|
|
Contents
- 1 Opening Remark
- 2 Calendar and Events
- 3 Work Distribution
- 3.1 Overall Construction (Wei Pan, Bo Ding)
- 3.2 G/LSA and PSO (Yuwei Cui)
- 3.3 GUI (Xiaomo Yao)
- 3.3.1 Jun. 15, 2009
- 3.3.2 Jul. 26, 2009
- 3.3.3 Jul. 27, 2009
- 3.3.4 Aug. 4, 2009
- 3.3.5 Aug. 13, 2009
- 3.3.6 Aug. 22, 2009
- 3.3.7 Aug. 26, 2009
- 3.3.8 Aug. 27, 2009
- 3.3.9 Aug. 31, 2009
- 3.3.10 Oct. 5, 2009
- 3.3.11 Oct. 16, 2009
- 3.4 I/O SBML (Jiahao Li)
- 3.5 I/O Database and Miscellaneous(Yu He)
- 3.5.1 Apr. 3, 2009
- 3.5.2 Apr. 15, 2009
- 3.5.3 May 15, 2009
- 3.5.4 May 20, 2009
- 3.5.5 Jul. 15, 2009
- 3.5.6 Aug. 5, 2009
- 3.5.7 Sept. 1, 2009
- 3.5.8 Sept. 3, 2009
- 3.5.9 Sept. 6, 2009
- 3.5.10 Sept. 20, 2009
- 3.5.11 Sept. 28, 2009
- 3.5.12 Oct. 8, 2009
- 4 Future Plan
- 5 Closure - End or And?
|
It's time to move from paper to website. We might recall the ever first plan and ambition when we first set feet on the software designing track. The following documentations record word by word the way how we are here.
Calendar and Events
Group Meetings: Slides and Records
- Feb. 11, 2009
- [http://home.ustc.edu.cn/~heyu3/090211.txt (abstract)]
- Feb. 12, 2009
- [http://home.ustc.edu.cn/~heyu3/090212.txt (abstract)]
- Feb. 21, 2009
- [http://home.ustc.edu.cn/~heyu3/090221.txt (abstract)]
- [http://home.ustc.edu.cn/~heyu3/BioCADthought.ppt (slides)]
- Feb. 28, 2009
- [http://home.ustc.edu.cn/~heyu3/Virtual_E._coli.ppt (slides)]
- Mar. 7, 2009
- [http://home.ustc.edu.cn/~heyu3/WholecellSimulation.pdf (slides)]
- Mar. 15, 2009
- [http://home.ustc.edu.cn/~heyu3/Dry_Week_4.txt (abstract)]
- [http://home.ustc.edu.cn/~heyu3/Week4.pptx (slides)]
- Mar. 29, 2009
- [http://home.ustc.edu.cn/~heyu3/Onaviewpointofstochasticprocess.pdf (slides)]
- [http://home.ustc.edu.cn/~heyu3/info-week6.pdf (slides)]
- Apr. 8, 2009
- [http://home.ustc.edu.cn/~heyu3/090408.txt (abstract)]
- Apr. 18, 2009
- [http://home.ustc.edu.cn/~heyu3/Software.ppt (slides)]
- Apr. 21, 2009
- [http://home.ustc.edu.cn/~heyu3/Simuweek7.pptx (slides)]
- Apr. 24, 2009
- [http://home.ustc.edu.cn/~heyu3/FlowChart.pptx (slides)]
- May 16, 2009
- [http://home.ustc.edu.cn/~heyu3/090516.pptx (slides)]
- May 22, 2009
- [http://home.ustc.edu.cn/~heyu3/SBML_zqliu.ppt (slides)]
|
Work Distribution
Overall Construction (Wei Pan, Bo Ding)
Week 2
“What can we do for the experimentalists?” I saw my team members. It’s the third time for me to take part in iGEM. Interestingly, in this time I am the only person with the background of biology, while nearly everyone’s background is solo. It is really a mixture. I hope there will be chemical reactions among us.
“Design”, the key word floated in everyone’s mind during the discussion. The network inference from the microarray expression profile, for protein concentration profiles, from promoter sequence data, etc, can be done in the DREAM (Database for Reverse Engineering Analysis and Method) project. Meanwhile, the network design can also be done to the required profile. So we will start from here!
So the aims for our project:
1.Present a structure frame for network design
2.Redesign the known networks to test the strategy
3.In silico, design a network with a interesting behavior
4.Realize the network in the cell
And the problems are really hard for our projects. Actually, everything we want to do is we have never done before. So we will try our best.
Week 4
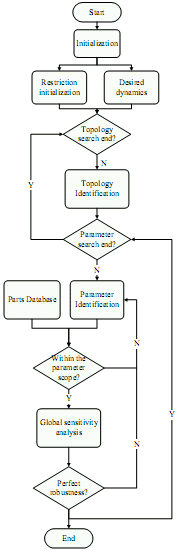
The problems we faced may be harder than our imagination. First, the search space is too large to traverse. The number of potential topologies is , while K is the number of interaction forms and n is the number of species. It is an astronomy number if n >=3. Second, the non-linear system identification is still an unsolved problem in the present. The parameters can not be recognized exactly even the topologies are meeting the requirements. Some approximation should be done mathematically. First, the networks we designed are restricted to the simple network, which means the reactors are at most two kinds in one reaction. The searching space can be shrunk in this way, while all the dynamic behaviors can be simulated with this simple model. Second, we may need some restrictions to the interactions forms and the parameters. The more information is, the better system we can get by the design process.
Week 6
The frame of the software has been constructed as indicated in right figure.
week 9
From the analysis, we should get the information of a reaction from three aspects: topology, formula and the parameters. The optimized algorithms are required with the three different layers problems. For the bottom layer, the parameters are optimized with the known forms of formula. For the upper layer, we need genetic algorithm to look for the formula forms that meet the requirement. And for the top layer, genetic algorithm is also employed to optimize the topology. And I will work with the selection of the formula and the topology.
week 16
The genetic algorithm has been implemented with c++ and tested with some benchmarks. Next, we discussed about the data structures for the program, including the input form and input data, the output form and output data, the description of a reaction, and the items considering in formula. According to the discussion results, I began to implement the whole frame for the program. There are three classes in the program.
week 20
The frame of the program has been implemented. The oscillator system is tested.
week 22
information is integrated. And the adaptation system is tested
week 23
PSO Algorithm is proposed and implemented.
week 25
The multiple initial values system identification is integrated. And the toggle switch system is tested.
week 27
Wikiwikiwiki~ Fast learning needed to digest so much info...
|
G/LSA and PSO (Yuwei Cui)
Feb. 12, 2009
|
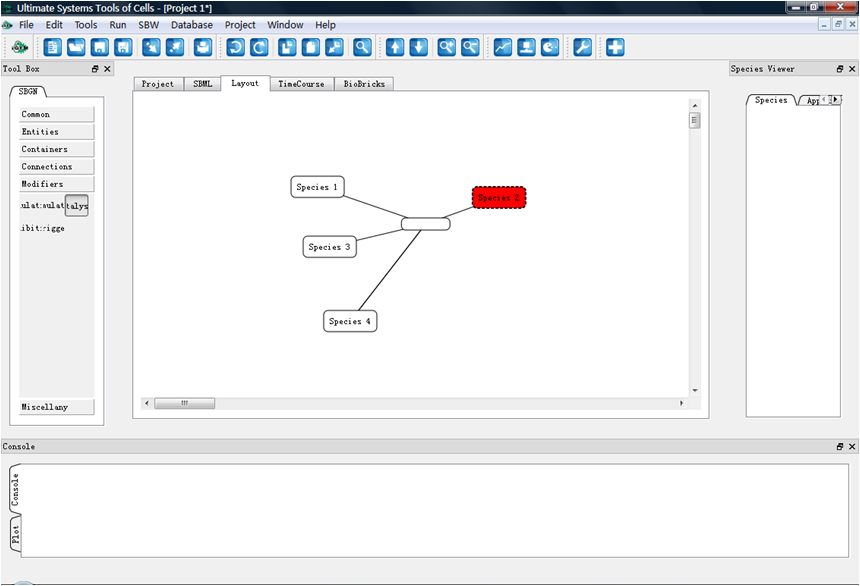
GUI (Xiaomo Yao)
Jun. 15, 2009
|
Start GUI design
|
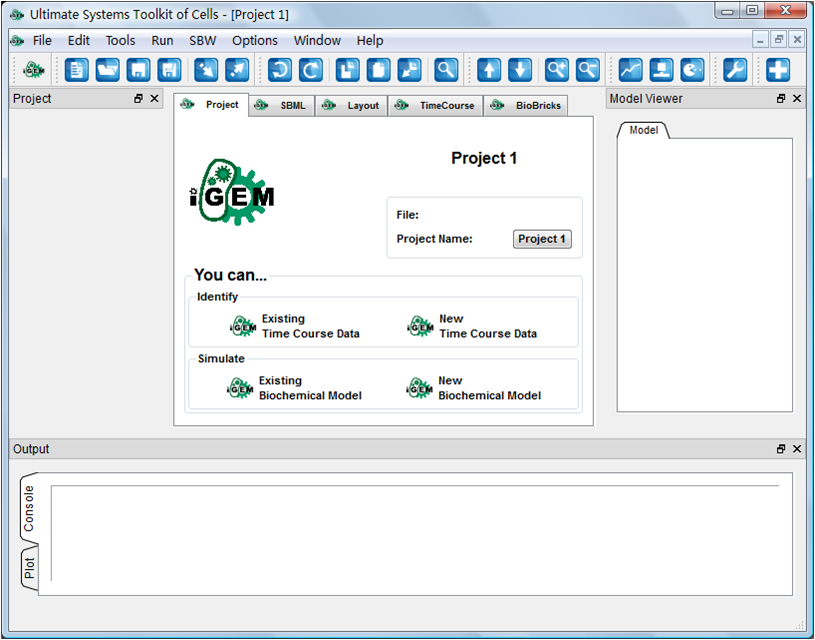
Jul. 26, 2009
Start GUI project
Vedio Commu.
|
Jul. 27, 2009
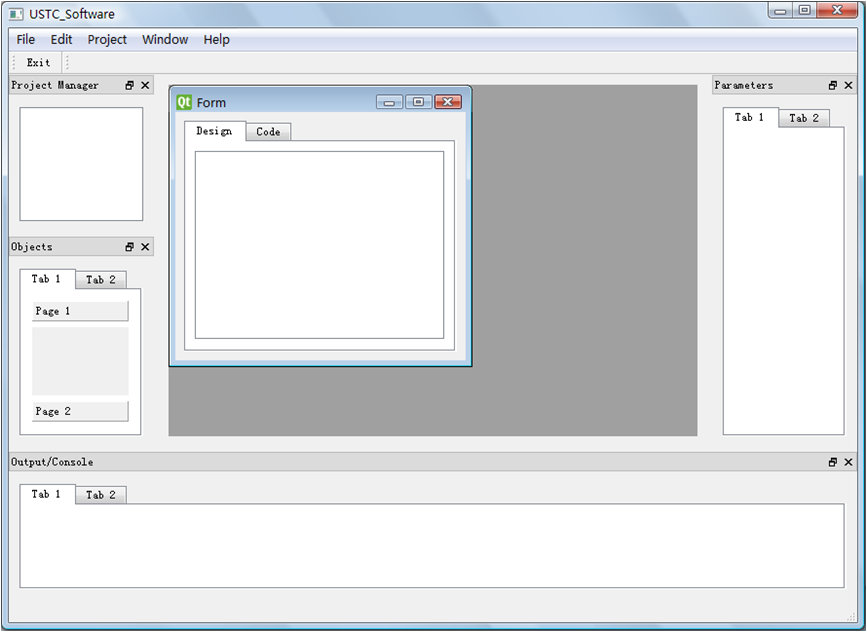
Form created, with MDI & dock windows
GUI version 090727 released
|
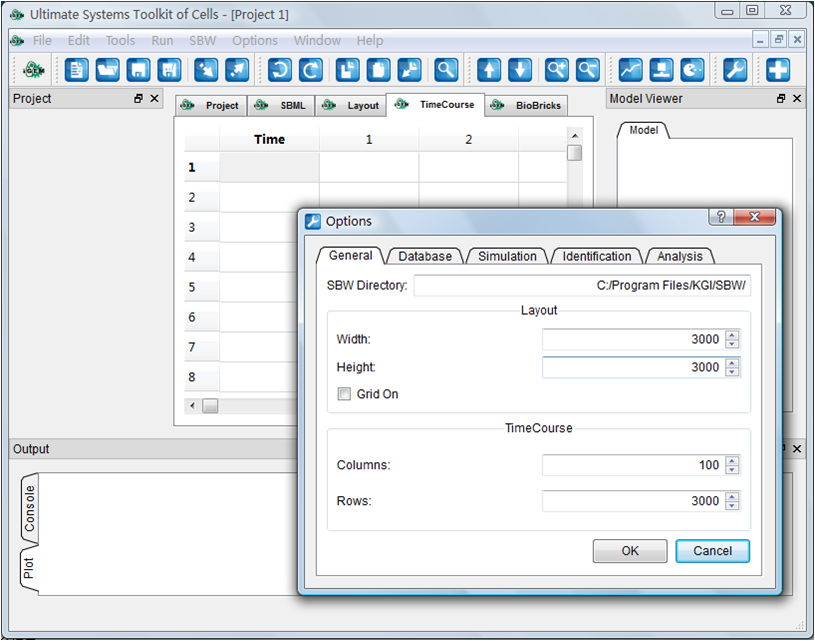
Aug. 4, 2009
XML editor finished
|
Aug. 13, 2009
Start developing layout drawing canvas
|
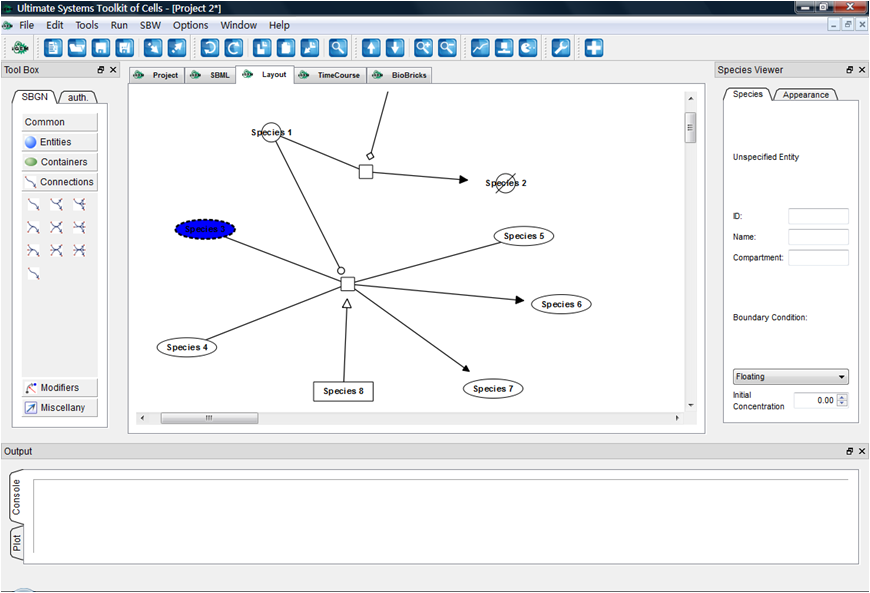
Aug. 22, 2009
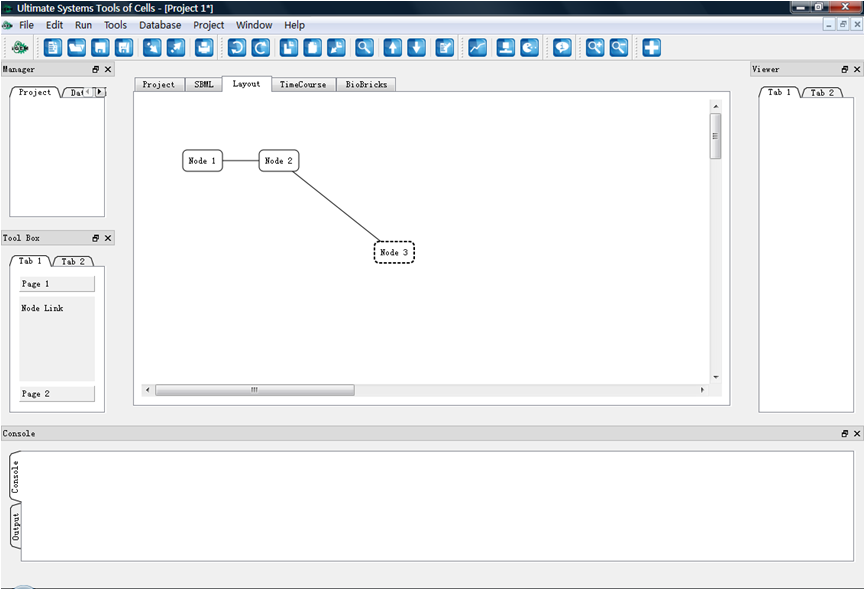
Layout drawing canvas developed; Reactions and modifiers enabled
|
Aug. 26, 2009
Layout drawing canvas completed
|
Aug. 27, 2009
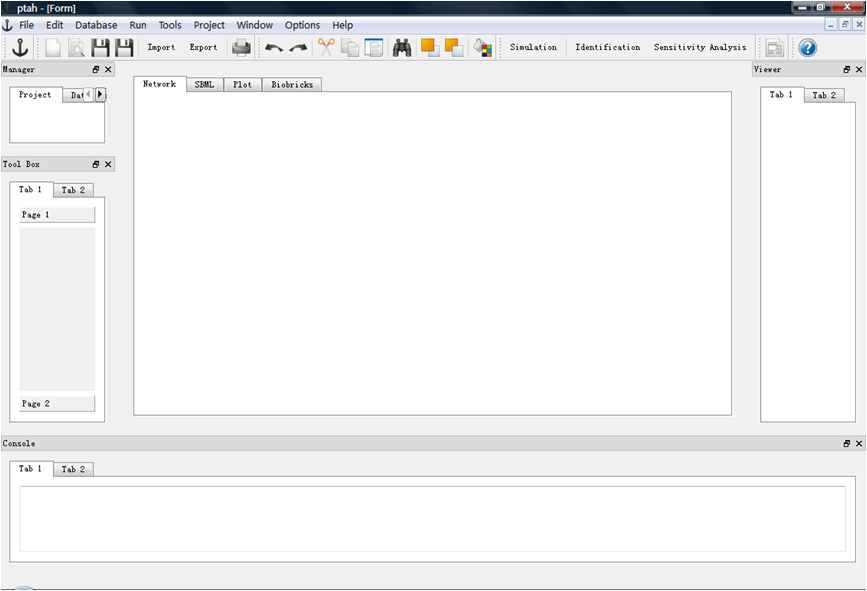
GUI version 090827 released
|
Aug. 31, 2009
Start programming interface with identification/analysis algorithms
|
Oct. 5, 2009
Interface developed
|
Oct. 16, 2009
Interface completed
GUI version 091017 released
|
|
I/O SBML (Jiahao Li)
Feb. 12, 2009
No matter what kind of software we will make, it is sure that one of the most important work is to find out the structure and functions of existing software. Preparing start from Athena and Tinkercell, two multi-function synthetic biological software.
OMG!! these software are great! They can do not only some simulating works, but also other create job! Their GUI also performs well. It seems hard to beyond or even achieve them in simulating functions. We have to find other highlights.
|
Mar. 14, 2009
Be aware of SBML, a standard structured file contains a certain chemical/biological system. The support of SBML related functions in Athena; Tinkercell and other software convince me the necessity of providing these functions in our software. It means our output can supply a standard interface to others.
May. 2, 2009
Start learning SBML and LibSBML. Well~~ download, unzip, install, OMG!! Quite a lot of codes, my teammates advise me to read the tutorial first. Yes, after reading, I realize the structure of SBML file and working process of LibSBML approximately. In terms of our software, some basic information of the SBML file is needed, such as amount of spceies and parameters, their names, information of each reaction, including the mathematical expressions of each kineticlaw. All these data need to extract from given SBML file. So it looks learning to use LibSBML is inevitable.
Jun. 6, 2009
Go on working with LibSBML, Successfully output the species names, parameters and other informations.
Need to face some problems when deal with the mathematical expressions, SBML uses ASTNode to store the formula, although it is not difficult to manage the "tree" data structure, I still need to make clear the functions in "class ASTNode".
My teammates need me to supply the information orderly, which means :
1. The information of one reaction should be stored by categories.
2. The terms(the formula related to one certain species) should be put together.
3. The whole information need to be packaged for next step.
Aug. 15, 2009
The readSBMLFile part finished, in this part, I pick each species list in the file orderly, and search all reactions, if the species appeared in "ListOfReactants"; "ListOfProducts" or "ListOfModifiers", i'll push the informations of the reaction into the vertor<term>, the data structure of vector<term> has been decided as "class val_func". After the traversing, i'll package these vectors<term>, and sent to the next step.
Aug. 31, 2009
The writeSBMLFile part finished, the experience of making read part make me facilitate to assemble this part. Besides some normal steps, I use "traversing binary tree" to make the mathematical expression fit each certain reaction. But there also some problems with open files from database part, I still need to work on this~~
Sep. 2, 2009
Solving the problems in writing part, the validating part work successfully.
I/O Database and Miscellaneous(Yu He)
Apr. 3, 2009
|
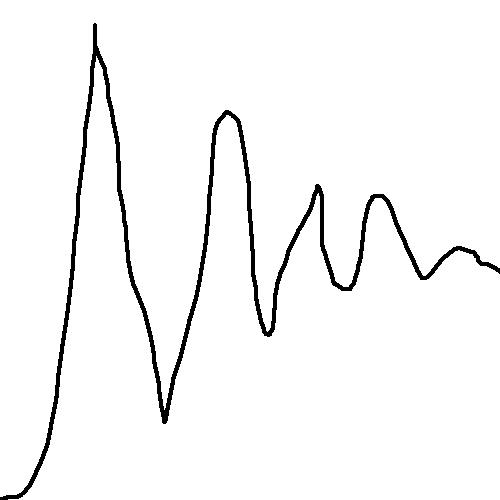
Release of bmptodat plug-in, in order to realize 'desire in, solution out'
Source Code File:FuncExtractor.cpp
Function: Extract function info from a .bmp file
Input: .bmp file path and name (less than 200 bytes) | .dat file path and name (less than 200 bytes)
Output: .dat file with range x[0,1], y[0,1], and step length of 0.01 (alterable).
Demo:
|
|
1.Draw the picture D:\Activities\iGEM\Codes\weigh.bmp like this:
|
|
2.Input like this:
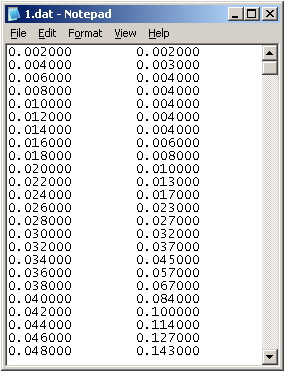
3.Get a .dat file at under folder D:\Activities\iGEM\Codes, named weigh.dat like this:
|
|
|
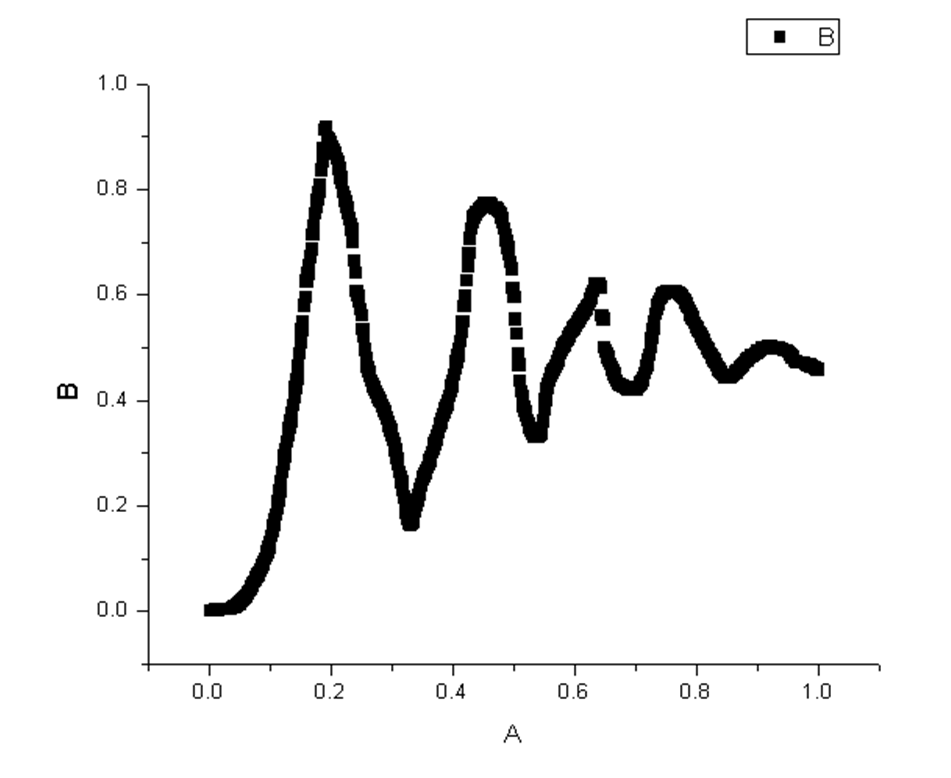
4.Testify the output weigh.dat file by Origin8.0:
|
|
To be improved:
1.Rescale-able range of x-axis and y-axis
2.User designated step length
3.Function of ‘smoothing’
|
Apr. 15, 2009
Release of matrixtoode plug-in
Source Code File:Matrix_ODEArray_conv_1.0.cpp
Function:Convert a interaction-style matrix into text-ified ODE Array
Input: a coefficient matrix through console window
Output: .txt file describing the ODE Array in matlab readable text.
Demo:
|
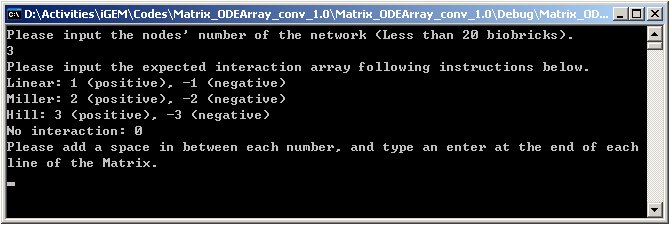
1.Input the coefficient matrix like this:
|
|
2.Output completed like this:
|
|
Note: Depreciated later on.
|
May 15, 2009
Today the core algorithm of reverse engineering is proposed. We discussed possible approaches to realize these system identification requirements.
The very first thing emerge in my mind is Metropolis method and Monte Carol sampling. If we define an error function which serves as 'judging criteria', and a random number generator used for 'correction' or 'exploration', then we can basically get it running from pure theoretical view.
May 20, 2009
Fourth order multi-dimensional Runge Kutta Method is realized. Tested also on Mathematica7.0 to precision of 6th. digits.
Jul. 15, 2009
Hooray!!! Raw program of system identification on an colony-evolution system, whose behavior is damped oscillation realized.(3 functions, 6 parameters)
Aug. 5, 2009
Raw simulation of an oscillation example on substential level. (6 functions, 5 parameters, nonlinear model).
Sept. 1, 2009
Realizes multiple (13 functions, 11 parameters) targets identification with basic metropilis method and gradient decrease method. Yet, it is still a linear func array.
Sept. 3, 2009
Database in development. It's really a challenge to adopt SQLite now. I don't have that much confidence, since there's little time left.
Sept. 6, 2009
Today I finally make up my mind to realize the database from 'ab-initio' on. Since the schedule is so tight, I still feel safer to work with familiar C language.
Sept. 20, 2009
Okay, the very basic, essential, elementary database in .dat form is finished...
Sept. 28, 2009
Inter-database search and sort function completed.
Oct. 8, 2009
Thanks to Grapeot's timely suggestion and help. The new version of bmptodat plug-in is successfully upgraded by taking color-depth into consideration. This eliminates the x-axis 'triple folding' bug due to differnt color-depth of 8 bits and 24 bits.
|
Future Plan
Our detailed future plan about ABCD Suite is discussed in form of FAQ HERE.
Closure - End or And?
We hold in hands the same motivation to adventure, unfold and appreciate the secret of life via the way of virtual evolution and simulation. The seven of us major from automation to engineering, mathematics to physics, grading from freshman to PhD candidates. Though there had been hard times, we have a faith. More hopefully, we’d express our gratitude to the kind support from the School of Life Science of USTC, which makes all impossible possible.
From the one-month brainstorm we collected our first proposal – construct virtual bacteria. Yet, for some practical reasons we then shifted to a second proposal, which narrowed to molecular level simulation and later on turned out to be the prototype of our present project.
From the one-month brainstorm we collected our first proposal – construct virtual bacteria. Yet, for some practical reasons we then shifted to a second proposal, which narrowed to molecular level simulation and later on turned out to be the prototype of our present project.
What do we desire to realize? In short, just tell us what you want your bio-device to behave, and we will return you with a list of eligible formation strategies for your design, of course, with bio-bricks.
Turgidly as the idea might appear to be, this is basically an adoption of the gist of reverse engineering and evolution. By intake the custom-designated behaviors, we search, sometimes traverse, the solution space formed by nearly-inexhaustible combination of Biobricks. In order to make the simulative process applicable and practical, we employed a couple of algorithms widely used in computational sciences like Metropolis method and Dijkstra Algorithm to cut down the time cost and optimize the final result. Analysis on sensitivity and robustness has also been carried out to escort a reliable output of the final list of Biobrick combinations.
Now we have an ABCD for reconstruction of bio-network, or rather, all ODE systems theoretically. But there still exist many more defects and more inspirations on a higher level. Therefore, instead of fixing 'The End' to the last line of our notebook, we'd rather define another 'and...' clause, claiming another start for greater good.
See ya iGEM2010!
|
 Automatic Biological Circuits Design - Team Logo: wanna know more about the hinding metaphors and inspirations in this little red square? Click to check out how much fun this year's iGEM has brought us!
|
|
 Teaching Affair Office, USTC
Teaching Affair Office, USTC
|
|
 School of Life Sicences, USTC
School of Life Sicences, USTC
|
|

Foreign Affair Office, USTC
|
|

Graduate School, USTC
|
|
 School of Information Science and Technology, USTC
School of Information Science and Technology, USTC
|
|
 School for the Gifted Young, USTC
School for the Gifted Young, USTC
|
|

|
 "
"