Team:Imperial College London/M3
From 2009.igem.org
(→40pxModule 3 Overview) |
(→40pxModule 3 Overview) |
||
| Line 19: | Line 19: | ||
<br> | <br> | ||
| + | {{Imperial/09/Division}} | ||
==Restriction Enzymes== | ==Restriction Enzymes== | ||
Revision as of 11:21, 18 September 2009

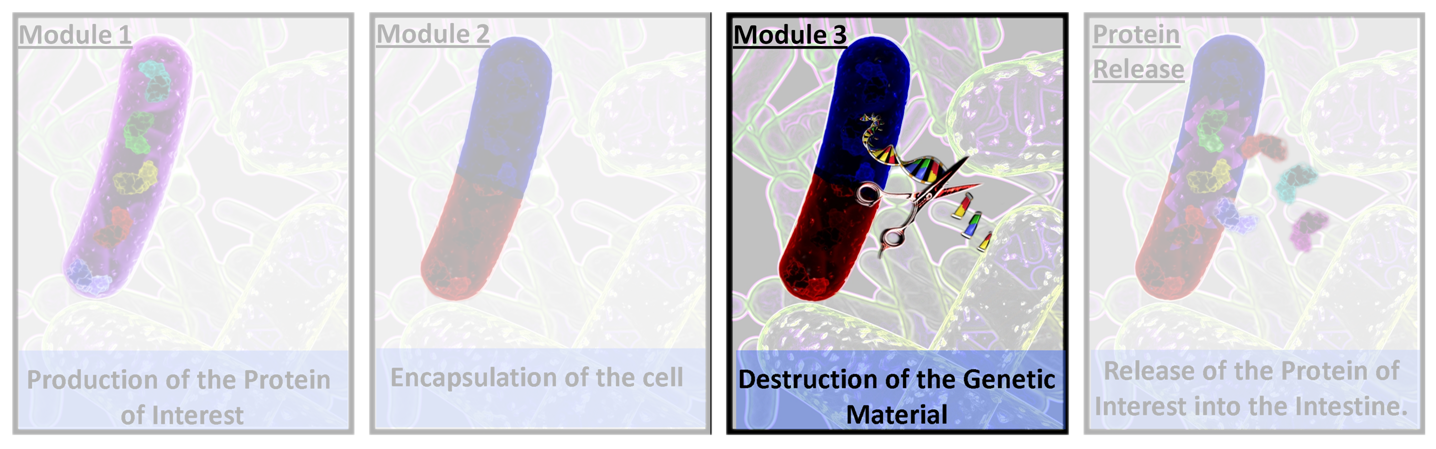
 Module 3 Overview
Module 3 Overview
Module 3 is the final module of the system. It programs the E.ncapsulator to produce restriction enzymes and destroy all its genetic material after encapsulation has finished. This prevents any possible pathogenic effects, and also allays health concerns of eating genetically modified organisms.
To learn more about the ethical implications of live organisms, please click on The E.ncapsulator 


Restriction Enzymes
After Module 2 has been completed, genome deletion is triggered by raising the temperature.
see thermoinduction under temporal control)
The cell then produces the restriction enzymes DpnII and TaqI. These specifically target and cut short 4 base DNA sequences. As the cutting is frequent, the genetic material contained within the cell will all be destroyed.
A distinct advantage of using restriction enzymes for our 'killing' mechanism is that the cell membrane is left intact afterwards, and the protein of interest will still be protected by the encapsulated cell. This renders the bacterium no more than an inanimate shell containing our protein drug of choice.
Dam methylation
To protect against DNA destruction due to basal levels of restriction enzyme production, we have made use of the native E. coli Dam methylase protection system. This methylates DNA. Therefore, only high levels of restriction enzyme (ie. after thermal triggering) will cleave the DNA.
 "
"