Team:Waterloo/Project
From 2009.igem.org
Contents |
Abstract:
Chromobricks: A Platform for Chromosome Engineering with BioBricks
The aim of our project is to develop a fully-featured platform for chromosome engineering, allowing the in vivo assembly of a synthetic chromosome from interchangeable parts, followed by selective degradation of the native chromosome. We have designed a proof-of-concept for chromosome-building that will use the site-specific integrase of phage ΦC31 to integrate a BioBrick into a defined locus of the E. coli genome. Six pairs of integrase-targeted att sites have been designed to be non-cross-reactive in order to support repeatable cassette-exchange reactions for chromosome building. We have also written software to model the integrase-mediated rearrangement of DNA molecules containing att sites, to aid the design of more elaborate chromosome-building systems. To selectively degrade the native chromosome we designed a nuclease-based, inducible genome-degradation system. In its simplest form, our system can be used to integrate biological devices into a chromosome in situations requiring stable copy number and selection-free maintenance.
Project Details
Background
Chromosome Engineering provides many benefits over plasmid based engineering, including precise copy number and stable maintenance without selection.Imprecise copy number plasmid-based systems can add unnecessary noise to synthetic biology systems, limiting the potential complexity of these systems. However, a plasmid can easily be isolated, engineered with restriction enzymes and reintroduced into a host strain. Chromosomes cannot be engineered in this way.
Traditionally, methods to engineer chromosomes in bacteria have been based on homologous recombination, which is an inefficient process. Other methods have taken advantage of recombinant lysogenic phage, such as the E. coli phage λ. These phage integrate their genome into the bacterial chromosome via a site-specific recombination. Genes that have been added to the small phage genome will be carried along. The mechanisms of phage genome integration have been studied and there are some well characterized systems. These systems can be separated from the phage itself. They can be used to engineer DNA molecules in vitro or in vivo without actually infecting the bacteria with a phage (Groth & Calos, 2004).
The phage integration system usually involves an integrase/recombinase enzyme, the phage attachment site (attP), the bacterial attachment site (attB), and for the reverse reaction an excisionase enzyme. The integrase enzyme catalyzes a stand exchange between attP and attB sites on the DNA yielding attL and attR sites which are composites of attP and attB sites. If there is excisionase and integrase present the reaction will go in the reverse direction. The well known λ integration system is the basis of Invitrogen's Gateway® technology. Gateway® technology uses the activity of the λ integrase and excisionase to carry out cassette exchange reactions, where sections of two molecules each flanked by respective att sites are exchanged. The system is used to engineer plasmids as an alternative to traditional cloning using restriction enzymes. For our project, the goal of engineering chromosomes, without conflicting with the widely adopted Gateway® system, led us to choose the integrase from the Streptomyces phage C31 (ΦC31).
The ΦC31 Integrase
The ΦC31 integrase catalyzes a unidirectional stand exchange between a 39 bp attP site and a 34 bp attB site (Groth et al., 2000). The enzyme works by making a synapse between attP and attB, making double strand breaks producing a 2 bp sticky overhang, exchanging the strands, and re-ligating them in the recombinant configuration (attL and attR).
Our BBa Compatible Cassette Exchange System
Due to the lack of any BBa compatible chromosome engineering tools we decided to build a cassette exchange system based on the ΦC31 integrase. The system can be broken down into three components: the landing pad strain (LPS), the helper plasmid, and the BBa donor vector.
The protocol to integrate a BioBrick into the chromosome of the LPS involves a conjugation of a section of the donor plasmid via bi-parental mating into the LPS which is also expressing ΦC31 integrase. Then the LPS strain is selected with naladixic acid, and the loss of the landing pad markers is selected with sucrose. The colonies would then be screened for kanamycin sensitivity. The LPS contains a Landing Pad cassette in the melA locus. This cassette consists of sacB,rfp,and nptII(kanamycin resistance) genes flanked by ΦC31 attP sites. After cassette exchange these markers are lost. The BioBrick will replace them.
The Goal of on Chromosome Assembly of Interchangeable Parts
With the design of non-cross reactive att sites it would be possible to regenerate the landing pad after a successful integration. This would be done by having a second non-cross-reactive pair of att sites downstream of our integrated BioBrick (attP2 attP2). A landing pad regeneration plasmid with attB2 sites flanking a landing pad would be introduced and a ΦC31 integrase reaction would integrate the landing pad. Another attP1 attB1 reaction could be carried out to integrate another BioBrick. The problem with an approach like this is that attL sites would build up. We decided to write software to try and solve this problem. The software is further discussed in the modelling section.
We have designed six different attB sites and six different attP sites, giving us six non-cross-reactive pairs. This is discussed further below.
Selective Chromosome Degradation
There are examples in the literature of whole genomes being reconstructed in a locus of an existing chromosome. One of these examples is assembly of the Synechocystis PCC6803 genome in the Bacillus subtilis 168 genome (Itaya et al., 2005). This was done through a tedious homologous recombination based approach.
We figured that in the future improved chromosome engineering systems would be used to assemble synthetic genomes in host cells. After that it would be possible to circularize the synthetic genome and degrade the native genome. This would essentially change the species of the cell.
As a proof of concept we have designed an inducible nuclease construct that will express a restriction endonuclease (PmeI) and an exonuclease (T7 gene 6 exonuclease). As long as the synthetic chromosome does not contain a PmeI site it will not be degraded.
The Experiments
Cassette Exchange
The goal of this aspect of the project was to create a system capable of integrating a BioBrick into the chromosome of E. coli at a defined locus. The steps required for integration, baring initial cloning of the BioBrick, would be carried out in vivo.
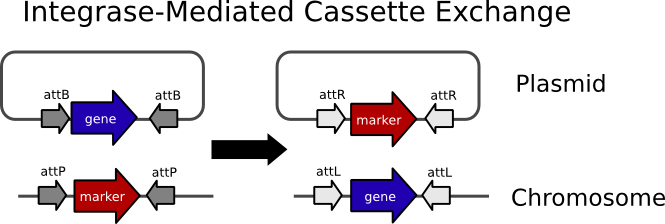
The basic idea behind cassette exchange (hereinafter IMCE) is pictured below.
We designed our system so that a plasmid carrying a BioBrick would by transferred into the Landing Pad Strain (LPS) via conjugation in such a way that it would be lacking an origin of replication. The integration reaction would take place causing the markers to be lost due to their transfer to a non-replicating molecule. In the case of our system the markers are nptII (kanaymycin resistance), sacB(sucrose sensitivity), and rfp. That way integrants can be selected on media containing sucrose. The design also calls for the landing pad strain to be nalidixic acid resistant so that it can be selected for after conjugation. Therefore integrants can be selected for in one step on a nalidixic acid sucrose plate.
BBa Donor Plasmid
Landing Pad Strain
We have designed a cassette system that will allow us to easily detect occurrence of Integrase Mediated Cassette Exchange (IMCE). The cassette is a landing pad flanked by attP’s that is integrated on to the chromosome of the host strain, E.coli K12 MM294A, via homologous recombination (The host strain must have RecA activity for homologous recombination to occur). The landing pad contains several screening and selection markers, including RFP, kanr, and sacB. In the cell, these markers produce easily detectable phenotypes that allow us to track the occurrence of cassette exchange. Red Fluorescent Protein, or RFP, is a widely used fluorescence marker that can be used for tracking protein expression in vivo. Kanamycin resistance is conferred by the kanr gene and can be used as a selection marker. SacB encodes the protein levansucrase, which polymerizes sucrose and confers sucrose sensitivity. The entire landing pad construct is built on a suicide vector derived from pKNG101, so that, after the landing pad is integrated on to the chromosome, vector backbone will be lost due to its inability to replicate.
Integrase expression plasmid
Non-cross-reactive att sites
We set out to design and test 6 different att site pairs that would be non-cross-reactive, or "orthogonal". This would allow for increased specificity when inserting DNA pieces into the chromosome. It would also allow for repeatable insertion of DNA pieces into the chromosome as long as a build up of attL sites was not an issue.
First we discuss the mechanism of the integrase; then we move on to the design and testing of our "orthogonal att sites".
Integrase Mechanism
The ΦC31 integrase catalyzes a unidirectional strand exchange between a 39 bp attP site and a 34 bp attB site. The enzyme works by making a synapse between attP and attB, making double strand breaks producing a 2 bp sticky overhang, exchanging the strands, and re-ligating them in the recombinant configuration (attL and attR). This reaction is not reversible with only the integrase present. The att sites can be mutated to produce different overhangs. A mutant attB with different a different overhang (A.K.A core dinucleotide)has been shown, in a related system, not to recombine with attP (Ghosh et al., 2003). Instead of a recombination activity the integrase showed a topoisomerase activity (reducing supercoiling on a plasmid) suggesting the double stranded breaks were made but the DNA would not re-ligate in the recombinant configuration (Ghosh et al., 2003). When identical mutations are made to the overhangs the recombination activity is restored (Ghosh et al., 2003), implying that making orthogonal att sites (non cross-reactive site pairs) for the ΦC31 integrase is possible.
The sequence of the overhangs also has implications in the directionality of the recombination, which can lead to different end results of recombination (excision or inversion for example)(Ghosh et al., 2003). The palindromic overhangs will lead to loss of directional specificity. Given the 16 possible core dinucleotide sequences six of them should be completely non-cross reactive while maintaining direction specificity.
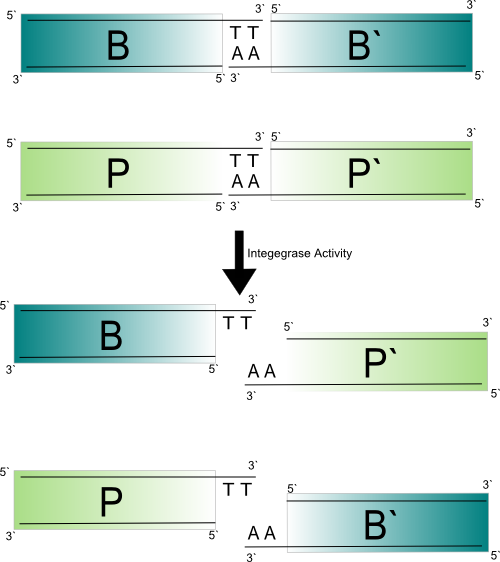
The integrase mechanism is shown below. The integrase protein forms a dimer about each att site. When the att sites are brought together, presumably through the two dimers recruiting each other, a tetramer is formed. When the tetramer is formed between an attB and an attP the small n-terminal catalytic domain is activated. A double strand is made. For each protein a covalent linkage between a serine residue and its respective 5` phosphate is made. The complex rotates 180 degrees and re-ligates the DNA in a recombinant configuration.
Design of new att sites
The above figure shows that the completion of the integrase mediated recombination reaction relies on the successful base-pairing of the two base-pair 3` overhangs between attB and attP halves. If there is a mutation that disrupts this the integrase complex will continue to rotate and the products will be reformed. As mentioned earlier, the base-pairing of the 3` overhangs produced by the core dinucleotides also contrains which orientation the att sites can recombine in. If the core dinucelotide is palindromic directional specificity is lost (Ghosh et al., 2003). This also implies reactivity between att sites with reverse compliment core dinucleotides.
To make "orthogonal att sites" for phiC31 integrase sites with different core dinucleotides (3` overhangs) were designed. We avoided palindromic core dinucleotides (GC,TA etc) and reverse compliment core dinucleotides(TT with respect to AA). Of the sixteen possible dinucleotides four are palindromic. Twelve are not palindromic, and will confer directionally specific recombination. Those twelve consist of 6 pairs of sense and reverse compliment dinucleotides. That leaves six mutually exclusive dinucleotides that should display no cross-reactivity at all. We chose TT, GG, TG, AC, GA, and CT dinucleotides.
The att sites are shown below with the core dinucleotide bolded. The only difference between orthogonal att sites is the core dinucleotide.
attB34 TT: 5`-GGTGCCAGGGCGTGCCCTTGGGCTCCCCGGGCGCG- 3`
attP39 TT: 5`-CCCCAACTGGGGTAACCTTTGAGTTCTCTCAGTTGGGGG- 3`
Design of experiment to characterize new att sites
To test the functionality and quantify the efficiency of recombination between six different att site pairs an in vivo recombination experiment was designed.
Complimentary oligonucleotides were used to synthesize the att sites, and to serve as linkers for ligation into different vectors.
- The attP sites were to be cloned into a very narrow host range vector that only replicates in select strains of E. coli (lambda pir lysogens). This vector carries streptomycin resistance.
- The attB sites were cloned into pSB1A2, which carries ampicillin resistance. This vector replicates in most E. coli strains.
-The attB plasmid was to be transformed into E. coli DH5alpha carrying the integrase expressing plasmid.
- The attP plasmid was to be conjugated from E.coli SM10 into the above DH5alpha (attB and integrase plasmid carrying) strain.
-DH5alpha is nalidixic acid resistant SM10 is not. The streptomycin resistance carrying attP plasmid does not replicate in DH5alpha
- The cells from the conjugation mixture would be plated on media containing naladixic acid and streptomycin. The only way a streptomycin resistance can be maintained is thorough integrase mediated recombination.
- The number of colonies relative to a positive control (conjugation into DH5alpha lambda pir) would provide efficiency of recombination numbers.
Conjugation was chosen instead of transformation because of conjugation's consistent efficiency between experiments. Transformation efficiencies vary widely between experiments.
Selective Chromosome Degradation
Results
References
Ghosh, P., Kim, A. I. & Hatfull, G. F. (2003). The orientation of mycobacteriophage Bxb1 integration is solely dependent on the central dinucleotide of attP and attB. Mol Cell 12, 1101-1111.
Groth, A. C., Olivares, E. C., Thyagarajan, B. & Calos, M. P. (2000). A phage integrase directs efficient site-specific integration in human cells. Proc Natl Acad Sci U S A 97, 5995-6000.
Groth, A. C. & Calos, M. P. (2004). Phage integrases: biology and applications. J Mol Biol 335, 667-678.
Itaya, M., Tsuge, K., Koizumi, M. & Fujita, K. (2005'). Combining two genomes in one cell: Stable cloning of the Synechocystis PCC6803 genome in the Bacillus subtilis 168 genome. Proceedings of the National Academy of Sciences 102, 15971-15976.
 "
"