estimated KIPTG ~1200 using [7] and our value of KLacI above. TetR to TetO have a lower affinity to each other than LacI to LacO. However, the in-vitro values suggest that TetR still binds with a strong affinity to TetR. Thus the KTetR value was roughly estimated to be up to 10 times KLacI. The in-vitro values for cI to its operon seem to suggest that the in-vivo KcI value is of the same order of magnitude, but possibily smaller, than KTetR.
Now we have:
KLacI = 700 molecules per cell
KcI = 7000 molecules per cell
KTetR = 7000 molecules per cell
References
[1] Mitchel Lewis (2005) The Lac repressor. C. R. Biologies 328 (2005) 521–548
[2] Falcon C.M and Matthews K.S. (2000) Operator DNA sequence Variation Enhances High Affinity Binding by Hinge Helix Mutants of Lactose Repressor Protein. Biochemistry. 39, 11074-11084
[3] Uri Alon, An introduction to systems Biology, p244
[4] Nucleic Acids Res. 2004; 32(2): 842–847. Two mutations in the tetracycline repressor change the inducer anhydrotetracycline to a corepressor Annette Kamionka, Joanna Bogdanska-Urbaniak, Oliver Scholz, and Wolfgang Hillen*
[5] Volume 272, Number 11, Issue of March 14, 1997 pp. 6936-6942, The Role of the Variable Region in Tet Repressor for Inducibility by Tetracycline, Christian Berens , Dirk Schnappinger and Wolfgang Hillen
[6] http://parts.mit.edu/igem07/index.php?title=ETHZ/Parameters
[7] Detailed map of a cis-regulatory input function – Y. Setty*,†, A. E. Mayo*,†, M. G. Surette‡, and U. Alon*,†,§
 Back Back
|
Continue 
|
 "
"