Team:Todai-Tokyo/Notebook/LDLR
From 2009.igem.org
| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Notebook | Protocols | Ethics |
|---|
Plan
Aim: Create yeast cells that express LDLR on their cell membrane
Methods:
- Clone the LDLR gene.
- Create biobrick of LDLR.
7/6
Overview:
1.PCR LDLR gene from plasmid containing LDLR cDNA
2.gel-purify DNA from the PCR product
3.insert the DNA to vector (iGEM parts)
4.Transform into E. coli and select for ampicillin resistance
5.check for Transformation success using colony PCR by LDLR primers
Constructs to be created:
pGal1-Kozak-LDLR-terminator
Obtaining DNA:
Resuspended DNA in the following wells with 10ul water:
plate 1 7D
pGal1(including Kozak sequence)
[http://partsregistry.org/wiki/index.php?title=Part:BBa_J63006 Gal1 promoter]
Transformed 1ul of each of the above into DH5a competent cells:
- Mix 1ul of DNA with 100ul of competent cells on ice.
- Leave on ice for 30 minutes.
- Heat shock at 42ºC for 45 seconds.
- Leave on ice for 2 minutes.
- Add 500ul of LB and incubate at 37ºC for 1 hour.
- Plate on LB-ampicillin plates.
7/7
Miniprep of E.coli cells containing LDLR gene with Promega, Wizard Plus SVMiniprep DNA Purification System
Successful
7/19
PCR of LDLR
0.4ul 50uM 5’primer
0.4ul 50uM 3’primer
1.6ul 2.5mM dNTP
2ul 10×Pfu Ultra 2 buffer
0.05ul LDLR plasmid
0.5ul Pfu Ultra
15.05ul MilliQ water
↓
Performed PCR using the following program:
1. 95ºC 2min
2. 95ºC 30sec
3. 55ºC 30sec
4. 72.5ºC 60sec
5. repeat 2-4 29times
6. 25ºC forever
PCR successful
We cut the region of LDLR out of the gel and purified PCR product using Promega kit.
infusion of LDLR
4ul PCR product (12.35ug/ml)
3ul vector (iGEM parts plate1 D-3, 14.95ug/ml)
2ul 5×infusion reaction buffer
1ul infusion enzyme
↓
37ºC 15min
50ºC 15min
↓
TE buffer up to 20ul
↓
added 10ul of the sample to DH5α (090614) and put on ice for 15 min.
↓
42ºC 45sec
↓
added 500ul LB broth to the tube.
↓
placed it on LB ampicilin plate.
7/20
colony PCR
put a small amount of single colony into each tube with 5ul MilliQ water
↓
95ºC 5min
↓
PCR reaction
1ul 10×buffer
0.8ul 2.5mMdNTP
0.08ul Ex-Taq
0.1ul 5’-primer
0.1ul 3’-primer
2.92ul MilliQ water
↓
added the PCR reaction to each tube.
↓
Performed PCR using the following program:
1. 95ºC 2min
2. 95ºC 30sec
3. 55ºC 30sec
4. 72.5ºC 120sec
5. repeat 2-4 29 times
6. 25ºC forever
PCR unsuccessful・・・.
7/25
gal1 Sequencing by BIG DYE
1.8ul 5xB.D.3.1.buffer
0.4ul B.D.3.1.
6.3ul MilliQ water
1ul plasmid(0.15ug/ul)
0.5ul 5'or3'primer(3.2pmol/ul)
↓
PCR Program
1.96ºC 2min
2.96ºC 10sec
3.55ºC 5sec
4.60ºC 3min
5.go to 2.29times
6.25ºC forever
↓
add 0.5ul PHOSPHATASE ALKALINE shrimp
↓
37ºC 1hr incubate
↓
add 1ul 3MNaOAc
↓
add 25ul EtOH
↓
20000xg 4ºC 10min centrifugation
↓
put off supernatant
↓
dry tubes
↓
add 15ul HiDi
↓
put them in the sequence machine
7/26
colony PCR again, using 7/20 protocol.
PCR program
1.95ºC 2min
2.95ºC 30sec
3.55ºC 30sec
4.72.5ºC 150sec
5.go to 2. 29times
but,unsucessful.
7/27
Miniprep of E.coli cells containing LDLR gene(that yesterday picked up and cultured) with Promega, Wizard Plus SV Miniprep DNA Purification System
using AGE, found a colony that has LDLR infusion plasmid
7/30
- seaquensing of LDLR
August
PCR of GFP for fusion
program
1.95ºC 2min
2.95ºC 30sec
3.55ºC 30sec
4.72.5ºC 20sec
5.go to 2. 29 times
6.25ºC forever
We found that the length of these PCR product till now was longer than that of LDLR.
- read the Sequence of LDLR cDNA→the result didn't conform with NCBI datebase
- change PCR program→didn't get the PCR product whose length was correct
8/1
8/2
- Sequencing of LDLR
8/4
The following were sequenced using the labeled primers:
- 1)F-spe1-Gall p-LDLR-5
- 2)F-pst1-Gall p-LDLR-3
Results:
Sequence read failed
8/5
- Sequencing of LDLR
8/6
- PCR of LDLR1
- PCR of LDLR2
LDLR1 and LDLR2 is combination of Gal1, LDLR and GFP.
8/7
- PCR of LDLR
using the labeled primers:
- 1)F-spe1-Gal1-LDLR-5'
- 2)F-spe1-Gal1-LDLR-3'
- PCR of GFP
using the labeled primers:
- 1)F-spe1-Gal1-LDLR-GFP-5'
- 2)F-pst1-Gal1-LDLR-GFP-3'
8/8
- Transformation of P3 21D
- Transformation of P1 23L
- PCR of LDLR
using the labeled primers:
- 1)F-spe1-Gal1-LDLR-5'
- 2)F-spe1-Gal1-LDLR-3'
using the following program;
program
1.95ºC 2min
2.95ºC 30sec
3.52~55ºC 30sec
4.72.5ºC 60sec
5.go to 2. 29 times
6.25ºC forever
Result;
The length of LDLR is about 2583 bp.
- PCR of LDLR
using the labeled primers:
- 1)F-spe1-Gal1-LDLR-5'
- 2)F-spe1-Gal1-LDLR-3'
using the following program;
program
1.95ºC 2min
2.95ºC 30sec
3.57ºC 30sec
4.72.5ºC 60sec
5.go to 2. 29 times
6.25ºC forever
The change point of this program is the temperature at third step.
This temperature is 57ºC but previous is 52~55ºC.
result;
The length of LDLR is 2500~3000bp.
8/9
- PCR of LDLR
8/10
- Sequencing of LDLR
The following were sequenced using the labeled primers
1)1D-LDLR-5'
2)1D-LDLR-3'
Results:
Sequence read failed
- PCR of LDLR(1st time)
using the labeled primers:
1)F-spe1-Gal1-LDLR-GFP-5'
2)F-spe1-Gal1-LDLR-GFP-3'
using the following program;
program
1.95ºC 2min
2.95ºC 30sec
3.55ºC 30sec
4.72.5ºC 60sec
5.go to 2. 29 times
6.25ºC forever
Result;
PCR is sucessful.
- PCR of LDLR(2nd time)
using the labeled primers is same as 1st time's:
using the following program;
program
1.95ºC 2min
2.96ºC 30sec
3.55ºC 30sec
4.60.0ºC 3min
5.go to 2. 29 times
6.25ºC forever
8/12
- Sequencing of LDLR
Result;
NNNNN can't read.
- electrophorese LDLR+GFP
RESULT:the band across 3000-3500bp not found. -> need to rePCR LDLR+GFP.
8/13
- measure the absorbance of LDLR2cDNA
->534ug/mL
+PCR
objects:LDLR2, yqiT, GFP
program
1.95ºC 2min
2.96ºC 30sec
3.55ºC 30sec
4.60.0ºC 3min
5.go to 2. 29 times
6.25ºC forever
- electrophorese the above 3
result -> other than GFP not found <- maybe DNAs dissolved by UV or contamination;
8/14
- PCR
objects: LDLR GFP
LDLR
primer
F_S_GalI_LDLR_5'
F_PstI_GalI_LDLR_3'
GFP
primer
F_S_GalI_LDLR_GFP 5'
F_P_GalI_LDLR_GFP 3'
- electrophorese PCR products
->failed (LDLR+GFP)
8/15
rePCR LDLR+GFP electrophorese LDLR+GFP
8/19
1 Colony PCR of LDLR
Using the labeled primers:
1) J63010(P17D)_seq_5’
2) J63010(P17D)_seq_3’
Result:
successful
2 PCR of LDLR for fusion
Using the labeled primers:
1) F_S_Gal1_LDLR_5’new
2)F_LDLR_3’new
3 PCR of GFP for fusion
Using the labeled primers:
1) F_S_Gal1_LDLR_GFP_5’
2) F_S_Gsl1_LDLR_GFP_3’
8/21
1 PCR of LDLR
Using the labeled primers:
1) F_S_Gal1_LDLR_5’new
2)F_LDLR_3’
2 infusion of LDLR and GFP
8/22
1 Miniprep ofLDLR
8/23
1 Sequencing of LDLR
Using the labeled primers:
1) J_P17D_5’
2) J_P17D_3’
8/25
1 PCR of LDLR
Using the labeled primers:
1) F_S_Gp_LDLR_out_5’
2) F_P_Gp_LDLR_out_3’
8/26
1 PCR of LDLR
Using the labeled primers:
1) F_S_Gp_LDLR_out_5’
2) F_P_Gp_LDLR_out_3’
2 Sequencing of LDLR cDNA
Using the labeled primers:
1) F_S_Gp_LDLR_out_5’
2)F_P_Gp_LDLR_out_3’
Result : failed
8/27
1 PCR of LDLR
Using the labeled primers:
1) F_S_Gp_LDLR_out_5’
2) F_P_Gp_LDLR_out_3'
2 Sequencing of LDLR cDNA
Using the labeled primers:
1) F_S_Gp_LDLR_out_5’
2)F_P_Gp_LDLR_out_3’
Result : failed
8/30
- PCR of LDLR
using the labeled primers:
1)F-spe1-Gal1-LDLR-GFP-5'
2)F-spe1-Gal1-LDLR-GFP-3'
8/31
Sequencing of LDLR
The following were sequenced using the labeled primers
1)1D-LDLR-5'
2)1D-LDLR-3'
Results:
Sequence read failed
September
We got new LDLR cDNA!
- look for the best PCR program
→DMSO 5% or 10%
→annealing temparature 55ºC or 60ºC
result
the best program
10% DMSO
1.95ºC 2min
2.95ºC 30sec
3.60ºC 30sec
4.72.5ºC 1min
5.go to 2. 29 times
6.25ºC forever
purify the PCR product using Promega kit and insert it in iGEM part(plate1-7D:Gal1 promoter)
And, insert it and GFP in iGEM part(plate1-7D:Gal1 promoter)
9/15
1. PCR + Gel Extraction
- LDLR
2. PCR
- LDLR (annealing temperature:57)
- LDLR (annealing temperature:60)
3. PCR
- LDLR: tested different concentrations of template (do not contain DMSO)
- 1. X 1/10
- 2. X 1/5
- 3. X 1 (normal)
- 4. X 5
- 5. X 10
9/16
1. PCR
- LDLR: tested different concentrations of template
- 1. X 1/10 + 20%DMSO
- 2. X 1/5 + 20%DMSO
- 3. X 1 (normal) + 20%DMSO
- 4. X 5 + 20%DMSO
- 5. X 10 + 20%DMSO
2. PCR
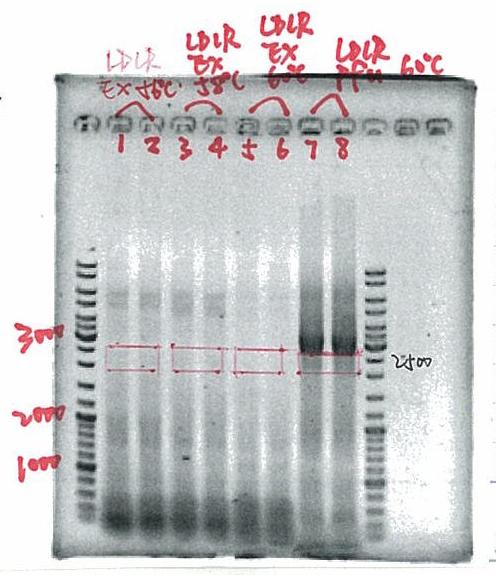
- LDLR: tested different conditions
- 1. cDNA1, old primer, 20% DMSO
- 2. cDNA1, old primer, no DMSO
- 3. cDNA1, new primer, 20% DMSO
- 4. cDNA1, new primer, no DMSO
- 5. cDNA2, old primer, 20% DMSO
- 6. cDNA2, old primer, no DMSO
- 7. cDNA2, new primer, 20% DMSO
- 8. cDNA2, new primer, no DMSO
- 9. control
9/24
1. PCR
- GEP
- YCplac111
9/25
- Miniprep of E.coli cells containing GFP gene (those yesterday picked up and cultured) with Promega, Wizard Plus SV Miniprep DNA Purification System
- read the Sequencing of LDLR cDNA
Results:
Sequence read failed
- PCR of LDLR cDNA(Pfu UltraII)
Results:
PCR failed
9/26
- PCR of LDLR for infusion
Using the labeled primers:
1) F_S_Gal1_LDLR_5’new
2)F_pstI_Gal1p_LDLR_3’new
Results:
PCR failed
- PCR of LDLR for fusion
Using the labeled primers:
1) F_S_Gal1_LDLR_5’new
2)F_LDLR_3’new
Results:
PCR failed
- PCR of GFP for fusion
Using the labeled primers:
1) F_S_Gal1_LDLR_GFP_5’
2) F_p_Gal1_LDLR_GFP_3’
Results:
PCR failed
9/27
- PCR of LDLR
Using the labeled primers:
1) F_S_Gp_LDLRout_5'
2)F_P_Gp_LDLRout_3'
Results:
PCR failed
- PCR of LDLR
Using the labeled primers:
1) F_S_Gal1_LDLR_5'new
2)F_P_Gal1p_LDLR_3'
Results:
PCR failed
- PCR of LDLR
Using the labeled primers:
1) F_S_Gal1_LDLR_5'new
2)F_LDLR_3'
Results:
PCR failed
- read the Sequencing of LDLR cDNA
Results:
Sequence read failed
10/5
- check of plasmids including LDLR by AGE
1.LDLR3 091004
2.LDLR5 091004
3.LDLR+GFP 1 091004
4.LDLR+GFP 2 091004
1&2->OK,3&4->wouldn't have LDLR+GFP
1.LDLR3 091004
2.LDLR5 091004
3.LDLR+GFP 1 091004
4.LDLR+GFP 2 091004
5~10.YCplacIII(whose restriction site may be changed)
primer
1~4:P1-7Dseq.5'or3'
5~10:YCplacIII seq. primer5'or3'
- PCR for LDLR+GFP fusion(Pfu UltraII)
template:HEP-LDLR cDNA
primer:
F_S_Gal1_LDLR_5'new
F_P_GAl1_LDLR_GFP_5'
10/7
- PCR for GFP fusion(Pfu UltraII)
primer
F_S_Gal1_LDLR_GFP_5'
F_P_Gal1_LDLR_GFP_3'
10/8
1. PCR of Gal1+LDLR
P1-7D seq5'
P1-7D seq3'
(Pfu ULtraII)
10/10
- YCplacIII quick change
PCR
YCplacIII5'
YCplacIII3'
10/11
- PCR
template:HEP-LDLR cDNA
primer:
F_S_Gal1_LDLR_5'new
3'-GFP
LDLR5
LDLR3
10/12
1. sequencing
- LDLR
- P2,10F
- YCplac111
- Gal1
2. PCR LDLR from HEP
10/13
1. PCR
- LDLR + GFP
- LDLR
10/15
1. Sequencing
- LDLR
10/16
1. PCR
- LDLR
- YCplac111
2. Restriction Enzyme digestion
- YCplac111 with E,P
- YCplac111 with X,P
3. Colony PCR
- LDLR + GFP
- LDLR + P1,7D
10/18~
1. Sequencing
- LDLR
2. PCR of LDLR
tested the following conditions;
- 1. DMSO 10%, 60ºC(for annealing)
- 2. DMSO 15%, 55ºC
- 3. DMSO 15%, 60ºC
- 4. DMSO 20%, 55ºC
- 5. DMSO 20%, 60ºC
- 6. DMSO 10%, 58ºC
3. Ligation
- YcplacIII + LDLR
4. PCR of LDLR
tested the following conditions;
- 1. DMSO 10%, 60ºC(for annealing)
- 2. DMSO 15%, 55ºC
- 3. DMSO 15%, 60ºC
- 4. DMSO 20%, 55ºC
- 5. DMSO 20%, 60ºC
- 6. DMSO 10%, 58ºC
5. PCR of LDLR
tested the following conditions;
- 1. DMSO 10%, 60ºC(for annealing)
- 2. DMSO 15%, 55ºC
- 3. DMSO 15%, 60ºC
- 4. DMSO 20%, 55ºC
- 5. DMSO 20%, 60ºC
- 6. DMSO 10%, 58ºC
- 7. 40ul system, DMSO 10%, different primer (YcplacIII)
6. Gel electrophoresis
- lane 2~15; LDLR
7. sequencing
- LDLR+P1 7D
8. Ligation
- LDLR and YcplacIII
| Home | The Team | The Project | Parts Submitted to the Registry | Modeling | Notebook | Protocols | Ethics |
|---|
 "
"