Team:Freiburg software/Project
From 2009.igem.org
[http://www.youtube.com/watch?v=hM44QasLyfw Watch a high resolution verion here]
Contents |
Introduction
Motivation
Today, science communication is still pretty outdated, making little use of the collaborative world wide web. There has started a process of wiring results in synthetic biology research. We want to push this process and go one step further: Why only store and share results? Why not making use of the web to collaborate? Scientist could make transparent the whole process of creating data, they could even create data together. Today, the Web offers all the technologies needed. Make use of it!
Google Wave
Google Wave is "a personal communication and collaboration tool" developed by Google. It is a web-based service, computing platform, and communication protocol designed to merge e-mail, instant messaging, wikis, and social networking. It has a strong collaborative and real-time focus and provides several ways to extend its functionality.
Wave basically consists of an open communication protocol similar to email, as well as of client- and server-software. Like email, the protocol aims to be open, decentralized and easy to adapt, but includes modern achievements like multi-user-, real-time-communication and rich formated text with embedded data as well. At the moment, the only working server and client software for Wave is also written by Google, but with the protocol being open-source already, other - Google independent - servers and clients will soon be available.
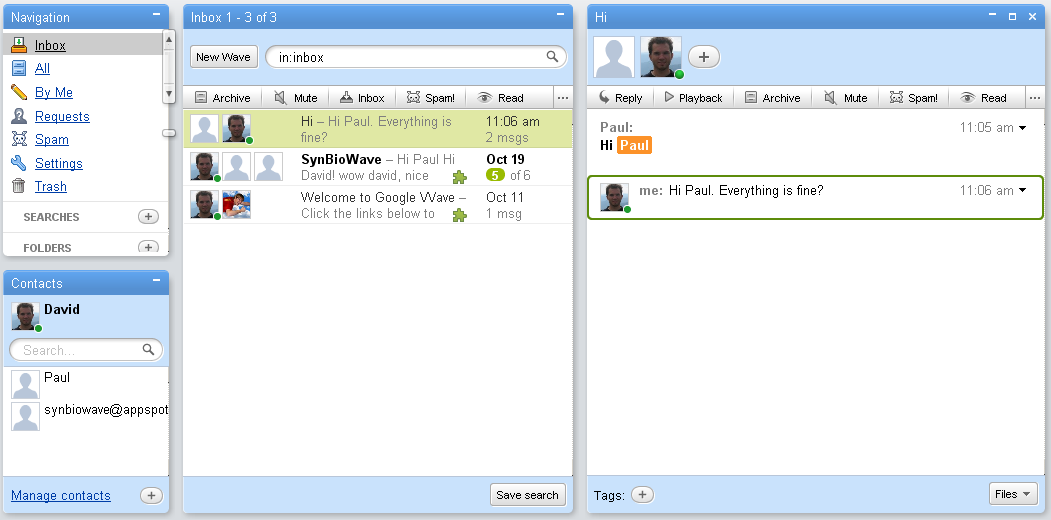
Practically a typical Wave-conversations - called a wavelet - normally works like this: User Alice creates a new Wavelet. She than invites his friend Bob to join the conversation. Bob accepts and can now write Messages to the wave. Each message creates a so called Blip. What differentiates Wave from normal instant-messaging is, that if Alice and Bob decide to write an Document together, they can start to edit the same Blip together. Each change they make to the text there is shown to the other on in real time. If they want, they can also use a build-in playback feature similar to the version-history in wikis to review the changes made to the wavelet.
Additional Google has published an API for writing so called Robots and Gadgets for Wave. While Robots are small programs written in Python or Java, which can participate in Wave similar to normal users, Gadgets are small Webpages that can be embedded into a wave-conversation.
That makes Google Wave quite interesting for Systemic Biology: Synthetic biologist work with text most of the time, Google Wave is made for collaborative text creation. Synthetic biologist need to work together in order to create artificial sequences, protein and later maybe hole genomes, Google Wave is made for working together with any embedded data. Synthetic biologist need automated features for their work, Wave offers them via robots.
BioJava
[http://www.biojava.org BioJava] is an open source project dedicated to providing a Java framework for processing biological data. It provides analytical and statistical routines, parsers for common file formats and allows the manipulation of sequences and 3D structures. The goal of the BioJava project is to facilitate rapid application development for bioinformatics.
Using the scalable, cross-platform, network-aware power of Java technology, researchers at Great Britain's famed Sanger Institute for genetic study have spawned BioJava--an open-source project dedicated to providing genomic researchers with a Java technology-based developer's toolkit.
BioJava offers bioinformatics developers over 1200 classes and interfaces for manipulating genomic sequences, file parsing, CORBA interoperability, and more. The facility is already being used at major research and pharmaceutical centers, and in over 85 countries around the world.
BioJava provides objects for all kind of synthetic biologists needs, which there are:
- Basic manipulation of sequences, like translations, building complements and doing multiple sequence alignments.
- Viewing sequences in different formats, like feature added sequence presentations and circular views.
- Import and export sequence data in different file formats.
- Accessing databases like BioSQL and [http://www.ensembl.org Ensembl].
Concept
Our concept is to create a collaborative software suite called SynBioWave for synthetic biology purpose. SynBioWave is a Google Wave extension using BioJava to add synthetic biology functionality, giving synthetic biological research access to the collaborative and interactive web 2.0. Using SynBioWave, scientists can share their results in Waves or even conduct research together from different places around the world. Users can add and modify sequences within conversations while others observe the progress or even interact. Participants can be invited to a conversation any time and track back the collaboration process using the playback function, which fully supports all biosynthetic contents.
SynBioWave makes use of Wave's powerful communication and collaboration functionality and is designed to be be easily extended with new synthetic biology functionality. Mashing up the reinvention of the email with a major library for processing synthetic biology data, raises science collaboration to a new level.
Our small team of three developers will not be able to create a full-value synthetic biology software by iGEM Jamboree 2009. Our goal is to lay the foundation for a robust software suite and to demonstrate the benefits of this wave approach for synthetic biological research. Moreover we implement some basic biological functionality to demonstrate this concept.
SynBioWaves' key features
- open source, free web application accessible from every computer connected to the internet
- strong communication and collaboration functionality
- basic synthetic biology functionality
- easy to extended with additional synthetic biology functionality
The road to success
TODO: Grafik
For addressing a wide audience of users and contributers, SynBioWave is published under a free licence. This will attract other developers creating new functions or modify the software for their own purpose.
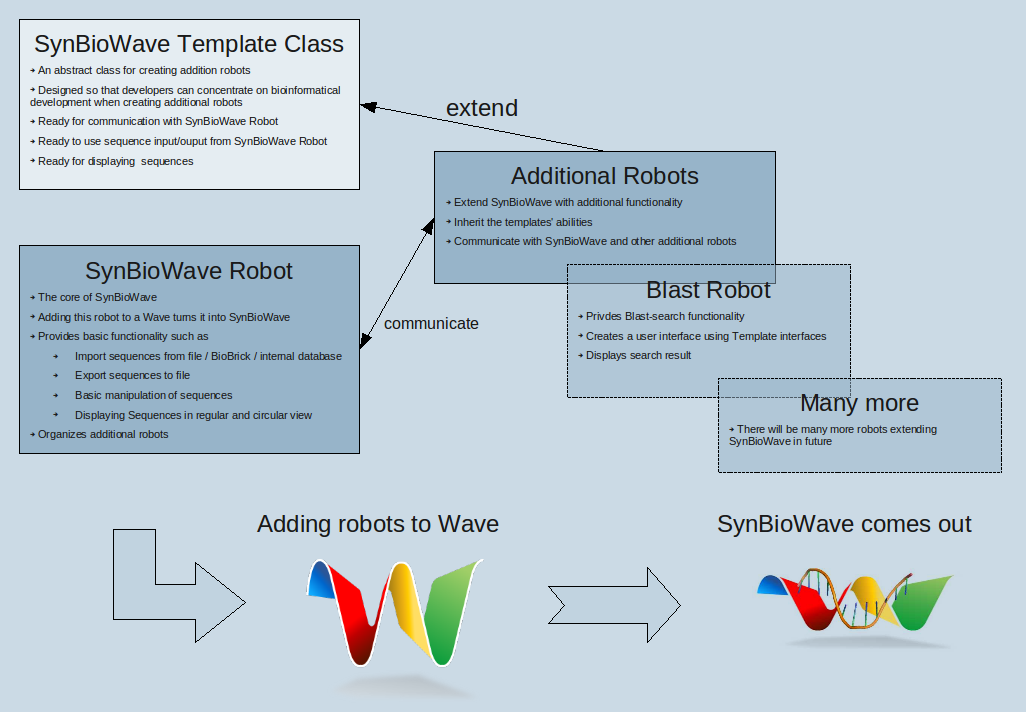
One of the key goals of SynBioWave is the feature of easy-extendibility. We want to create a framework that allows other developers to contribute new biosynthetic functionality with a minimum knowledge of Wave development. For this purpose SynBioWave offers an abstract robot class which can be regarded as a template for biosynthetic functions. This concept has very nice side-effect: SynBioWave can be easily customized by adding and removing robots which represent certain function.
SynBioWave will not only be a simple mashup, a synthetic biological software running inside a wavelet. It will be a perfect symbioses of Wave and BioJava. The look and feel of SynBiowave will perfectly fit in the Wave concept. Waves real-time-editing, multi-user-editing functions as well as the playback function must work in harmony with SynBioWave. This sounds quite trivial. But looking at the current Wave extensions gives reason to be concerned about this.
Benefits of the symbioses
Building SynBioWave as mashup of Wave and BioJava brings up certain benefits:
- no need for building the whole application from scratch
- get the communication and collaboration features for nothing (wave)
- get multi-user editing for nothing (wave)
- get real-time editing for nothing (wave)
- as a web application, SynBioWave can be accessed from any computer connected to the internet.
- easy to setup. No local installation is needed
- get many key bio features for nothing (biojava)
- robust and high quality software basis
- As Google's latest child, wave is going to be talked of a lot. As one of the first application using google wave, SynBioWave has probably a huge audience
challenge and difficulties
- With the begin of SynBioWave's development, wave is in a very unstable Alpha version available. Even the API is still weak and might change. There is not much documentation and discussion yet, many Bugs and "no-yet-implemented-features" often nearly drove us crazy.
- At the moment, Google forces developers to host Robots trough their AppEngine-project. AppEngine has every strict limitations build-in, making it incredible hard for us to implement even simple features like file up- and download. Nearly no existing bioinformatic Java-class works on AppEngine without modifications. When Google opens Wave for own robot-servers, these problems will instantly vanish. They have announced to do so in the foreseeable future.
- Developing a collaborative web application faces programmers special challenges and difficulties. For example, multi-user editing and real-time editing issues the challenge of synchronising user input. What happens, for example, if two users submit contrary input at the same time?
The Software
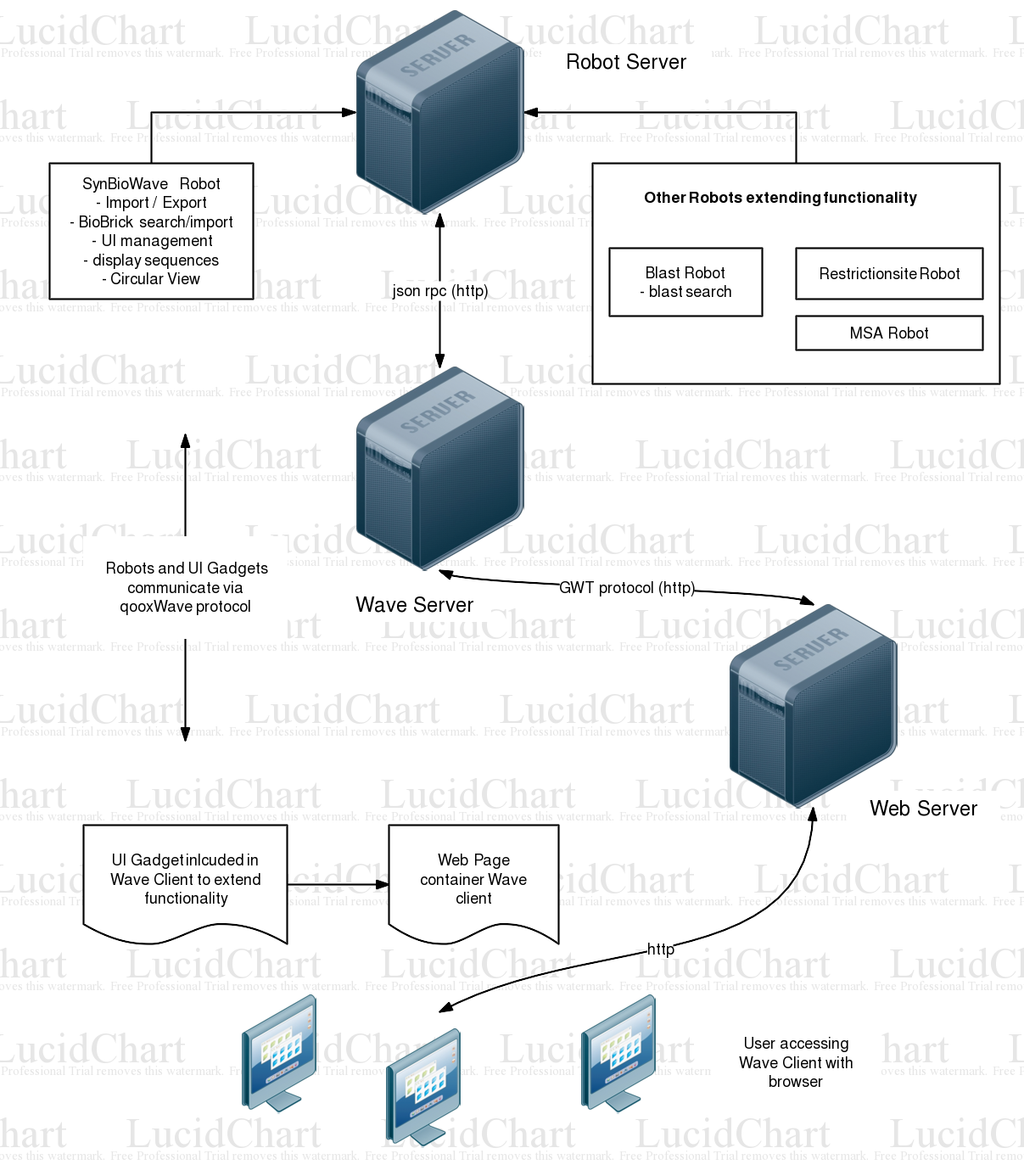
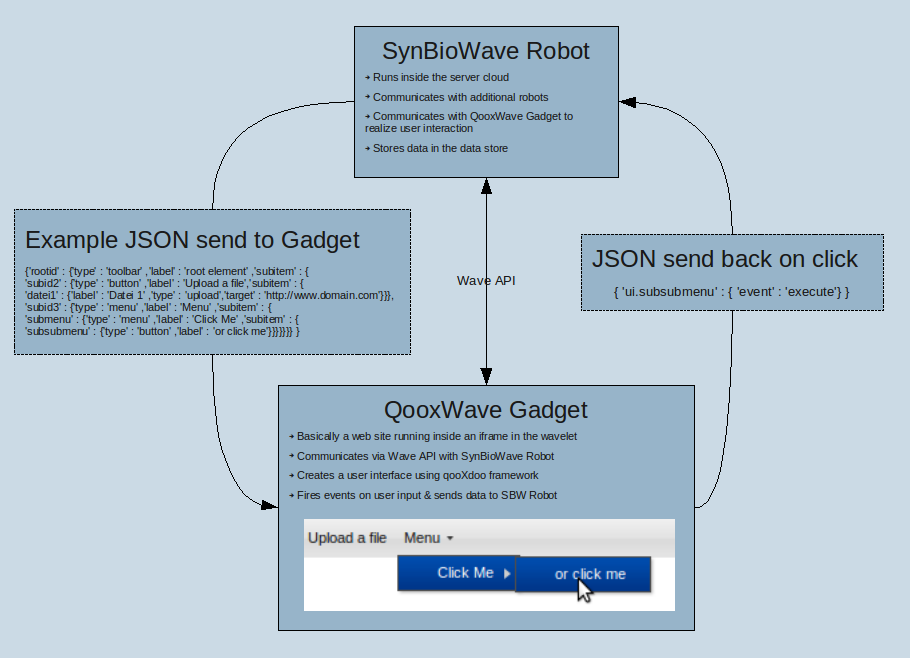
Architecture
SynBioWave is a Google Wave extension, turning Wave into a biosynthetic Software Suite. There are currently two available methods of extending Google Wave (See Google Wave for an introduction):
- Robots
- Gadgets
We are making use of both!
Robots
Robots are automated participants, able to modify the wavelets' contents they have joined and interact with real participants. Using Google's Java Client library for Robots gives us the possibility to implement biosynthetic functionality provided by the BioJava library. The SynBioWave Robot can be regarded as the core of our extension. Adding this robot to a wavelet activates SynBioWave. There are additional robots for specific functions, which can added to a wavlet to activate this certain function. The SynBioWave Robot is responsible for:
- Organization of all SynBioWave functions represented by additional robots
- extending Wave's user Interface with SynBioWave specific elements using the qooxWave protocol
- storing the important informations written to the wave in an external database
- Providing basic functionality like
- Import/export of sequences
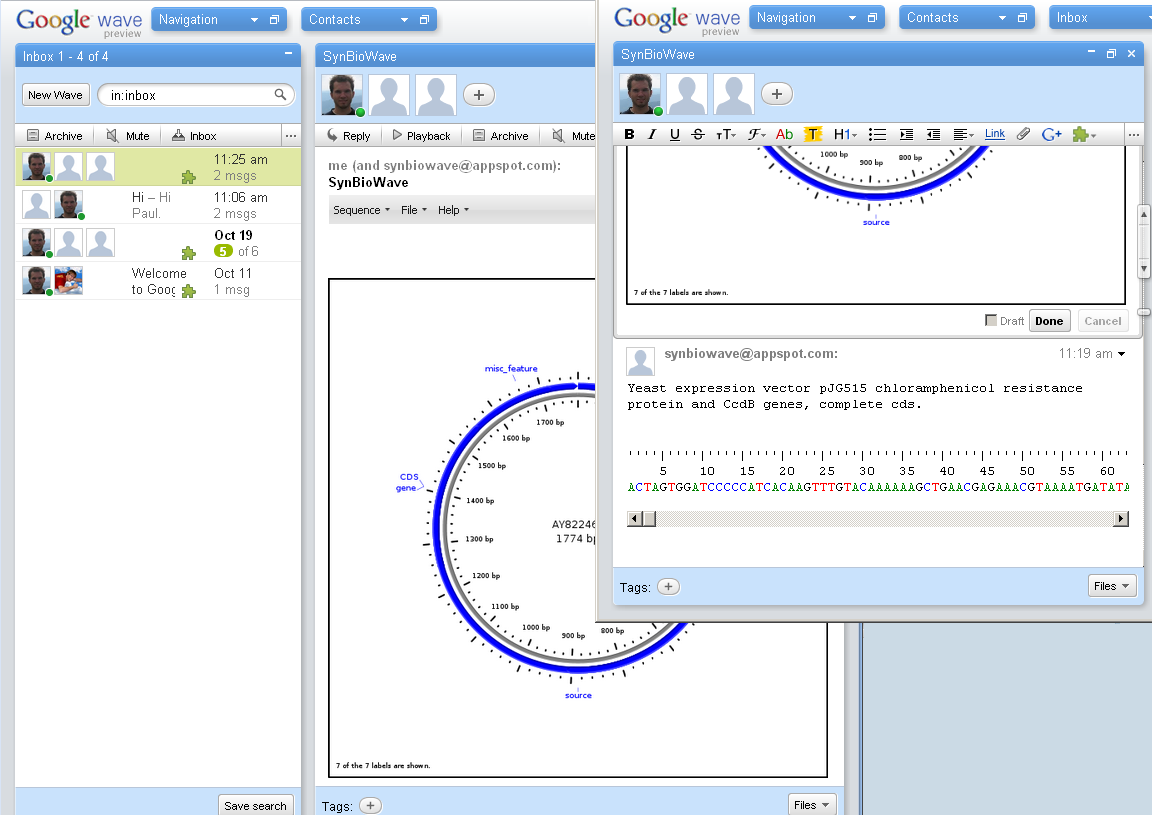
- Rendering and displaying of sequences
- Circular view
- BioBrick communication (searching and importing biological parts)
Additional robots extending SynBioWave functionality
To extend SynBioWave with new biosynthetic functions, one can use additional robots. Adding such a robot to an active SynBioWave wavelet (a wavelet that contains the SynBioWave Robot), enables this function to all users participating in this wave. For SynBioWave users it is very easy to customize and extend SynBioWave. A User searches the robot representing a certain function and adds it to the wavelet he is working with. But this is not the only benefit! This modular architectures makes it very easy for other developers to create additional robots. To facilitate contribution, we provide an abstract SynBioWave Template class. Using this class simplifies additional robot creation a lot. The developer neither needs to worry about Wave development nor about SynBioWave integration (a minimal understanding of both is still needed). There for, one can concentrate on biosynthetic development.
Organizing Robots Work
As we were testing this concept of multiple Robots in one and the same conversation, it pointed out quite clearly that we need to organize the way the robots work: With different Robots all listening to different (or maybe even the same) Events, editing text, writing messages and so on we ended up in greater chaos most of the time. So we decided to extend our Framework to provide a standard in- and output for robots. For the input part, we created a menu, which dynamically displays all the functions of the robots in the current wave. As Google has not foreseen this need of a standardized and easy usable input-method for Wave-robots we had to create this from scratch up, and implemented it by creating a Gadget displaying the menu, a Java-Class Robots can use to create Menus and the so called qooxWave protocol to communicate the menu between the robots and the gadgets. To organize the output of the robots, we created a standardized sequence-display. At the moment the user can choose if he likes at have the sequences either directly in the wave, which feels more "natural" or inside a Gadget, which is currently much more usable because of strict limitations of text-formatting inside Google Wave. Most likely all Sequences will be written directly in the wave in the future as Google has announced to boost die layout options of Wave dramatically.
Gadgets
qooxWave protocol
The qooxWave protocol is introduced to realise an easy to use interface for creating graphical user interfaces (GUI) inside a wave from a robot. The general goal is to provide one abstract robot class for implementing new function into SynBioWave (each function is provided by a robot; see ...). The programmer who uses this class does not need to worry about the client side GUI implementation. This protocol ensures some easy to use server side function, that automatically build the client side GUI.
The protocol defines a a server-to-client and client-to-server communication. This communication contains
- a server to client communication for building client-side GUI elements from server side (so a server side robot can create a client side button for example)
- a client to server communication for reporting events in the client side GUI (so the server knows when the user clicks a button for example)
The communication is realised via JSON-Strings that are stored inside a gadgets state object. Both, server (robot) an client (gadget) can access the state object and they can both react on changes of this state object (this is provided by the google wave API).
Building a GUI - Server to Client Communication
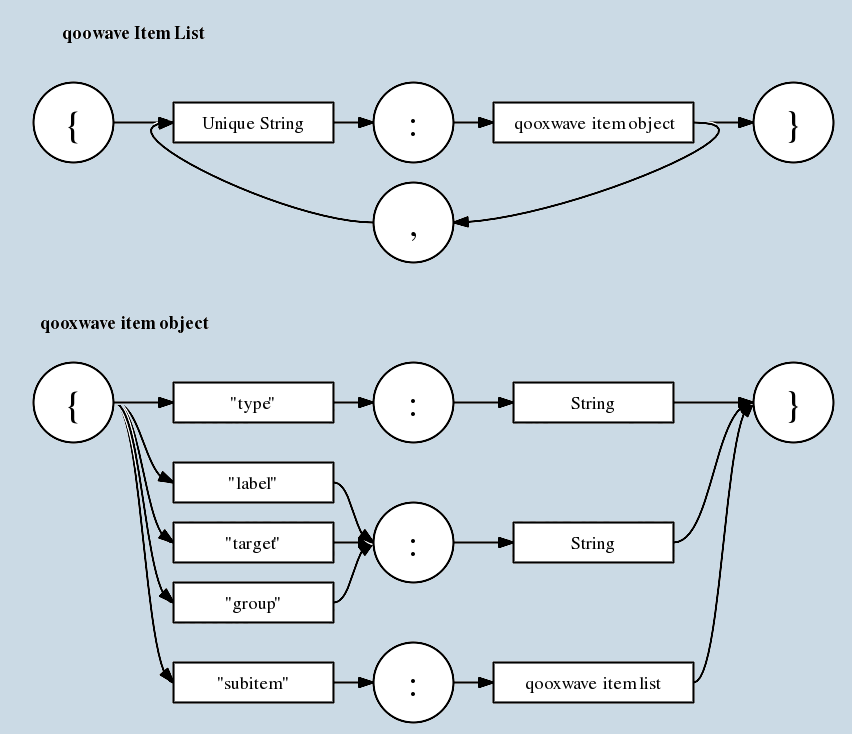
The JSON-object for building a GUI is stored as a String in wave.getState().get("ui.structure") and defined as follows:
{ 'type' : 'ui', 'subitem' : ITEMLIST} where ITEMLIST is associative Array with ids as keys and ITEM objects as values:
{ 'mySubitemId1' : ITEM, 'mySubitemId2' : ITEM, ...}
where ITEM is of the form
{
'label' : STRING <i>label</i>,
'type' : ENUM('toolbar', 'button', 'menu', 'checkbox', 'radio', 'label', 'form') [,
'subitem' : ITEMLIST ]
}
This protocol has a recursive structure. Each item can have an itemlist as subitem. This provides a basis for creating complex menus, forms and ui elements. The type specifies the element. We will extend the available types from time to time. Have a look at Team:Freiburg_software#Item_Structure for the available types and which combinations are allowed.
Item Structure
This section gives an overview of the available types for the items described above. It also depends on the type which other keys can be used for the object and which types are allowed for the subitem list.
- 'type' : 'ui'
- available keys:
- available subitem types: toolbar, menu, button, checkbox, form, label
- 'type' : 'toolbar'
- available keys: subitem
- available subitem types: button, checkbox, menu
- 'type' : 'menu'
- available keys: label, subitem
- available subitem types: button, checkbox, menu
- submits event/data: TODO
- 'type' : 'button'
- available keys: label
- available subitem types: -
- submits event/data: TODO
- 'type' : 'checkbox'
- available keys: label, value (true|false)
- available subitem types: -
- submits event/data: TODO
- 'type' : 'radio'
- available keys: label, value (true|false), group (to group radio items to one radiogroup)
- available subitem types: -
- submits event/data: TODO
- 'type' : 'label'
- available keys: label
- available subitem types: -
- 'type' : 'form'
- available keys: label, subitem
- available subitem types: textfield, textarea, checkbox, radio, upload, download
- submits event/data: many, especially on uploads, downloads | TODO
- 'type' : 'textarea'
- available keys: label, value
- available subitem types: -
- 'type' : 'textfield'
- available keys: label, value
- available subitem types: -
- 'type' : 'upload'
- available keys: label, target
- available subitem types: -
- submits event/data: TODO | submitted, completed
- 'type' : 'download'
- available keys: label, target
- available subitem types: -
- submits event/data: TODO | ?
Building GUI Events - Client to Server Communication
The JSON-object for submitting a GUI-Event from client to server is stored as a String in wave.getState().get("ui.idofelemt") and defined as follows: ...
Example of qooxWave
{'type':'ui','label':'ui','subitem':{'0':{'type':'toolbar','label':'toolbar','subitem':
{'0':{'type':'menu','label':'DNA','subitem':{'0':{'type':'menu','label':'Import','subitem':
{'0':{'type':'button','label':'from File'},'1':{'type':'button','label':'from Database'}}},'1':
{'type':'button','label':'Translate'},'2':{'type':'button','label':'Rev-Complement'}}},'1':
{'type':'menu','label':'Protein','subitem':{'0':{'type':'button','label':'Import'},'1':
{'type':'button','label':'Export'},'2':{'type':'button','label':'Backtranslate'}}},'2':
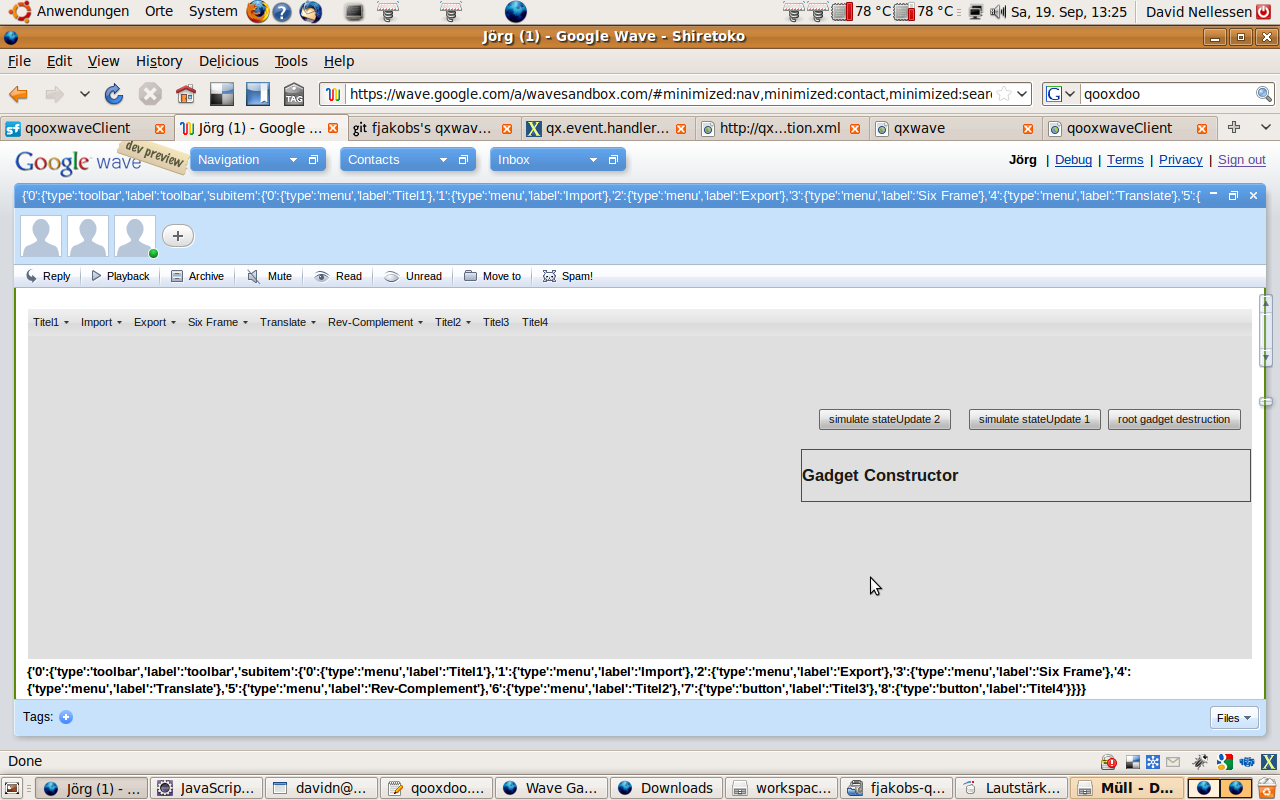
{'type':'button','label':'Titel3'},'3':{'type':'button','label':'Titel4'}}}}}
The above JSON-String creates the following GUI:
Extending google wave I/O
Over the past few decades rapid developments in genomic and other molecular research technologies and developments in information technologies have combined to produce a tremendous amount of information related to molecular biology. This huge amounts of data lays the ground for the work of synthetic biologists. By analyzing,modifying and extending this sequences, the synthetic biologists are enabled to build new functionalities and potentially whole genomes. So one basic feature for the work in synthetic biology is to have fully access to this pool of sequence data, provided in different formats. As only file sharing was intended for google wave, we were forced to extend the build in servlet functionality of the Google Wave Java API with the needs for file import/export as well as methods for database access.
Sequence file import/export
The iGEM Release - Version 0.1
Our current release both proves the concept of SybBioWave
Road-map
Currently we are planing to continue focusing on the improvement of the framework while Google Wave is still a Preview Version and adapt it to the new functions Wave will hopefully get in the future.
Version 0.2
By the time Wave enters a real public and final beta state, we will release Version 0.2 of SynBioWave, which will have a mostly completed, stable and we'll documented Framework and enables developers to create SynBioWave-Robots even more easily than the iGEM-Release.
Improvements planed:
Framework
- Improved Robot-Robot-Communication, both via Wave and direct URL-connections (1)
- Bidirectional Robot-Gadget-Communication (1)
- More simple menu creation in robots i.e. changing the MenuItem-class to an interface and create a class implementation for every available menu item.
- Integrate Callback-functions for menu items
- Improved sequence-display and manipulation, either in a proper-styled(1) Inline-Blip(1) with scrollbar(1) or in a "wave-y" Gadget.
- Improving the Model-View-Controller-Concept of the menu-Gadget
- Heavily improving the usability with lots of testing and feedback from biologist(1)
- Improving the Documentation
BioBrick-Robot
- Integration of Assembly-Algorithms
- Support for different Biobrick-Standarts
- Direct Parts-Upload to the iGEM-Server
Blast-Robot
- Make it do something useful with the received Blast-hits
Other
- Some more Robots as prove of concept and examples for developers
Version 0.3
Versions 0.3 will make SynBioWave attractive to even more Developers by offering them the possibility to write SynBioWave-Robots in Python and further simplifies the creation of Robots with a SynBioWave-Eclipse-plugin. With hopefully many more Robots available at that point, this could be the first release that can be used in labs.
- A Python-implementation of the SynBioWave-Framework
- An Eclipse-Plugin for SynBioWave-Developing
- Many many more Robots
Later Versions
- Support for own Wave- and Robot-Servers (1)
- All functions typically needed in Synthetic Biologie (1)
(1) : currently not supported by Google Wave and/or Google AppEngine, but announced for the future.
 "
"