|
|
| Line 1: |
Line 1: |
| | {{:Team:USTC/Template/Header}} | | {{:Team:USTC/Template/Header}} |
| | + | {|valign = "top" align = "center" border = "0" width = "800px" |
| | + | |
| | = Problem & Solution = | | = Problem & Solution = |
| | [[Image:barcode.jpg|thumb|right|200px|Barcode & its Scanner]] | | [[Image:barcode.jpg|thumb|right|200px|Barcode & its Scanner]] |
| Line 53: |
Line 55: |
| | | | |
| | = Results = | | = Results = |
| - | {|align="justify"
| |
| - | |[[Image:tool_result.png|500px|left|thumb|cross experiment result]]
| |
| | | | |
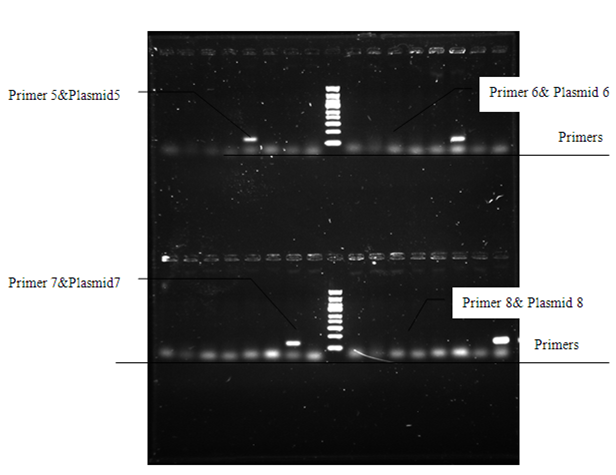
| - | |<p>In the electrophoretogram, only the combination of primer 5 & plasmid 5, primer 6 & plasmid 6, primer 7 & plasmid 7 lead to PCR results. Along with other electrophoretograms we conclude that this set of primers can be used as a set of barcodes.</p> | + | |- |
| | + | |align = "center" width = "400px"| |
| | + | [[Image:tool_result.png|350x350px|thumb|cross experiment result]] |
| | + | |align = "justify" cellspacing = "10"| |
| | + | <p> In the electrophoretogram, only the combination of primer 5 & plasmid 5, primer 6 & plasmid 6, primer 7 & plasmid 7 lead to PCR results. Along with other electrophoretograms we conclude that this set of primers can be used as a set of barcodes.</p> |
| | | | |
| | | | |
| Line 62: |
Line 66: |
| | | | | | |
| | | | |
| - | |} | + | |
| | + | <!--[[Image:tool_result.png|500px|left|thumb|cross experiment result]] |
| | + | --> |
| | | | |
| | = Application = | | = Application = |
Problem & Solution
How to distinguish the different biobricks in a biobrick pool in experiment?
To take a DNA sequencing?---it's costly and time-consuming.
We are faced with such a problem to determine the final outputs of E.ADEM (E.coli Automatic Directed Evolution Machine)[1]. An inspiration comes from the supermarket checkout system where thousands of commodities are tagged by barcodes. This universal commercial ID both shortens the check-out time and adds the accuracy. By comparing our problem with this system, what we need to do can be concluded as to design barcodes for biobricks, BioBrick-Barcode, as we call it.
Where is the scanner?
PCR (Polymerase Chain Reaction)[http://en.wikipedia.org/wiki/PCR], which is so conveniently conducted, can serve as the scanner while the primers it needs are the analogue of barcode sequences.
Solution:
To check the final outputs in the system of E.ADEM (E.coli Automatic Directed Evolution Machine), we need to design a set of DNA oligo-sequences primers as the barcodes to go through the scanner of PCR and help determining the final evolutionary products.
Design
- A set of barcodes under a certain kind of barcode reader has to satisfy its conditions in order to be picked out respectively. For BioBrick-Barcodes, the oligo-sequences must meet several conditions as we considered:
1. Since the BioBrick-Barcodes function as PCR primers, they must satisfy the basic conditions as high-quality primers:
a. Each primer should be 20-30 nucleotides in length;
b. Contain approximately equal numbers of 4 bases, with a balanced distribution of G&C residues;
c. Hold a low propensity to form stable secondary structures;
d. Forward and reverse primers can work properly together;
2. As a set of barcodes, each of the primers should be perceivably different in order to be identified by the scanner:
a. Any one of the primer cannot lead to the PCR of other DNA sequences(a plasmid with a certain target sequence.)
b. All of the primers should be specific for the target sequence without combining with other sites.
Primer3[http://primer3.sourceforge.net/releases.php] is an open-source primer-design software, and we modified its C codes in parts to design the BioBrick-Barcodes. Since Primer3 is mainly used to pick primers from a DNA template while the BioBrick-Barcodes are supposed to be randomly produced, and a set of BioBrick-Barcodes should be diverse to avoid overlapping PCR, we changed three functions and structure of Primers.
1. Add a function to randomly produce the DNA oligo-sequences which replaced the previous function used to pick
segments from the template.
2. Modified the structure used to store information of each primer.
3. Use the Libray_Mispriming to choose diverse primers.
We design a set of ten diverse primers by using the modified software. Apparently, we need to conduct experiments to test its feasibility.
1.Synthesize the DNA oligo-sequence;
2.Construct plasmids;
3.Conduct a 10*10 cross experiment: Each primer reacts with all the plasmids;
Expectation: only complementary couple leads to PCR results.
Results
|
|
In the electrophoretogram, only the combination of primer 5 & plasmid 5, primer 6 & plasmid 6, primer 7 & plasmid 7 lead to PCR results. Along with other electrophoretograms we conclude that this set of primers can be used as a set of barcodes.
|
|
Application
BioBrick barcodes are a set of artifical DNA parts designed for PCR. They can be used alone as a high-quality PCR primer. Also, they can be used in a group as a convenient tool to identify the biobrick parts just as we do.
We create a new software to design PCR primers without a template. The primers chosen from a random oligo-sequences can enlarge the application scope of PCR.
|
 "
"