Team:Aberdeen Scotland/WetLab/andgate/cloning
From 2009.igem.org
| Line 5: | Line 5: | ||
As our individual modules would be ideally combined at a later point, we had to make sure that our final vector would be compatible with both the quorum sensing module and LacI-Lach module, thus we settled on pSB3T5 as our final vector. | As our individual modules would be ideally combined at a later point, we had to make sure that our final vector would be compatible with both the quorum sensing module and LacI-Lach module, thus we settled on pSB3T5 as our final vector. | ||
| - | + | <br><br> | |
[[Image:University_Aberdeen_2009_ANDgateStep1.PNG|center|400 px]] | [[Image:University_Aberdeen_2009_ANDgateStep1.PNG|center|400 px]] | ||
| - | + | <br><br> | |
Our first step was to join CI repressor with a ribosome binding site, green fluorescence protein and the double terminator. For this step we decided to use biobricks C0051, E0840 and clone into the pSB1AC3. As CI repressor gene was upstream we digested it with EcoRI and SpeI restriction enzymes. The downstream part was digested with XbaI and PstI, therefore vector was cut with EcoRI and PstI. Our experience with plasmid digests led us to use high copy number. This initial step was the creation of biobrick | Our first step was to join CI repressor with a ribosome binding site, green fluorescence protein and the double terminator. For this step we decided to use biobricks C0051, E0840 and clone into the pSB1AC3. As CI repressor gene was upstream we digested it with EcoRI and SpeI restriction enzymes. The downstream part was digested with XbaI and PstI, therefore vector was cut with EcoRI and PstI. Our experience with plasmid digests led us to use high copy number. This initial step was the creation of biobrick | ||
<html><a href="http://partsregistry.org/Part:BBa_K182100">K182100</a></html> | <html><a href="http://partsregistry.org/Part:BBa_K182100">K182100</a></html> | ||
| - | + | <br><br> | |
[[Image:University_Aberdeen_2009_ANDgate_Step2.PNG|center|400 px]] | [[Image:University_Aberdeen_2009_ANDgate_Step2.PNG|center|400 px]] | ||
| - | + | <br><br> | |
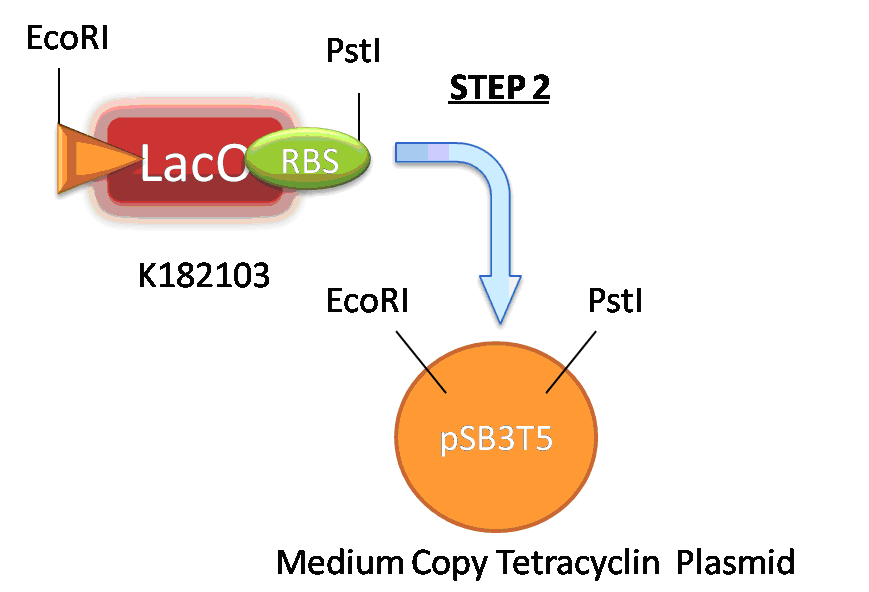
| + | Our second step required that we first create the actual biobrick K182102 using the method of fusion PCR. The promoter K182102 was composed of biobricks I751502 and B0034. As there was no DNA for biobrick I751502 we ordered four oligonucleotides. The forward primer was composed of the pSB3T5 sequence followed by half sequence of I751502. The reverse primer was created as part of pSB3T5, B0034 and half sequence of I751502. However first PCR product was necessary to extent therefore for the next PCR we used biobrick prefix and biobrick suffix with addition of random five nucleotide at their ends. Once the biobrick K182102 was prepared we digested it with EcoRI and PstI restriction enzymes as well as vector pSB3T5. | ||
<html> | <html> | ||
Revision as of 13:03, 20 August 2009
University of Aberdeen - Pico Plumber
Cloning Strategy
As our individual modules would be ideally combined at a later point, we had to make sure that our final vector would be compatible with both the quorum sensing module and LacI-Lach module, thus we settled on pSB3T5 as our final vector.
Our first step was to join CI repressor with a ribosome binding site, green fluorescence protein and the double terminator. For this step we decided to use biobricks C0051, E0840 and clone into the pSB1AC3. As CI repressor gene was upstream we digested it with EcoRI and SpeI restriction enzymes. The downstream part was digested with XbaI and PstI, therefore vector was cut with EcoRI and PstI. Our experience with plasmid digests led us to use high copy number. This initial step was the creation of biobrick
K182100
Our second step required that we first create the actual biobrick K182102 using the method of fusion PCR. The promoter K182102 was composed of biobricks I751502 and B0034. As there was no DNA for biobrick I751502 we ordered four oligonucleotides. The forward primer was composed of the pSB3T5 sequence followed by half sequence of I751502. The reverse primer was created as part of pSB3T5, B0034 and half sequence of I751502. However first PCR product was necessary to extent therefore for the next PCR we used biobrick prefix and biobrick suffix with addition of random five nucleotide at their ends. Once the biobrick K182102 was prepared we digested it with EcoRI and PstI restriction enzymes as well as vector pSB3T5.
 Back Back
|
Continue 
|
 "
"