Team:KULeuven/Modeling/Key Antikey
From 2009.igem.org
(→References) |
(→Mathematical Model) |
||
| Line 35: | Line 35: | ||
| 0.00848 s<sup>-1</sup> | | 0.00848 s<sup>-1</sup> | ||
| estimate | | estimate | ||
| - | | [https://2009.igem.org/Team:KULeuven/Modelling/Vanillin_Production#References [ | + | | [https://2009.igem.org/Team:KULeuven/Modelling/Vanillin_Production#References [2<html>]</html>] |
|- | |- | ||
| k<sub>translation</sub> | | k<sub>translation</sub> | ||
| 0.167 s<sup>-1</sup> | | 0.167 s<sup>-1</sup> | ||
| estimate | | estimate | ||
| - | | [https://2009.igem.org/Team:KULeuven/Modelling/Vanillin_Production#References [ | + | | [https://2009.igem.org/Team:KULeuven/Modelling/Vanillin_Production#References [2<html>]</html>] |
|- | |- | ||
! colspan="4" style="border-bottom: 1px solid #003E81;" | Key Lock Parameters | ! colspan="4" style="border-bottom: 1px solid #003E81;" | Key Lock Parameters | ||
| Line 47: | Line 47: | ||
| 0.00237 s<sup>-1</sup> | | 0.00237 s<sup>-1</sup> | ||
| Rate of unlocking the RIBOLOCK through key | | Rate of unlocking the RIBOLOCK through key | ||
| - | | [https://2009.igem.org/Team:KULeuven/Modelling/Key_Antikey#References [ | + | | [https://2009.igem.org/Team:KULeuven/Modelling/Key_Antikey#References [3<html>]</html>] |
|- | |- | ||
| k<sub>lock</sub> | | k<sub>lock</sub> | ||
| 0.00416 s<sup>-1</sup> | | 0.00416 s<sup>-1</sup> | ||
| Rate of locking of unlocked RIBOLOCK-Key complex. | | Rate of locking of unlocked RIBOLOCK-Key complex. | ||
| - | | [https://2009.igem.org/Team:KULeuven/Modelling/Key_Antikey#References [ | + | | [https://2009.igem.org/Team:KULeuven/Modelling/Key_Antikey#References [3<html>]</html>] |
|- | |- | ||
| k<sub>open</sub> | | k<sub>open</sub> | ||
| 7.5 s<sup>-1</sup> | | 7.5 s<sup>-1</sup> | ||
| Rate of unlocking RIBOLOCK when no key is present (LEAK). | | Rate of unlocking RIBOLOCK when no key is present (LEAK). | ||
| - | | [https://2009.igem.org/Team:KULeuven/Modelling/Key_Antikey#References [ | + | | [https://2009.igem.org/Team:KULeuven/Modelling/Key_Antikey#References [3<html>]</html>] |
|- | |- | ||
| k<sub>close</sub> | | k<sub>close</sub> | ||
| 500 s<sup>-1</sup> | | 500 s<sup>-1</sup> | ||
| Rate of locking of leaked RIBOLOCK. | | Rate of locking of leaked RIBOLOCK. | ||
| - | | [https://2009.igem.org/Team:KULeuven/Modelling/Key_Antikey#References [ | + | | [https://2009.igem.org/Team:KULeuven/Modelling/Key_Antikey#References [3<html>]</html>] |
|- | |- | ||
! colspan="4" style="border-bottom: 1px solid #003E81;" | Annealing | ! colspan="4" style="border-bottom: 1px solid #003E81;" | Annealing | ||
| Line 69: | Line 69: | ||
| infinity | | infinity | ||
| Rate annealing is so high we consider it to be infinite. | | Rate annealing is so high we consider it to be infinite. | ||
| - | | [https://2009.igem.org/Team:KULeuven/Modelling/Key_Antikey#References [ | + | | [https://2009.igem.org/Team:KULeuven/Modelling/Key_Antikey#References [4<html>]</html>] |
|} | |} | ||
</center> | </center> | ||
Revision as of 13:21, 9 September 2009
Contents |
Key Lock Antikey
Overview
The Key and Antikey system performs a subtraction of the blue light signal and the vanillin receptor signal. The result controls the vanillin production. The biology behind the subtraction involves the annealing of complementary RNA strands, the Key and the Antikey. This reaction is favoured over the reaction between the Key and the Lock leading to vanillin synthesis. In this way we try to perform the subtraction before inducing production of vanillin.
This biological equivalent of a subtraction can only yield a positive number, so one can only subtract a small from a big amount. Because we can only actively produce vanillin, we have to subtract the measured quantity of vanillin, the amount of anti-key produced by the vanillin receptor from the wanted quantity of vanillin, the amount of key produced by the blue light sensor.
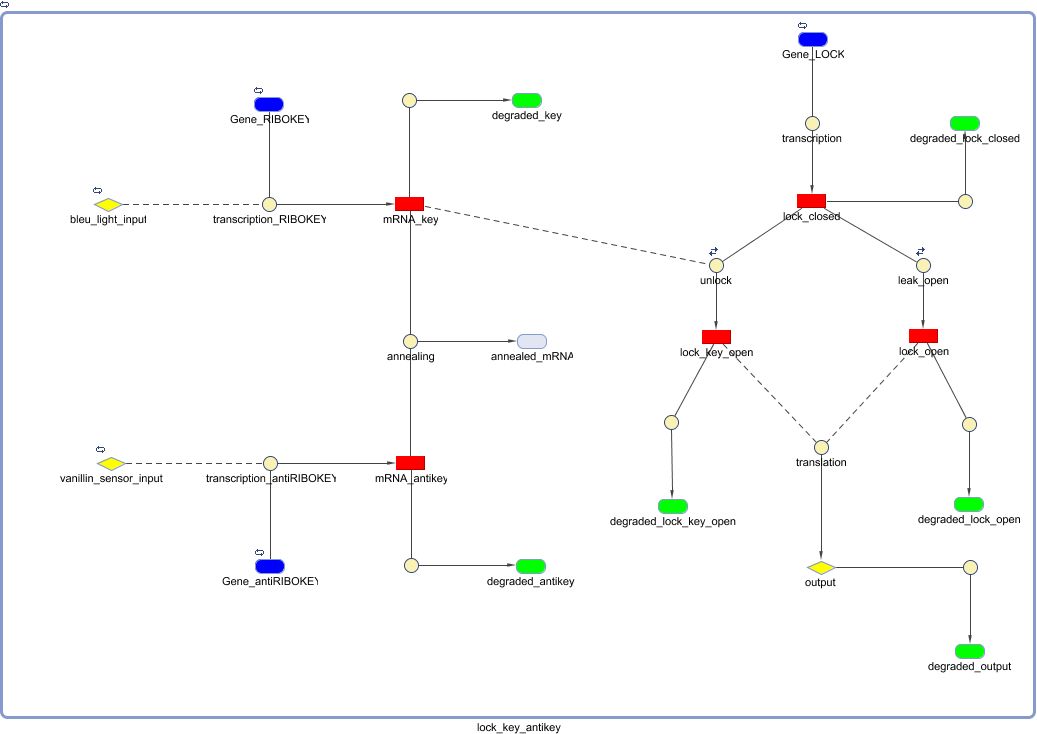
Biological Model
Mathematical Model
| Name | Value | Comments | Reference |
|---|---|---|---|
| Degradation Rates | |||
| dmRNA | 2.3105E-3 s-1 | [1] | |
| Transcription Rates | |||
| ktranscription | 0.00848 s-1 | estimate | [2] |
| ktranslation | 0.167 s-1 | estimate | [2] |
| Key Lock Parameters | |||
| kunlock | 0.00237 s-1 | Rate of unlocking the RIBOLOCK through key | [3] |
| klock | 0.00416 s-1 | Rate of locking of unlocked RIBOLOCK-Key complex. | [3] |
| kopen | 7.5 s-1 | Rate of unlocking RIBOLOCK when no key is present (LEAK). | [3] |
| kclose | 500 s-1 | Rate of locking of leaked RIBOLOCK. | [3] |
| Annealing | |||
| Kannealing | infinity | Rate annealing is so high we consider it to be infinite. | [4] |
Simulation
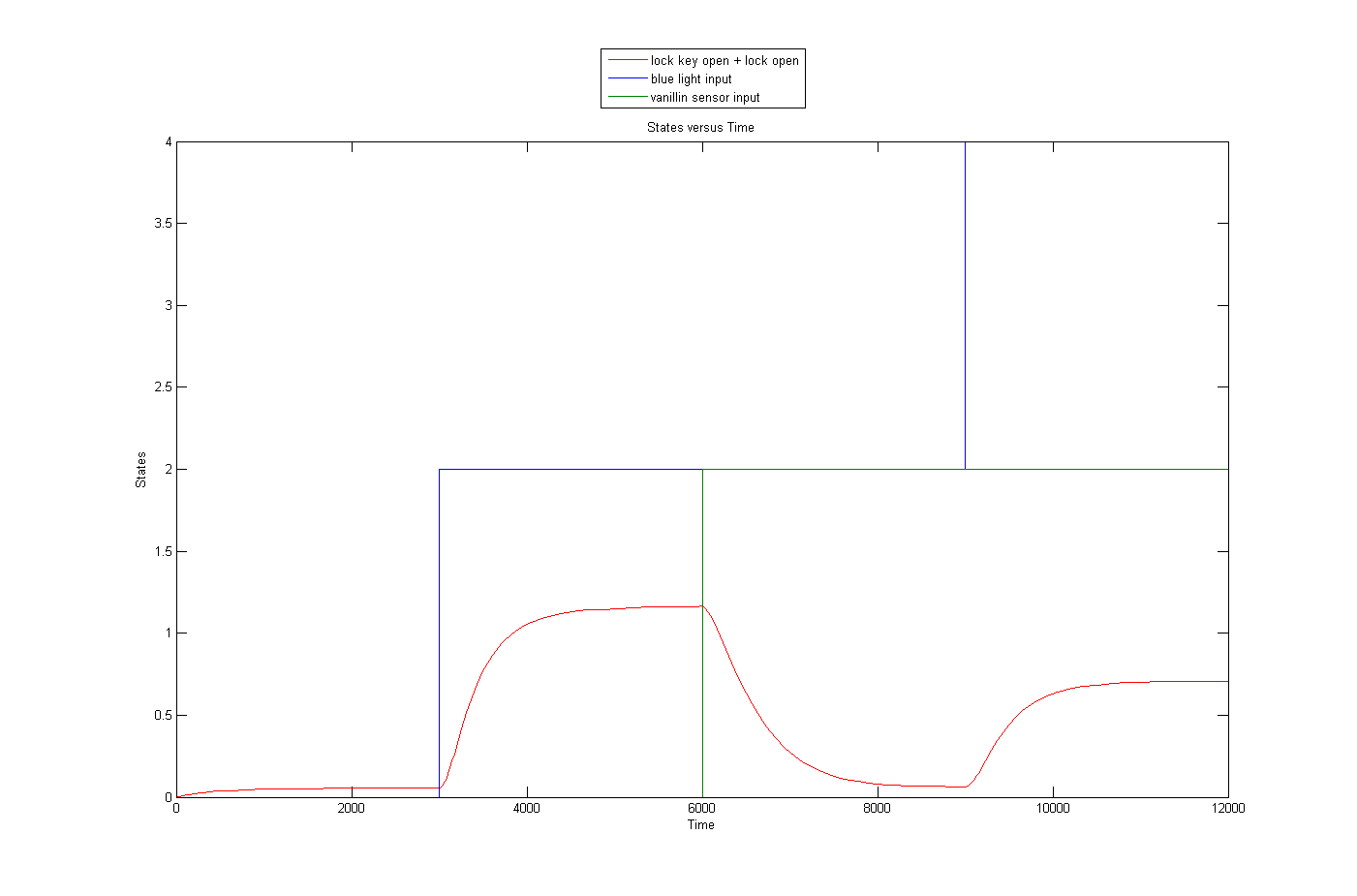
The input/output behaviour of the key/lock system was investigated using simulations. In the following figures, time is measured in seconds and quantities in molecules.
In the figure below, the blue light input is represented in blue, the vanillin input in green and the amount of unlocked key in red.
Because of the nonlinear relationship between input and (anti)key (Michaelis-Menten), the steady state level of key depends on the level of both blue light sensor and vanillin sensor input and not just the difference between the two.
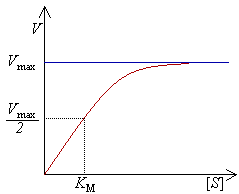
The most important conclusion is that for good functioning of the differentiator the transcription of both the key and the antikey are operated in the linear region (See figure Michaelis-Menten). It's therefore important that the inputs of this system remain in the same magnitude of the Km of the production of the (anti)key.
Also, the half life of the key and antikey have to be as short as possible, otherwise we subtract the integrated amounts of the signals and not the signals themselves. This should not pose any problem since the half life of non translational mRNA strands is merely 5 minutes, substantially faster than the system requires. This reasoning remains valid for all species in the cell, where the speed of the cell is limited by the slowest degrading species.
References
[1] J.A. Bernstein et al., “Global analysis of mRNA decay and abundance in Escherichia coli at single-gene resolution using two-color fluorescent DNA microarrays,” Proceedings of the National Academy of Sciences of the United States of America, vol. 99, Jul. 2002, pp. 9697–9702
[2] S.L. Gotta et al., “rRNA Transcription Rate in Escherichia Coli,” Journal of Bacteriology, vol. 173, Oct. 1991, pp. 6647-6649
[3] https://2008.igem.org/Team:KULeuven/Model/Filter
 "
"