Team:Virginia Commonwealth/Design
From 2009.igem.org
GMcArthurIV (Talk | contribs) (→Promoters classes) |
GMcArthurIV (Talk | contribs) |
||
| Line 4: | Line 4: | ||
| | | | ||
| | | | ||
| - | == | + | ==UP-element design== |
| - | === | + | Work has been done to identify a consensus UP element on some promoters which interacts with an alpha subunit on RNAP (Estrem ST, Gaal T, Ross W, Gourse RL, Identification of an UP element consensus sequence for bacterial promoters, ''PNAS'', (1998), 95, 9761-9766.). This UP element has been shown to significantly increase the RNA polymerase-recruiting power of promoters and is, therefore, a transcriptional enhancer. |
| + | |||
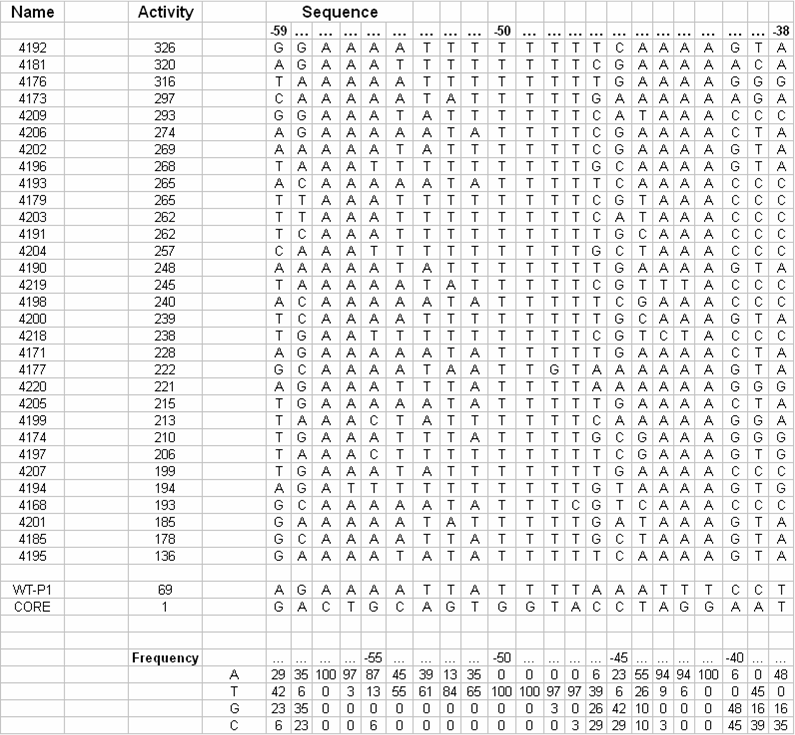
| + | [[Image:VCU 2009 tabulated UP-elements.png|thumb|center|400px|Tabulation of UP-elements in ''E. coli'' and nucleotide frequency at each position (adapted from Estrem ST, Gaal T, Ross W, Gourse RL, Identification of an UP element consensus sequence for bacterial promoters, ''PNAS'', (1998), 95, 9761-9766.)]] | ||
| + | |||
| + | We are working on creating this UP element as a modular BioBrick part and characterizing its activity using the promoter characterization method that we develop. We will also manipulate this element using a bottom-up design approach to get UP elements of varying degree's of strength. Thus increasing the level of control we have over gene expression. | ||
| + | ===Design strategies=== | ||
| + | Promoter design will be approached from the bottom up. First, a consensus promoter sequence will be studied and accepted by researching the work done by others on various promoters. This consensus will be our starting point as we identify which nucleotide and nucleotide sequences are most important to RNAP binding affinity. We will use direct synthesis to construct promoter sequences that we will test. | ||
| + | ==Future work== | ||
*Constitutive promoters | *Constitutive promoters | ||
**There are several types of constitutive promoters that we can focus on. These promoters are separate in regard to their respective sigma binding domains. Constitutive promoters that interact with the sigma 70 domain of RNAP are most common most prevalent being as how sigma 70 RNAP is most common at normal cellular growth conditions. The alternatives would be sigma 54 and sigma 32 binding factors, these types of polymerases are more active when the cell is under certain strain, such as heat shock of nitrogen starvation. | **There are several types of constitutive promoters that we can focus on. These promoters are separate in regard to their respective sigma binding domains. Constitutive promoters that interact with the sigma 70 domain of RNAP are most common most prevalent being as how sigma 70 RNAP is most common at normal cellular growth conditions. The alternatives would be sigma 54 and sigma 32 binding factors, these types of polymerases are more active when the cell is under certain strain, such as heat shock of nitrogen starvation. | ||
**There has already been considerable work done with sigma 70 RNAP including an established promoter library of varying promoter strengths available on the parts registry. We will develop and implement a method to fully characterize and standardize these promoters' activity and upload it to the registry. | **There has already been considerable work done with sigma 70 RNAP including an established promoter library of varying promoter strengths available on the parts registry. We will develop and implement a method to fully characterize and standardize these promoters' activity and upload it to the registry. | ||
| - | <br /> | + | <br /> |
| - | + | ||
| - | + | ||
| - | + | ||
*T7 Promoters | *T7 Promoters | ||
**T7 phage promoters have long been utilized as a way to create orthogonal gene expression. There is very little variation in the strength of T7 promoters available via the BioBrick registry. Our goal is to identify a consensus T7 promoter sequence and use our bottom-up design approach to create a library of T7 promoters which we will characterize and upload to the registry. | **T7 phage promoters have long been utilized as a way to create orthogonal gene expression. There is very little variation in the strength of T7 promoters available via the BioBrick registry. Our goal is to identify a consensus T7 promoter sequence and use our bottom-up design approach to create a library of T7 promoters which we will characterize and upload to the registry. | ||
| - | |||
| - | |||
| - | |||
|- | |- | ||
| | | | ||
Revision as of 15:59, 21 October 2009
UP-element designWork has been done to identify a consensus UP element on some promoters which interacts with an alpha subunit on RNAP (Estrem ST, Gaal T, Ross W, Gourse RL, Identification of an UP element consensus sequence for bacterial promoters, PNAS, (1998), 95, 9761-9766.). This UP element has been shown to significantly increase the RNA polymerase-recruiting power of promoters and is, therefore, a transcriptional enhancer. We are working on creating this UP element as a modular BioBrick part and characterizing its activity using the promoter characterization method that we develop. We will also manipulate this element using a bottom-up design approach to get UP elements of varying degree's of strength. Thus increasing the level of control we have over gene expression. Design strategiesPromoter design will be approached from the bottom up. First, a consensus promoter sequence will be studied and accepted by researching the work done by others on various promoters. This consensus will be our starting point as we identify which nucleotide and nucleotide sequences are most important to RNAP binding affinity. We will use direct synthesis to construct promoter sequences that we will test. Future work
| |
 "
"