Team:Imperial College London/Drylab/Protein Production
From 2009.igem.org

Contents |
Introduction
This module consists of producing the drug protein of interest. In order to describe the function of the module, two models have been developed.
Model for protein drug production:
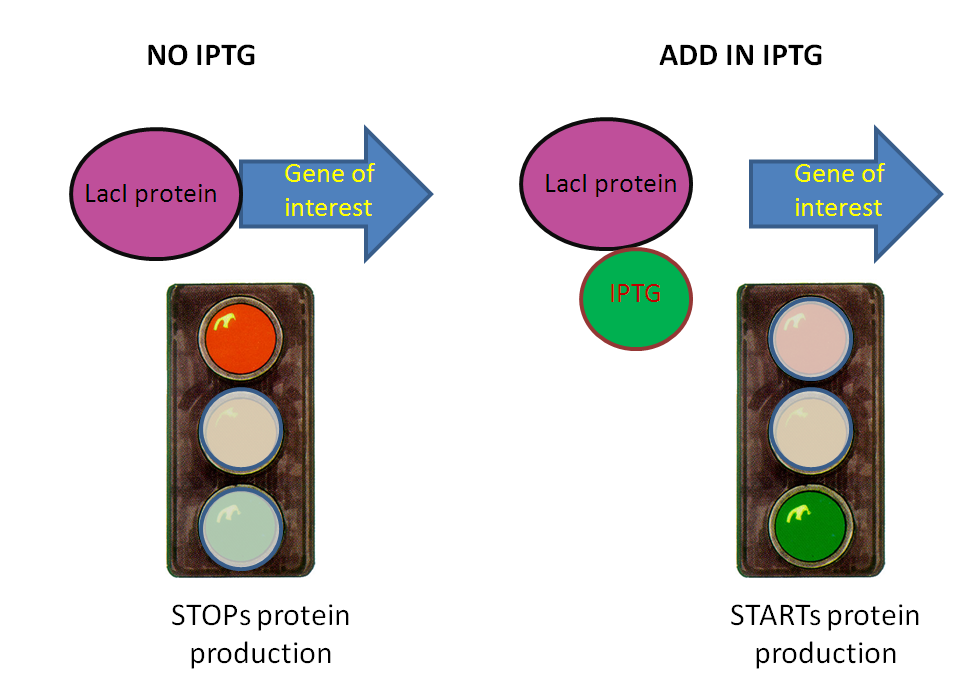
Based on the Genetic circuit, a LacI-IPTG inducible promoter is responsible for kickstarting the production of the drug.
- In the absence of IPTG, LacI represses the production of the drug (Cellulase or PAH)
- When IPTG is introduced, the LacI repressing pathway is “de-repressed”, and some output protein is produced.
Our goals
The modelling aims to provide an overview and better understanding of the M1 system’s function by:
- Characterizing the system.
- Modeling to account for several factors that may reduce/hinder the production of the protein drug such as:
- Lac promoter leakiness
- IPTG toxicity
- Stability of output protein
This module is an integral part of the design, as large-scale commercialization of the drug of interest depends on finding the optimal conditions for protein production. This model has also helped the Wetlab to plan some of the experiments.
More information on the model can be found in the Drylab hub
Model for drug protein enzyme kinetics
The protein drug of interest is an enzyme. It will bind to specific substrates and increase the rate of their conversion into products. Therefore, by monitoring either the substrate concentration or the product concentration, we can indirectly see the activity of the enzyme. This is quantified by measuring by enzyme activity in the cellulase assay and PAH assay. Enzyme activity is the rate of substrate utilisation /product formation per unit time.
Our Goals
This model aims to better understand the enzymatic action of the drug by:
- Characterizing its enzymatic activity
- Subsequently,by modeling the relationship between the quantity of protein being produced and the enzyme activity using a simple Michaelis-Menten enzyme kinetics model will enable us to take into account factors that could impair enzymatic activity
This model will allow an understanding of the activity of the enzymes and allow the WETLAB to have an idea of the magnitude of the enzymatic activity. More importantly, after the results from enzyme assays are obtained, the model should provide a means of relating the activity output data to the concentration of the enzyme.
More information on the model can be found in the Drylab hub
 "
"