Team:Imperial College London/M3/RestrictionEnzymes
From 2009.igem.org
(→Restriction Enzymes) |
|||
| Line 1: | Line 1: | ||
{{Imperial/09/TemplateTop}} | {{Imperial/09/TemplateTop}} | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
=[[Image:II09_Thumb_m3.png|40px]]<font size='5'><b>Module 3: Genome Deletion Overview</b></font>= | =[[Image:II09_Thumb_m3.png|40px]]<font size='5'><b>Module 3: Genome Deletion Overview</b></font>= | ||
==Restriction Enzymes== | ==Restriction Enzymes== | ||
| Line 37: | Line 30: | ||
<html><a href="https://2009.igem.org/Team:Imperial_College_London/Stomach"><img style="vertical-align:bottom;" width=90px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Learnmore.png"></a></html><br><br> About the ethical implications of live organisms. | <html><a href="https://2009.igem.org/Team:Imperial_College_London/Stomach"><img style="vertical-align:bottom;" width=90px align="left" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Learnmore.png"></a></html><br><br> About the ethical implications of live organisms. | ||
| - | |||
| - | |||
| - | |||
--> | --> | ||
| + | <br> | ||
{{Imperial/09/Division}} | {{Imperial/09/Division}} | ||
| + | <center><b>Module 3 - Genome Deletion</b></center> | ||
| + | |||
| + | <html><center><a href="https://2009.igem.org/Team:Imperial_College_London/M3"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Homepageimage3.png"></a><a href="https://2009.igem.org/Team:Imperial_College_London/Temporal_Control/M3/DamMethylation"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Homepageimage3.png"></a><a | ||
| + | href="https://2009.igem.org/Team:Imperial_College_London/Temporal_Control/M3/Genetic"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Homepageimage3.png"></a><a | ||
| + | href="https://2009.igem.org/Team:Imperial_College_London/M3/Wetlab"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Homepageimage3.png"></a><html><a | ||
| + | |||
| + | href="https://2009.igem.org/Team:Imperial_College_London/M3/Modelling"><img style="vertical-align:bottom;" width="20%" src="http://i691.photobucket.com/albums/vv271/dk806/II09_Homepageimage3.png"></a><center></html> | ||
| - | |||
| - | |||
| - | |||
<html><table border="0" style="background-color:transparent;" width="100%"> | <html><table border="0" style="background-color:transparent;" width="100%"> | ||
<tr><td width="0%"></td> | <tr><td width="0%"></td> | ||
| - | <td width=" | + | |
| - | <td width=" | + | <td width="20%"><center><a href="https://2009.igem.org/Team:Imperial_College_London/M3"><b>Module 3 Overview</b></a></center></td> |
| - | <td width=" | + | |
| - | <td width=" | + | <td width="20%"><center><a href="/Team:Imperial_College_London/M3/DamMethylation"><b>DAM Methylation</b></a></center></td> |
| + | |||
| + | <td width="20%"><center><a href="https://2009.igem.org/Team:Imperial_College_London/M3/Genetic"><b>Genetic Circuit</b></a></center></td> | ||
| + | |||
| + | <td width="20%"><center><a href="https://2009.igem.org/Team:Imperial_College_London/Temporal_Control/M3/Wetlab"><b>WetLab</b></a></center></td> | ||
| + | |||
| + | <td width="20%"><center><a | ||
| + | href="https://2009.igem.org/Team:Imperial_College_London/M3/Modelling"><b>Modelling</b></a></center></td> | ||
| + | |||
<td width="1%"></td> | <td width="1%"></td> | ||
</tr></table></html> | </tr></table></html> | ||
{{Imperial/09/TemplateBottom}} | {{Imperial/09/TemplateBottom}} | ||
Revision as of 13:57, 8 October 2009

 Module 3: Genome Deletion Overview
Module 3: Genome Deletion Overview
Restriction Enzymes
In our system, the restriction enzymes DpnII and TaqI are produced.
These restriction enzymes will cleave DNA at recognition sites. This leads to a double-stranded breakage in DNA, which will subsequently result in cell death unless repair is performed in time.
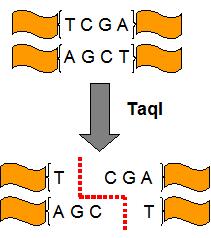
DpnII and TaqI are 4 base cutters, specifically targetting and cutting the sequences GATC and TCGA respectively (refer to diagram).
4 base cutters were chosen as they have a higher frequency of cleavage. Assuming equal distribution of nucleotides, the probability of cleavage is (1/4)4, which means that on average the four cutter will on average cut every 256 base pairs. Given that the genome of E.coli is around 4 million base pairs, it will become totally digested.
With more cleavages, the repair system would not be able to cope with multiple cleavages, so the genetic material contained within the cell will all be destroyed, including any inserted DNA.
A distinct advantage of using restriction enzymes for our 'killing' mechanism is that the the genetic material is removed, but the cell membrane is left intact. Therefore, the protein of interest will still be protected by the encapsulated cell. This renders the bacterium no more than an inanimate shell containing our protein drug of interest.





 "
"