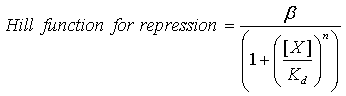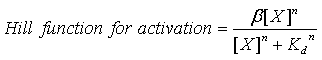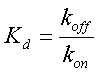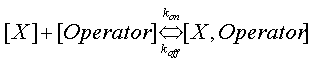Team:Aberdeen Scotland/parameters/invest 1
From 2009.igem.org
Revision as of 10:46, 7 August 2009 by Nick Smart (Talk | contribs)
University of Aberdeen - Pico Plumber
iGEM 2009
Dissociation Constants
Introduction
Our model uses hill kinetics; we have three repression hill functions of the form:
It also has one activation hill function of the form:
And one repression / induction hill function of the form
Where β is the maximal transcription rate, [X] is the concentration of protein X and Kd is the dissociation constant for molecule X to the operator in question, [S] is the concentration of the inducer, S and Ks is the dissociation constant for the inducer to the repressor, X. Kd is defined as follows.
| Parameter | Value | Value (molecules per cell) | Description |
| KLacI | 0.1 - 1 [pM] OR 800 [nM] | 0.00004-0.0004 molecules OR 322 molecules | LacI repressor dissociation constant |
| KIPTG | 1.3 [µM] | 522 molecules | IPTG-LacI repressor dissociation constant |
| KtetR | 179 [pM] | 0.07 molecules | TetR repressor dissociation constant |
| KCI | 8 [pM] OR 50 [nM] | 0.003 molecules OR 20 molecules | CI repressor dissociation constant |
| KAHL | 0.09 - 1 [µM] | 402 molecules | AHL-LuxR activator dissociation constant |
 "
"