Team:TUDelft/Research Proposal
From 2009.igem.org
(→References) |
(→References) |
||
| Line 171: | Line 171: | ||
<li id="cite_note-5"><cite id="CITEREFElbashir2001" class="article" style="font-style: normal;"> | <li id="cite_note-5"><cite id="CITEREFElbashir2001" class="article" style="font-style: normal;"> | ||
Elbashir SM, Harborth J, Lendeckel W, Yalcin A, Weber K, Tuschl T., 2001. [http://www.ncbi.nlm.nih.gov/pubmed/11373684 Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells]. Nature, 411(6836):494-498.</cite> | Elbashir SM, Harborth J, Lendeckel W, Yalcin A, Weber K, Tuschl T., 2001. [http://www.ncbi.nlm.nih.gov/pubmed/11373684 Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells]. Nature, 411(6836):494-498.</cite> | ||
| - | ==Conjugation== | + | <head>==Conjugation==</head> |
<li id="cite_note-6"><cite id="CITEREFThorsted1998" class="article" style="font-style: normal;"> | <li id="cite_note-6"><cite id="CITEREFThorsted1998" class="article" style="font-style: normal;"> | ||
Thorsted, P. B., D. P. Macartney, P. Akhtar, A. S. Haines, N. Ali, P. Davidson, T. Stafford, M. J. Pocklington, W. Pansegrau, B. M. Wilkins, E. Lanka, and C. M. Thomas. 1998. [http://www.ncbi.nlm.nih.gov/nuccore/113911681?ordinalpos=1&itool=EntrezSystem2.PEntrez.Sequence.Sequence_ResultsPanel.Sequence_RVDocSum Complete sequence of the IncPbeta plasmid R751: implications for evolution and organisation of the IncP backbone]. J. Mol. Biol. 282:969-990.</cite> | Thorsted, P. B., D. P. Macartney, P. Akhtar, A. S. Haines, N. Ali, P. Davidson, T. Stafford, M. J. Pocklington, W. Pansegrau, B. M. Wilkins, E. Lanka, and C. M. Thomas. 1998. [http://www.ncbi.nlm.nih.gov/nuccore/113911681?ordinalpos=1&itool=EntrezSystem2.PEntrez.Sequence.Sequence_ResultsPanel.Sequence_RVDocSum Complete sequence of the IncPbeta plasmid R751: implications for evolution and organisation of the IncP backbone]. J. Mol. Biol. 282:969-990.</cite> | ||
Revision as of 20:39, 14 July 2009
Research Proposal
Contents |
Part 1: delay device
Time delay genetic circuit This kind of circuit has the ability to integrate signals and trigger events after a delay from the initial detection event. There are two approaches to construct a time delay genetic circuit, these are:
- “protein-based” transcriptional regulators
- “RNA-based” posttranscriptional regulators[1]
Based on different literature, we have four different configurations for the time delay genetic circuit. The first two are protein-based and the last two RNA-based.
Negative Feedforward
Based on Hooshangi et al.[2] and the project requirements, the next scheme can be proposed.
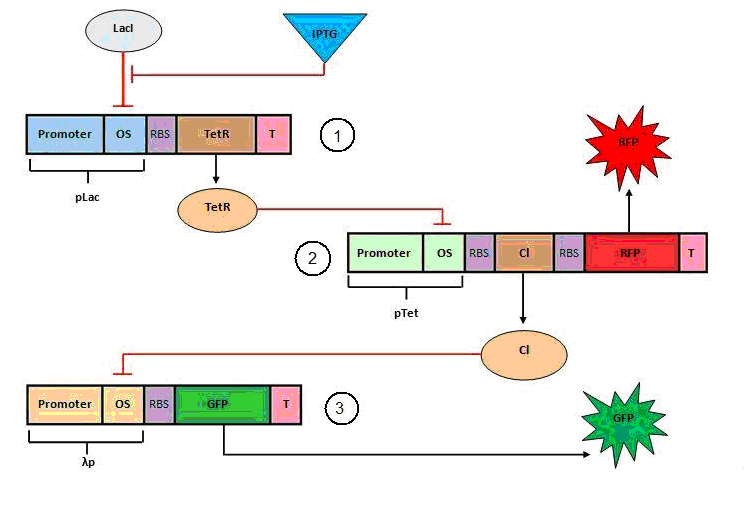
Figure 1: Genes 1 and 3 are present in the cell and they are produced all the time (black). Genes 2 and 4 (green) are in the self destructive plasmid (SDP). ![]() indicates repression.
indicates repression.
Description
Protein 1 starts the circuit activating (or repressing the repressor of) the promoter of gene 2, for proof of concept gene 1 can be changed for a signal molecule. The concentration of protein 2 will increase and achieve the threshold concentration to repress gene 3. At this point the production of protein 3 stops. As the concentration of protein 3 decreases due to its degradation, the repression of gene 4 will not longer exist and the production of protein 4 (restriction enzyme or fluorescent protein) starts.
Parameters
In order to make a longer or shorter delay the protein production and degradation rates have to be considered. For the former, we can “play” with the ribosome binding site (RBS) and promoter strength. The degradation can be controlled by protein stability, induction of degradation and up-down regulation of degradation enzymes.
Biobricks
- new RBS
- new transcriptional factors with different stabilities
- new promoters with different strength
Take into account
- There are many of biobricks 1 and 3 in the registry
- Time frame
- Change LVA tag to decrease degradation
Modeling
Differential equations involving production rate, degradation rate (and dilution rate)
Positive feedforward
Based on the book of Uri Alon[3] and the project requirements, the next scheme can be proposed.
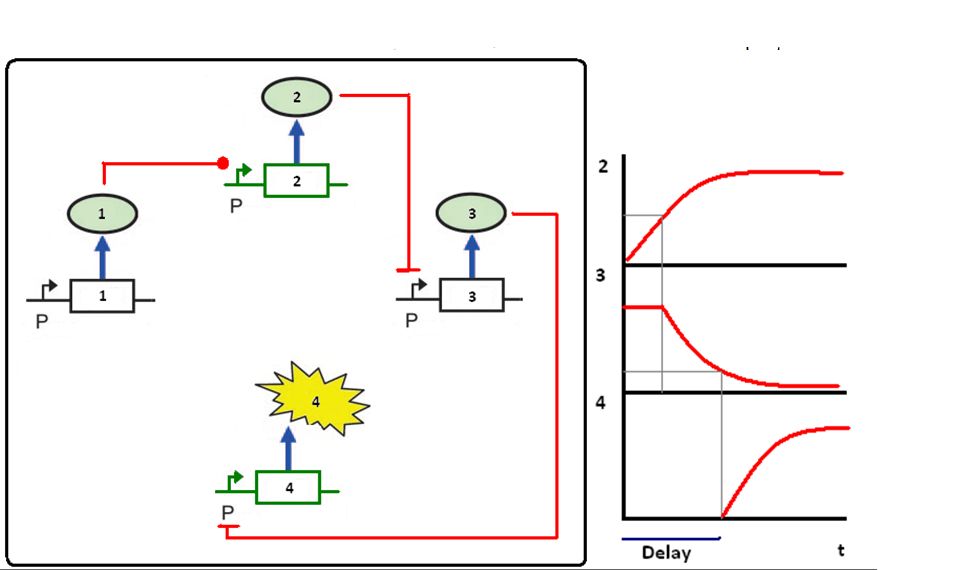
Figure 2: Genes 1 and 3 are present in the cell and they are produced all the time (black). Genes 2 and 4 (green) are in the self destructive plasmid (SDP).
Description
Protein 1 starts the circuit activating the promoter of gene 2, for proof of concept gene 1 can be changed for a signal molecule. The concentration of protein 2 will increase and achieve the threshold concentration to induce gene 3. As the concentration of protein 3 increases, the induction of gene 4 (restriction enzyme or fluorescent protein) starts.
Parameters
In order to make a longer or shorter delay the protein production and degradation rates have to be considered. For the former, we can “play” with the ribosome binding site (RBS) and promoter strength. The degradation can be controlled by protein stability, induction of degradation and up-down regulation of degradation enzymes.
Biobricks
- new RBS
- new transcriptional factors with different stabilities
- new promoters with different strength
Take into account
- There are many of biobricks like 1 and 3 in the registry
- Positive feed is always related to leaking problems
- Time frame
- Change LVA tag to decrease degradation
Modeling
Differential equations involving production rate, degradation rate (and dilution rate)
Riboregulator
In bacteria, several factors affect translation initiation, including ribosomal recognition of the mRNA’s RBS and the start codon (AUG). Recognizing the importance of RNA interactions between the ribosome and RBS, and based on work on endogenous riboregulators, Isaacs et al.[4] sought to regulate bacterial gene expression by interfering with ribosomal docking at the RBS. From the outset, their objective was to create a modular post transcriptional regulation system that could be integrated into biological networks and implemented with any promoter or gene. This system was already used by Friedland et al[5].
Circuit 3 that we have in mind is based on the riboregulator and is shown in figure 3.

Figure 3: taRNA/RBS (riboregulator) regulated expression of protein 3. Genes 1 and 2 are on the conjugative plasmid present in all cells. S1: signal molecule. Gene 3 is present on the SDP. RBS: ribosome binding site. Cr: piece of RNA strand in front of RBS that is complementary to RBS. taRNA: trans-activating RNA, could also be small interference RNA, as long as it is complementary to cr. Product 3: Could be either GFP/luciferase (for testing) or endonuclease.
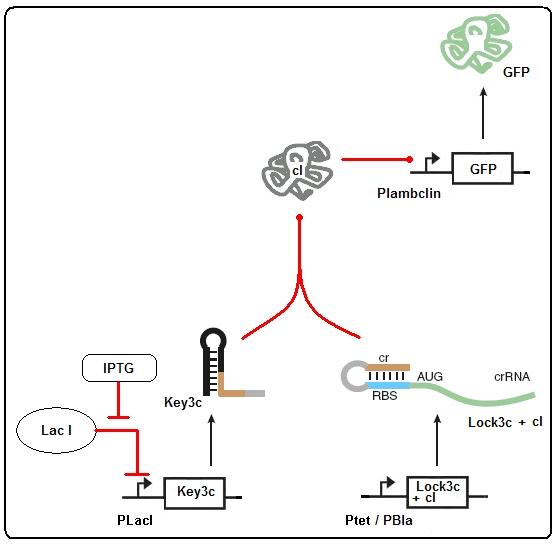
Figure 4: once the
Description
Either gene 1 or gene 2 is constitutively expressed (never both!) where the other is inducible. In the picture, gene 2 is constitutive and gene 1 is inducible by signal molecule S1. Gene 2 expresses constitutively the crRNA, on which the cr part of the RNA binds the RBS directly after transcription, causing RBS to be blocked for the ribosome and therefore blocks the RNA from translation. When the promoter of gene 1 is induced by S1, it produces the taRNA molecule which is complementary to the cr part of the crRNA. When the taRNA binds the crRNA it opens the RBS, which makes the crRNA ready for translation. The resulting product (the green folded protein 2) can then induce gene 3 to produce end-product 3.
Parameters and take into account
- taRNA should be better complementary to cr than cr to RBS, to make sure that taRNA causes the RBS to open up.
- One can play with promoter strengths for genes 1 and 3 (or when gene 1 is constitutively expressed, genes 2 and 3).
- The degradation rate of protein 2 (the green folded protein in figure 3) can influence the delay time.
- Instead of protein 2 inducing gene 3 expression, it might repress a repressor of promoter 3 (negative feed-forward, see figure 1), to get rid of leaky expression.
- Take into account “scars”
- Time frame
Biobricks
- new RBS
- new transcriptional factors with different stabilities
- new promoters with different strength
- crRNA for each RBS in the registry
- taRNA for each crRNA
Modeling
- Differential equations involving production rate, degradation rate (and dilution rate)
- Interaction crRNA’s – RBS and crRNA’s – taRNA’s
- Stability of taRNA’s
Small interfering RNA
This approach is based on the gene silencing by small interfering RNA explained in Elbashir et al.[6] and is given in the following scheme.
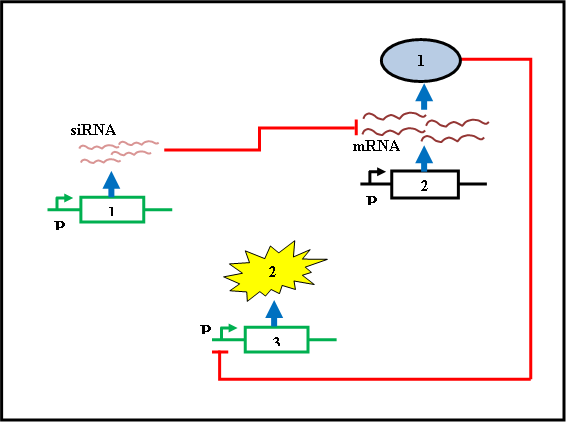
Figure 5: Genes 1 and 3 are present in the self destructive plasmid. Gene 2 is present in the cell and is always produced.
Description
Protein 1 is always produced in the cell and is controlled by the constitutive promoter on gene 2. This protein must be selected carefully since the gene 1 producing the siRNA must be decided based on this protein and the siRNA must be specific for silencing the mRNA of protein 1 without disturbing the other mRNAs of the host organism. The protein 1 will be in excess concentration and hence will inhibit the production of protein 2 (restriction enzyme or fluorescence protein) by repressing the promoter of gene 3. When the production of siRNA is induced by activating the promoter of gene 1, on reaching a threshold it will interfere with the translation of protein 1 and hence the repression of gene 3 is released to produce the protein 2.
Parameters and take into account
- Protein 1’s stability
- siRNA specificity
- Promoter strength.
Biobricks
- new RBS
- new transcriptional factors with different stabilities
- new promoters with different strength
- siRNA’s
Modeling
- Differential equations involving production rate, degradation rate (and dilution rate)
- Interaction siRNA’s - mRNA
Which biobricks we could use
| Part | Biobrick | well | Plate | Plasmid |
|---|---|---|---|---|
| pLac/PLacI | R0010 | 1D | 1 | pSB1A2 |
| PBla | I14018 | 18N | 1 | pSB2K3 |
| λp/Plambclin | I12006 | 11J | 2 | pSB2K3 |
| pTet/Ptet | R0040 | 6I | 1 | pSB1A2 |
| key3c | J23008 | 3F | 1 | J23006 |
| Lock3c | J23031 | 3L | 1 | J23006 |
| GFP | E0040 | 14K | 1 | pSB1A2 |
| mRFP1 | E1010 | 18F | 1 | pSB2K3 |
| cI | C0051 | 4E | 1 | pSB1A2 |
| RBS | B0034 | 2M | 1 | pSB1A2 |
| T (Double Terminator) | B0015 | 23L | 1 | pSB1AK3 |
| λp+RBS+GFP+T | S03335 | 85 | Box9 | pSB1A2 |
| λp+RBS+mRFP1+T | S03473 | 79 | Box9 | pSB1A2 |
Part 3: conjugation
Selecting the Conjugation system
There are many different conjugation systems, with F and IncP being among the most used by previous iGEM teams. The IncP incompatibility group is further divided into two groups IncP-alpha aka Birmingham aka RP4 plasmid and the IncP-beta R751 plasmid. There are several reasons why the IncP system would be better suited for us:
- Smaller plasmid size (R751 is 53423[7]) (RP4 is 60099[8]) (F is 99159[9])
- F has a self-imposed fertility inhibition system (new donors fertile for only about 6 generations), IncP systems have no self-imposed fertility inhibition system[10]
- F has two surface exclusion proteins (TraT, TraS), IncP has only one (trbK)[10]
Selecting genes to knockout
Entry exclusion
The goal of this part of the project is to allow a message plasmid to be transmitted between cells containing conjugation helper plasmids (modified R751). In nature this occurs but at very low levels, due to the presence of entry exclusion proteins[11]. These membrane-bound proteins block incoming transfers from cells containing the conjugative plasmid. Note that in order for the transfer to be blocked their presence is required only in the recipient. These entry exclusion proteins are present for two primary reasons: i) to prevent redundant transfer of the conjugative plasmid among a population of cells and ii) to prevent lethal zygosis[12]. In R751 the gene trbK encodes for the entry exclusion protein[13] in R751 and RP4. Its presence is not required for conjugation to occur[14]. Some papers have reported that knocking out the trbK genes in other plasmids did not affect the conjugation frequency: “transfer of the trbK mutant occurred at near-wild-type frequencies”[15]. Given this information we propose to knockout the trbK gene.
Lethal Zygosis
What is lethal zygosis: “We find that diaminopimelic acid in the recipient membrane is released into the medium during bacterial matings, indicating that membrane damage was inflicted on the recipient by the donor, probably for forming a channel for DNA transfer. When the damage is extensive, as in matings with an excess of Hfr bacteria, the F- bacteria are killed (lethal zygosis). The transfer of a large amount of DNA in Hfr matings appears to enhance the killing.[12]”
By knocking out the trbK gene we may be exposing our cells to lethal zygosis. If this is indeed the case than knocking out one of the critical mating pair formation genes (trbB, trbC, trbD, trbE, trbF, trbG, trbH, trbI, trbJ, trbL) from the trb operon would prevent this. The gene trbC was selected for this due to its small size and position on the operon. trbC is known to encode for the pilin subunits needed for pilus formation. Given this information we propose to knockout the trbC gene as well.
Experimental Procedures
Section 1: Helper Plasmid
Part 1A:
- Acquire R751 and/or RP4 plasmid
- Electoporate plasmid into cells and confirm conjugation. See conjugation protocol
- Characterize conjugation efficiency
Part 1B: oriT knockout
- Knockout oriT using either lambda red, knockout protocol used by Peking '07 or knockout protocol used by Berkeley '06
- Electroporate into cells
- Verify that conjugation stopped
- Electroporate PlasmidG as well and characterize conjugation efficiency
- If works send R751 oriT knockout plasmid to registry
Part 1C: trbK knockout
- Knockout oriT + trbK
- Electroporate into cells
- Verify that conjugation takes place among R+ cells using PlasmidG (see below)
- Characterize conjugation efficiency
- If works send R751 oriT + trbK knockout plasmid to registry
Part 1D: trbC knockout
- Knockout oriT + trbK + trbC
- Electroporate into cells and create culture for communication (cultureCom)
- Verify that no conjugation takes place in presence of PlasmidG
- If works send R751 oriT + trbK + trbC knockout plasmid to registry
Section 2: Message Plasmid
Part 2A: BioBrick Assembly
- Order DNA synthesis for BBa_K175000 (trbC) and BBa_K175001 (trbK)
- Amplify BioBricks needed BBa_E0840 (GFP generator), BBa_B0034 (strong rbs), BBa_J23100 (constitutive promoter), BBa_I714031 (oriT-R).
- Assemble: [oriT][promoter] and keep this for later
- Assemble PlasmidG: [oriT][promoter][GFP generator] + plasmid backbone with different resistance than helper plasmid and low copy number.
- Assemble [oriT][promoter][rbs][trbC][rbs][trbK] and keep this intermediate assembly product for possible use in integration stage
- Assemble [oriT][promoter][rbs][trbC][rbs][trbK][GFP generator] and keep this intermediate assembly product for possible use in integration stage
- Assemble PlasmidCKG: [oriT][promoter][rbs][trbC][rbs][trbK][GFP generator] + plasmid backbone with different resistance than helper plasmid and low copy number.
Part 2B: Full Communication testing
- Electroporate PlasmidCKG into some cells from cultureCom creating initiatorCells
- Select for presence of both message and helper plasmid
- Mix initiatorCells with cells not containing any plasmids and characterize conjugation efficiency
- Add initiatorCells to cultureCom and observe signal propagation, characterize rate of signal propagation.
- If signal propagation observed, do victory dance.
References
Delay device
- Smolke C, 2009. It’s the DNA That Counts. Science, 324:1156-1157.
- Hooshangi S, Thiberge S, Weiss R, 2005. Ultrasensitivity and noise propagation in a synthetic transcriptional cascade. PNAS, 102,10 :3581–3586.
- An introduction to systems biology: design principles of biological circuits. Uri Alon, Publisher CRC Press, 2006, pp 41-69.ISBN 1584886420
- Isaacs F, Dwyer d and Collins J, 2006. RNA synthetic biology. Nature Biotech., 24:545-554.
- Friedland A, Lu T, Wang X, Shi D, Church G and Collins J, 2009. Synthetic Gene Networks That Count. Science, 324:1199-1202.
- Elbashir SM, Harborth J, Lendeckel W, Yalcin A, Weber K, Tuschl T., 2001. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature, 411(6836):494-498. <head>==Conjugation==</head>
- Thorsted, P. B., D. P. Macartney, P. Akhtar, A. S. Haines, N. Ali, P. Davidson, T. Stafford, M. J. Pocklington, W. Pansegrau, B. M. Wilkins, E. Lanka, and C. M. Thomas. 1998. Complete sequence of the IncPbeta plasmid R751: implications for evolution and organisation of the IncP backbone. J. Mol. Biol. 282:969-990.
- Pansegrau, W., E. Lanka, P. T. Barth, D. H. Figurski, D. G. Guiney, D. Haas, D. R. Helinski, H. Schwab, V. A. Stanisich, and C. M. Thomas. 1995. Complete nucleotide sequence of Birmingham IncP alpha plasmids. Compilation and comparative analysis. J. Mol. Biol. 239:623-663.
- Frost, L., K. Ippen-Ihler, and M. Skurray. 1995. Analysis of the sequence and gene products of the transfer region of the F sex factor. Microbiol. Rev. 58:162-210.
- Plasmid Biology. Barbara E. Funnell and Gregory J. Phillips, eds. American Society for Microbiology Press, Washington, DC, 2004. Chapter 9.ISBN 1555812651
- Garcillán-Barcia MP, de la Cruz F. Why is entry exclusion an essential feature of conjugative plasmids? 2008;60:1–18. doi: 10.1016/j.plasmid.2008.03.002.
- Ou, J. T. 1980. Role of surface exclusion genes in lethal zygosis in Escherichia coli K12 mating. Molecular and General Genetics 178:573–581.
- Haase, J., Kalkum, M. & Lanka, E., 1996. TrbK, a small cytoplasmic membrane lipoprotein, functions in entry exclusion of the IncP alpha plasmid RP4. J. Bacteriol. 178, 6720–6729.
- Haase J, Lurz R, Grahn A M, Bamford D H, Lanka E., 1995. Bacterial conjugation mediated by plasmid RP4: RSF1010 mobilization, donor-specific phage propagation, and pilus production require the same Tra2 core components of a proposed DNA transport complex. J Bacteriol. 177:4779–4791.
- Li, P. L., Hwang, I., Miyagi, H., True, H., Farrand, S. K., 1999. Essential components of the Ti plasmid trb system, a type IV macromolecular transporter. J. Bacteriol. 181:5033-5041.
 "
"