Team:Imperial College London/Drylab/Genome deletion
From 2009.igem.org
(→Summary of Simulation results) |
(→References) |
||
| Line 79: | Line 79: | ||
===References=== | ===References=== | ||
| - | [1] | + | [1] Alon, U (2006) An Introduction to Systems Biology: Design Principles of Biological Circuits - Chapman & Hall/Crc Mathematical and Computational Biology |
[2]Jana NK, Roy S, Bhattacharyya B, and Mandal NC. Amino acid changes in the repressor of bacteriophage lambda due to temperature-sensitive mutations in its cI gene and the structure of a highly temperature-sensitive mutant repressor. Protein Eng 1999 Mar; 12(3) 225-33. | [2]Jana NK, Roy S, Bhattacharyya B, and Mandal NC. Amino acid changes in the repressor of bacteriophage lambda due to temperature-sensitive mutations in its cI gene and the structure of a highly temperature-sensitive mutant repressor. Protein Eng 1999 Mar; 12(3) 225-33. | ||
{{Imperial/09/TemplateBottom}} | {{Imperial/09/TemplateBottom}} | ||
Revision as of 23:14, 9 October 2009

Genome Deletion
Based on the genetic circuit, we know that
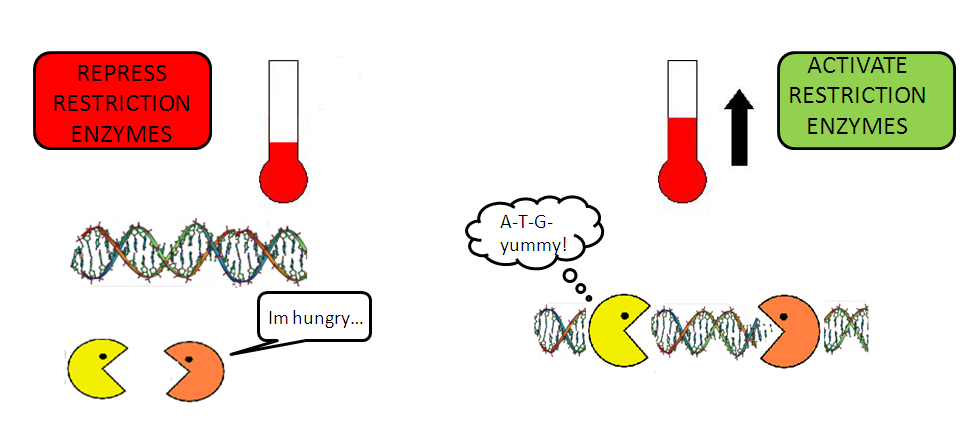
- The lambda cI promoter is repressed by the protein cI that is produced constitutively under the strong promoter J23114.
- At 28°C, functional protein cI will bind to the lambda cI promoter to repress it. Restriction enzymes DpnII and TaqI will not be produced.
- When there is an increase in temperature (from 28°C to 42°C [http://www.ncbi.nlm.nih.gov/pubmed/15652426?ordinalpos=13&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_DefaultReportPanel.Pubmed_RVDocSum [1]][http://www.ncbi.nlm.nih.gov/pubmed/10235623 [2]]), there will be a de-repression of lambda cI promoter, causing restriction enzymes DpnII and TaqI to be produced.
Contents |
Our goals
We aim to:
- Explore how temperature correlates to restriction enzyme concentration, and see how it affects the population of live cells, so as to characterise the effects of temperature on cell death.
- Develop a model of the number of dead cells as this correlates to our live and dead cells assay. (hyperlink to assay). From this, we can monitor the rate of killing and perform data analysis.
 about the model assumptions and predictions!
about the model assumptions and predictions!
The system
The system is made up of 6 ODEs based on the Module 3: Genome deletion genetic circuit. ODEs are also used in modelling the population of E. coli.
 about the equations and what they mean!
about the equations and what they mean!
Summary of Simulation results
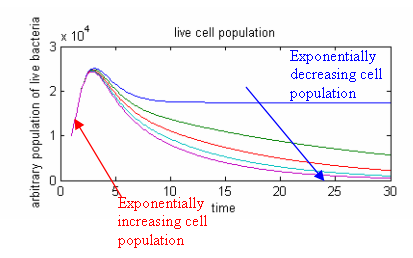
Temperature above threshold – Cell population goes to zero
- If the temperatures are above threshold, the population growth of live cells will be constrained by the concentration of restriction enzyme.
- The maximum live cell population at a given temperature will also be lower for higher temperatures
- If the lambda cI promoter is strong, there will be a higher rate of cell death than cell growth; we will see a decrease in live cell population. However, if lambda cI promoter is weak, there might not be enough killing, so we will only see a reduction in growth of cell population, ie: a slower increase in population.
Conclusions
References
[1] Alon, U (2006) An Introduction to Systems Biology: Design Principles of Biological Circuits - Chapman & Hall/Crc Mathematical and Computational Biology
[2]Jana NK, Roy S, Bhattacharyya B, and Mandal NC. Amino acid changes in the repressor of bacteriophage lambda due to temperature-sensitive mutations in its cI gene and the structure of a highly temperature-sensitive mutant repressor. Protein Eng 1999 Mar; 12(3) 225-33.
 "
"