Team:Aberdeen Scotland/parameters/invest 4
From 2009.igem.org
University of Aberdeen - Pico Plumber
Contents |
The Input-dependent model
For this model, we no longer assume a trivial process of repression of lacI being liften and QS being turned on. The LacI repression is now a repression / induction hill function with a Kd of 1200 molecules per cell for IPGT forming IPGT lacI complexes. We assume IPGT leaks into the cell from outside. We only set the outside concentration of IPGT. For quorum sensing all we set is an elevated HSL concentration outside which is free to defuse in. We now include a feedback loop into the QS mechanism. To do this we attach a lux box onto the plasmid for producing luxR and luxI, like so:
However, we are taking the production strengths of this lux box to be reduced in comparison to our lux box for producing X, Y and cI. The strengths are
Production max for X, Y and cI = 0.44 pops
taken from the iGEM page of ***** and personal communication with **** on ****
Production min for X Y and cI = 0.013 pops
taken from *****
Production max for QS feedback = 0.002 pops
Production min for QS feedback = 0.00015 pops
These values are taken from the paper “ FIOADAOIDFJAOFJASDIOJASOIJADOAJSDAFOJAFOAJSD”
An important new parameter is KP. This is the dissociation constant for “P” activating the lux boxes. P is the luxR/HSL complex . To take a break from the wordy descriptions we will now show some small sections of results for this model, which should be easily interpretable after following the section about the Monte Carlo analysis
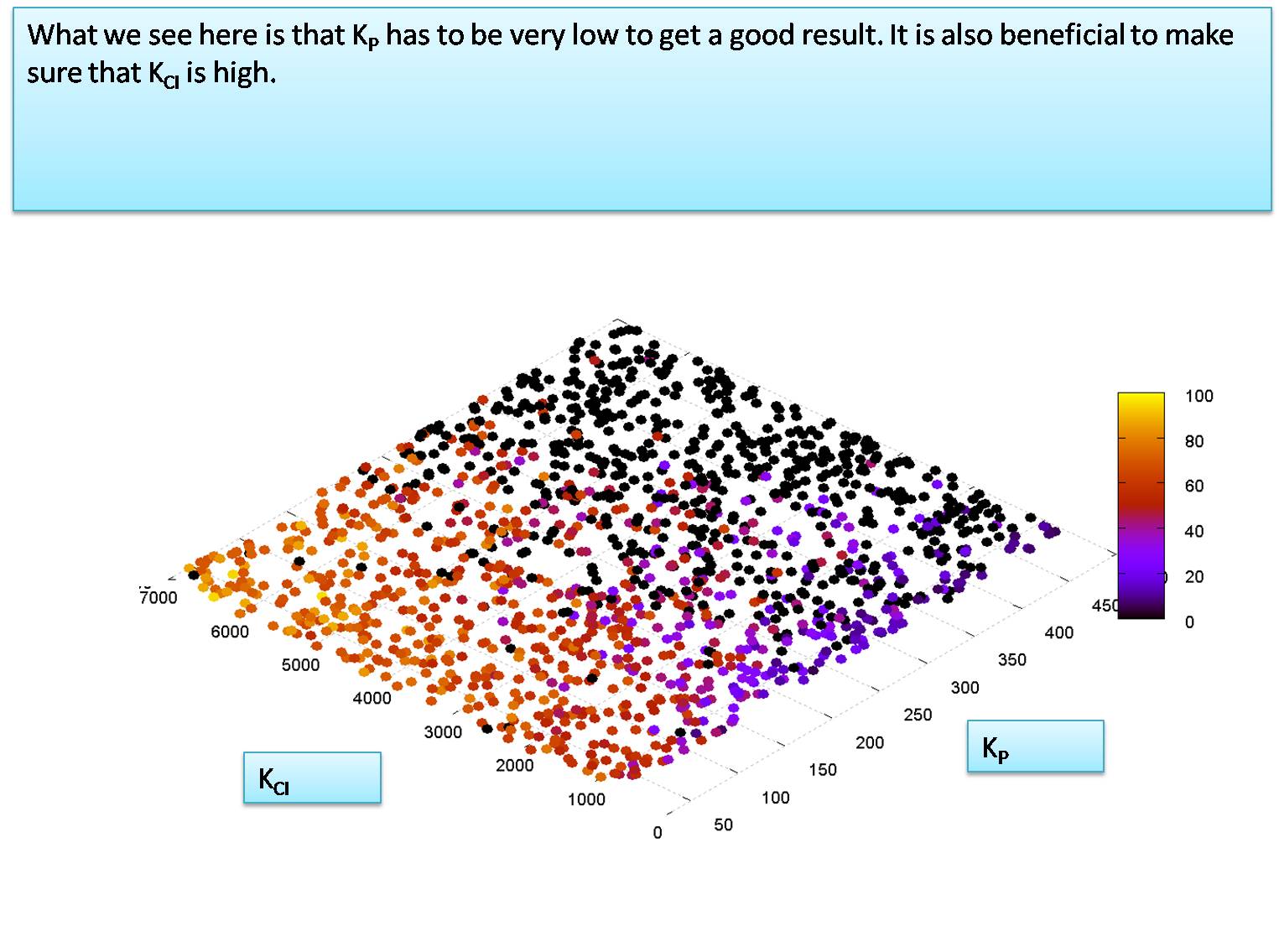
Level of KP required for system to function
here we use a Monte Carlo simulation to determine the value of KP in molecules per cell that is required for the system to function (note that the IPTG and HSL input levels added are both 10000 molecules)
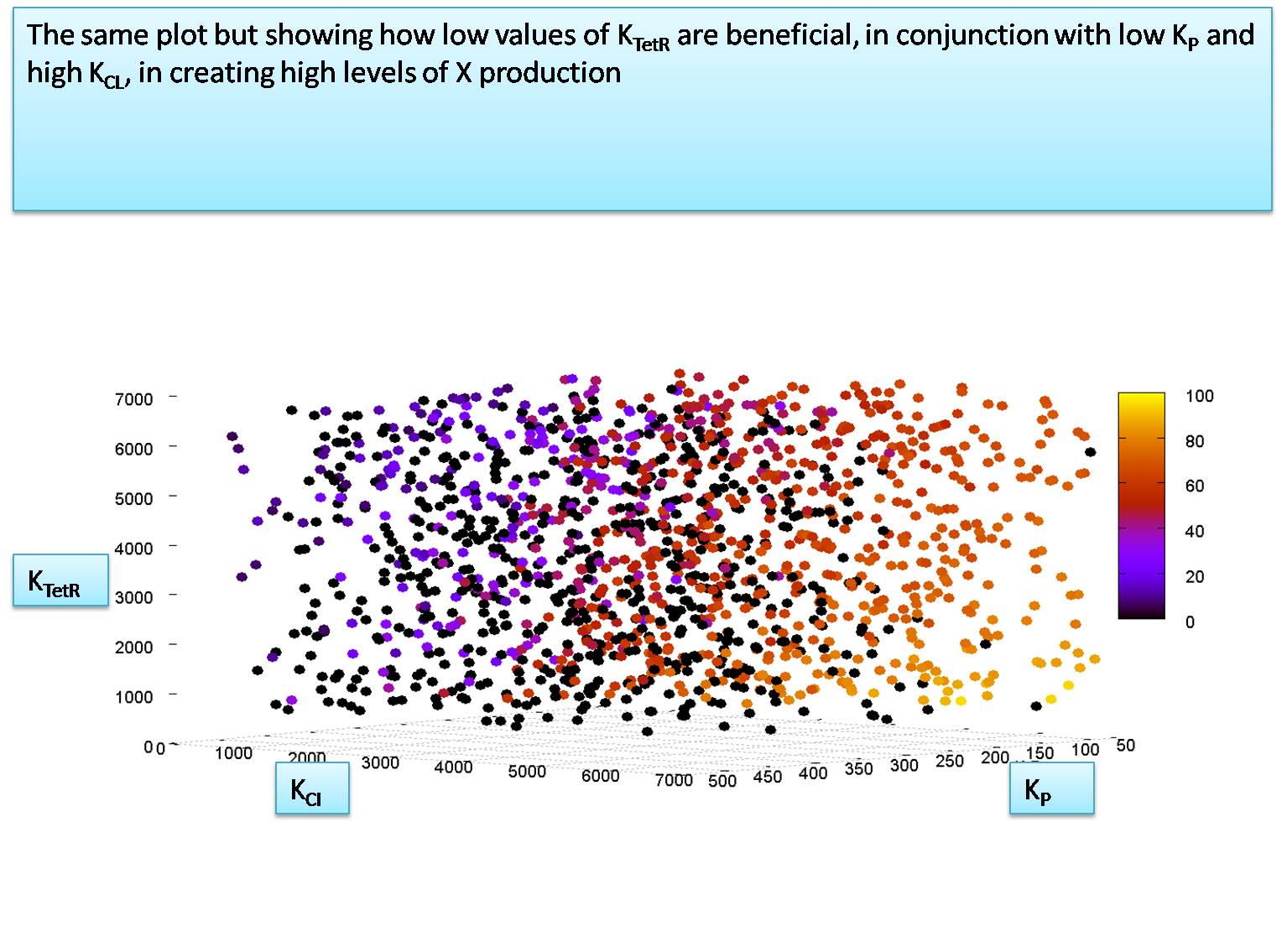
Relationship between KTetR and KCI when KP is 100
We notice that KP must be around 100 molecules to let the system function. We will now explore the relationship between KTetR and KCI at the value of KP=100 molecules per cell. for this simulation we set the HSL and IPTG input levels to 50,000 molecules. In our next section we will try to calculate the minimum levels of HSL and IPTG required to activate the system correctly.
Required input levels of IPTG and HSL
How the system should run
Here are the following assumptions that need to make
KP has to be ~50 molecules per cell KTetR has to be ~ molecules per cell KCI has to be ~ molecules per cell the input level of HSL has to be ~molecules the input level of IPTG has to be ~molecules
Relationship between KTetR and KCI when KP is 100
 "
"