Team:DTU Denmark/notebookuserfusion
From 2009.igem.org

| Home | The Team | The Project | Parts submitted | Modelling | Notebook |
|
Activities relating to our two sub-projects: - The Redoxilator - The USERTM assembly standard - Biobricks - Protocols |
Day-to-day activities Activities relating to the USERTM fusion assembly standard Construction of biobrick BBa_K194003 and yeast plasmid containing biobrick BBa_K194003 for demonstration. Two complentary oligoes were ordered so that they could anneal forming the biobrick BBa_K194003 holding a PacI restriction site flanked by two Nt.BbvCI-sites and with XbaI and SpeI sites in the end for upholding the biobrick standard. TCTAGAGGCTGAGGGTTTAATTAAGACCTCAGCGCAGTGGTGCGATCGCGACACTGCTACTAGT AGATCTCCGACTCCCAAATTAATTCTGGAGTCGCGTCACCACGCTAGCGCTGTGACGATGATCA The two primers were warmed up to 95 °C on PCR machine and slowly cooled down with 5°C every 5 minutes until it reached 25°C for the two primers to anneal correctly. Then the double stranded oligo was digested with XbaI and SpeI o/n (over night). In the morning the oligo was warmed up to 95 °C again and cooled down as former described. This method was chosen to denature the restriction enzyme before ligation as the oligoes would have been lost in a standard DNA purification. The oligo was inserted into two different plasmids: 1. pRS416 : in-house yeast plasmid with URA marker that we wished to use to demonstrate that Biobrick BBa_K194002 worked in yeast. The two plasmids were digested seperately using two enzymes, XbaI (20U) and SpeI (10U). The mixtures were incubated at 37 °C for eight hours after which more enzyme was added to ensure maximum digestion. After an additional two hours of incubation the digested plasmids were dephosphorilised to prevent reanealing during ligation. This was achieved by an Antartic Phosphatase treatment (25 U). The DNA was gel-purified and the resulting concentration was determined using fluorometrics (The Qubit Quantitation PlatformTM from invitrogen). The ligation was performed using T4-DNA ligase using a 3:1 ratio of fragment to vector and 9 ng DNA in total at a volume of 20 µl according to the manufactors recommendations. The actual ligation was carried out on a PCR machine for 1½ hours shifting the temperature from 10 °C to 30 °C at 30 second intervals as desribed by [1]. The final product was finally chemically transformed into competent E coli cells. Selection of transformants Ten random colonies were picked for overnight growth in liquid media. The next day the plasmids were purified and digestion were performed with PacI and XbaI to verify the presence of the biobrick. This was done as the presence of the biobrick is introduced a PacI site in the vector. The XbaI digestion was done to ensure that the biobrick had been inserted with the right directionality.
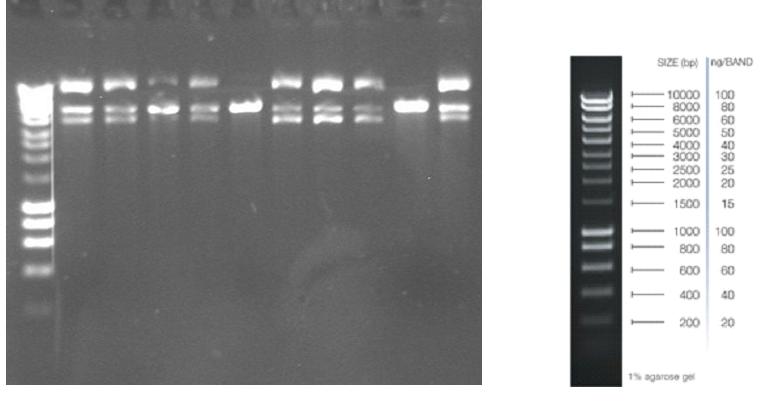
Figure 1. Left: PacI restriction assay, BBa_K194003 insertion into pRS416. Restriction performed on ten randomly picked, o/n grown and plasmid purified samples. Samples are run on a 1 % agarose gel. Digestion should leave a single band at approxiamately 6 kb. Colonies 5 and 9 are believed to contain the correct plasmid. Right: The ladder used in the corresponding experiment, Hypper Ladder 1. From the gel it was seen that the BBa_K194003 had been inserted correctly into pRS416. Now this was used to demonstrate that biobrick BBa_K194002 was functional. This was done by opening the USER cassette in BBa_K194003 with PacI and Nt.BbvCI, purifying the backbone from gel and adding two PCR fragments: 1. The constitutive TEF promoter (400bp) using the primers BGHA200/370
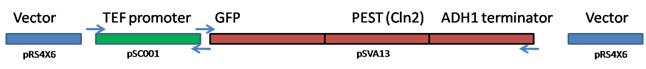
Figure 2. The cloning strategy for demonstrating that biobricks BBa_K194002 works. Two PCR products were fused using USER fusion and inserted into the pRS416 backbone by USER cloning. After treatment with USERTM mix, the mixture was transformed into E. coli. VerificationFollowing selection of transformants and overnight liquid cultures the plasmids was purified . The plasmids were linearized using AsiSI, which is also included in the BBa_K194003. Plasmids holding the fusion of TEF and BBa_K194002 will show to be 2kb longer than the ones without. The presence of the gene were only verified by PCR of the genes inserted. Transformation into yeast Yeast (CEN PK 113 5D (Ura-)) was transformed with this plasmid using the Ura marker present on pRS416 Yeast was also transformed with pSC011 kindly provided by Simon Carlsen, CMB containing the sequence from BBa_K194001 under the constitutive expression of the TEF promoter. pSC011 holds the same promoter and terminator as the pRS416 holding the BBa_K194003 and can therefore be used to analyze the difference of the addition of the Cln2 degradation signal. BioLector fermentation To characterize the yeast strains that we have constructed in the project we used the BioLector machine kindly provided by the Center for Biological Sequence Analysis at DTU.
We characterized: To minimize the background fluorescence from the media special media without X and Y was made. References [1] Lund, A. H., M. Duch, and F. S. Pedersen. 1996. Increased cloning efficiency by temperature-cycle ligation. Nucleic Acids Res. 24:800-801. doi:l50400 October 3rd Primer design software: Server uplink and HTML interface implemented. September 23rd Primer design software: Work on article commenced. September 22nd Primer design software: Data structure finalized. September 14th Primer design software: Statistical tests program performance. September 12th Primer design software: Preliminary data tests performed. September 4th Primer design software: Graphical output format settled. September 1st Primer design software: TM adjustment algorithm implemented. August 24th Primer design software: Fusion tail sorting algorithm pseudo code complete. August 22nd Primer design software: Work on fusion tail sorting algorithm. July 15th Primer design software: General program parameters determined. July 10th Primer design software: Preliminary meeting on primer design software. |
Work process Our team works parallel in smaller sub-teams. Some of us work hard in the lab, while others are in the process of developing software and the in silico model of the Redoxilator system. However, we constantly keep each other updated, and meet often to exchange ideas and take turns at the different tasks, thus exhausting all of our combined knowledge in every aspect of this project. |
| Comments or questions to the team? Please Email us -- Comments of questions to webmaster? Please Email us |
 "
"