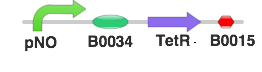Team:NTU-Singapore/Project/Approach
From 2009.igem.org
Our Approach
We wish to design a biological system to combat atherosclerotic plaque that can eventually be deployed in modified T-helper cells.
Due to time & equipment shortage during the iGEM period, we will only demonstrate the proof-of-concept system using the E.Coli K12 strain.
Research Methodology
In our project pLaqUe Out!, we will be dealing with 4 devices to arrive at a complete system.
The 4 devices are :
- Sensing device with nitric oxide sensing promoter (pNO)
- Degradation device for cholesteryl esterase (CHE) production
- Imagine device for infrared-fluorescent protein (IFP) production
- Dilation device for nitric oxide synthase (NOS) production
These devices are designed to work in sync to provide a robust systemic solution to plaque imaging & breakdown.
Sensing device
pNO is NorR-regulated sigma-54 promoter. NorR is a transcription factor which is highly responsive to the presence of nitric oxide. Therefore, we would be regulating the expression of a gene downstream of our pNO by varying the input of nitric oxide.
- The NO-sensitive promoter region of the NorV/W gene sequence (pNO), endogeneous in E.Coli, is used.
- This promoter will be the active NO-sensor for our system.
- pNO is ON in high [NO] conditions & OFF in low [NO] conditions.
- This promoter will be the active NO-sensor for our system.
- The pNO will be placed upstream of a TetR gene sequence.
- TetR is used to repress the other devices.
- In healthy [NO] conditions, pNO is activated and will facilitate expression of TetR.
- In low [NO] conditions, pNO will be deactivated and the transcription of TetR gene sequence will stop. Hence repression of other devices will stop.
In order to realize this sub-system, we will have our gene for pNO directly synthesized from GeneART.
We will characterize the pNO by the expression of downstream green-fluorescent protein (GFP). The protocols for this are available on the OpenWetWare (OWW) website and we will optimize them for our use. We will have sodium nitroprusside (SNP) as the source of nitric oxide, and we will vary the concentrations of SNP accordingly to obtain our desired input values.
Thus, we will be able to characterize the activity of our pNO via the GFP output in fluorescence time course (fluorescence versus time), absorbance time course (absorbance versus time) and fluorescence/OD600 versus time.
Please proceed here to see our detailed page on the Sensing device.
Degradation device
This sub-system will secrete cholesteryl esterase (CHE) which is able to catalyze the hydration reactions of cholesteryl esters into free cholesterol and fatty acids. The degradation of cholesteryl esters through these reactions will lead to the size reduction of the atherolsclerotic plaque.
- The pTet promoter will serve as the controller for this device.
- In high [TetR] conditions, implying high [NO] conditions, Degradation device is OFF.
- In low [TetR], implying low [NO] conditions, this device will be ON.
- LuxI gene sequence expresses LuxI protein.
- LuxI protein catalyzes conversion of SAM protein to AHL.
- This conversion has significance in the Dilation device.
- CHE gene sequence will express Cholesteryl Esterase enzyme.
- This enzyme will breakdown the cholesteryl esters in plaque.
This gene for CHE, originally obtained from Pseudomonas aeruginosa, will be directly synthesized from GeneART.
Since the ultimate objective of this sub-system is to characterize the functionality and activity of CHE, we will induce the expression of CHE via constitutive promoters such as Part BBa_J23101 and Part BBa_J23119 from the Registry.
With this construct, we can readily perform functionality test with the Amplex(R) Red Cholesterol Assay Kit from Invitrogen to quantitatively analyse the enzyme activity of CHE. Nevertheless, SDS-PAGE will also be carried out to confirm the expression of the cholesteryl esterase of our construct.
Please proceed here to see our detailed page on the Degradation device.
Modelling Approach
Our models will be mathematically deduced in the form of ordinary differential equations. These ODEs will relate the concentration of mRNA & protein output of the gene sequence of interest w.r.t time. We will use known and/or empirical constants such as maximum translation rate, maximum transcription rate, dissociation constants etc to assume for maximum efficiency case.
These equations for each construct are then re-constructed as a Simulink subsystem, which in turn are displayed as blackboxes in the Simulink version of the final system. We will also use [http://www.tinkercell.com/Home TinkerCell] to generate both deterministic & stochastic modelling predictions for our system.
The modelling process is almost entirely theoretical, except for abovementioned constants which are literature-derived.
- So firstly predicted outputs from the model can actually help us to envision the scale at which the system works. This is important to verify if the system can even ideally be used in the application that we want.
- Furthermore, by identifying larger deviations in output of the actual system compared to predicted model, the model can also help as a “debugger”.
- The models can also help us to enhance the system by identifying bottlenecks; identifying the assumptions made in simplifying calculations can indirectly point us to the potential sources of error in experimental verification.
References/Literature
See the complete list of references here.
 "
"