Team:Imperial College London/Drylab/Protein Production
From 2009.igem.org
(→Summary of simulation results) |
(→Summary of simulation results) |
||
| Line 36: | Line 36: | ||
*The more IPTG we add in, the higher the amount of output protein. | *The more IPTG we add in, the higher the amount of output protein. | ||
[[Image:II09_SIm_main_prot.jpg]]<br><br> | [[Image:II09_SIm_main_prot.jpg]]<br><br> | ||
| - | <html><a href="https://2009.igem.org/Team:Imperial_College_London/Drylab | + | <html><a href="https://2009.igem.org/Team:Imperial_College_London/Drylab/Protein_Production/Simulations"><img style="vertical-align:bottom;" width=90px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Learnmore.png"></a></html> about the simulations! <br><br><br> |
*The effects of IPTG toxicity were investigated and we found that for these concentration ranges, IPTG is not toxic to cells. Click on the link to see the analyzed results:[[Media:II09_IPTG_growth.ogg | IPTG growth curves]] | *The effects of IPTG toxicity were investigated and we found that for these concentration ranges, IPTG is not toxic to cells. Click on the link to see the analyzed results:[[Media:II09_IPTG_growth.ogg | IPTG growth curves]] | ||
*The constants in this model are arbitrary. We justify our usage of these values with a more detailed dynamical analysis of the system, which shows that it can only have fixed points[1]. [[Media:II09_Prot_stability analysis.ogg | System stability analysis (read about the equations before opening this file, it's intimidating!)]] | *The constants in this model are arbitrary. We justify our usage of these values with a more detailed dynamical analysis of the system, which shows that it can only have fixed points[1]. [[Media:II09_Prot_stability analysis.ogg | System stability analysis (read about the equations before opening this file, it's intimidating!)]] | ||
Revision as of 18:59, 7 October 2009

- Overview
- The model
- Simulations
Protein Production
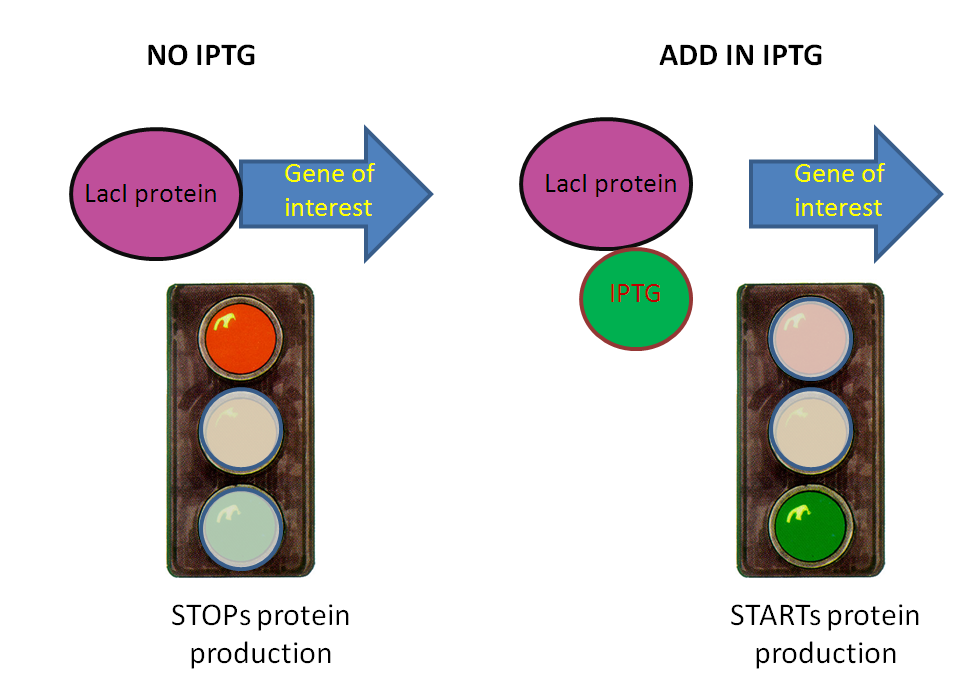
Based on the Genetic circuit, a LacI-IPTG inducible promoter is responsible for kickstarting the production of the drug.
- In the absence of IPTG, LacI represses the production of the drug (Cellulase or PAH)
- When IPTG is introduced, the LacI repressing pathway is “de-repressed”, and some output protein is produced.
Contents |
Our goals
The modelling aims to provide an overview and better understanding of the M1 system’s function by:
- Characterizing the system.
- Modeling to account for several factors that may reduce/hinder the production of the protein drug such as:
- Lac promoter leakiness
- IPTG toxicity
- Stability of output protein
This module is an integral part of the design, as large-scale commercialization of the drug of interest depends on finding the optimal conditions for protein production.
 about the model assumptions and predictions!
about the model assumptions and predictions!
The System
There are 6 differential equations that describe the behaviour of this system.
 about the equations and what they mean!
about the equations and what they mean!
Summary of simulation results
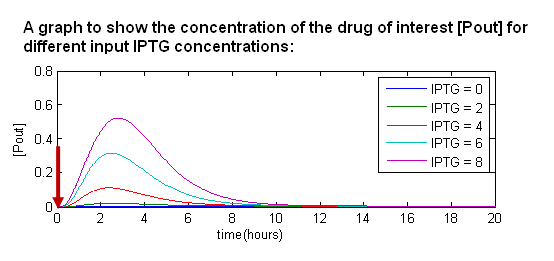
- When we introduce IPTG into the system, it temporarily removes LacI from the system. Hence, during this period of time, we produce the drug of interest.
- When the effects of IPTG wear off, the system returns to equilibrium.
- The more IPTG we add in, the higher the amount of output protein.
- The effects of IPTG toxicity were investigated and we found that for these concentration ranges, IPTG is not toxic to cells. Click on the link to see the analyzed results: IPTG growth curves
- The constants in this model are arbitrary. We justify our usage of these values with a more detailed dynamical analysis of the system, which shows that it can only have fixed points[1]. System stability analysis (read about the equations before opening this file, it's intimidating!)
Conclusions
NOTE: These will be better understood once the reader has gone through the details ("Learn More").
- The greater the strength of the Lac promoter, the greater the repressive action of LacI prior IPTG induction.
- The greater the Lac promoter leakiness (kleak) the greater the basal amount of expression of protein of interest, prior IPTG induction.
- The greater the amount of IPTG introduced, the greater the size of the bump in production of protein of interest.
- Here we assumed that the range of IPTG we have introduced is non-toxic for our cells. Growth curves will tell us whether IPTG does limit cell growth at the ranges we are interested in.
References
[1]Steven H. Strogatz (1994). Nonlinear dynamics and chaos: with applications to physics, biology chemistry and engineering. Addison Wesley. ISBN 0-201-54344-3
[2]3.Kuhlman T, Zhang Z, Saier MH Jr, & Hwa T (2007) Combinatorial transcriptional control of the lactose operon of Escherichia coli. - PNAS 104 (14) 6043-6048
 "
"