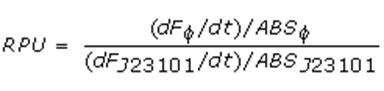Team:UNIPV-Pavia/Parts Characterization
From 2009.igem.org

|
|
|
Parts Characterization |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Here we describe the characterization results of 4 parts of our own design, 2 existing parts re-built because they were inconsistent and 7 existing parts taken from the Registry. When not reported differently, all the experiments have been performed according to Growth conditions and Data analysis sections.
Re-built existing parts (BBa_our part code/BBa_existing part code): Existing parts from the Registry:
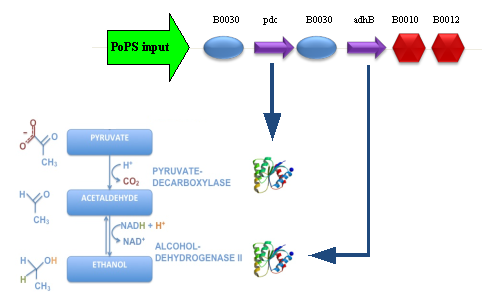
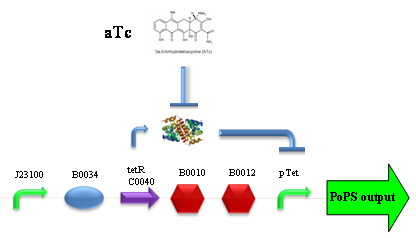
Our new partsBBa_K173003 - ethanol producing deviceDescriptionThis device takes PoPS as input and produces pyruvate decarboxylase (pdc) and alcohol dehydrogenase II (adhB) enzymes. Pyruvate decarboxylase (pdc, ) catalyses the decarboxylation of pyruvic acid to acetaldehyde and carbon dioxide, while alcohol dehydrogenase II (adhB, ) catalyses the acetaldehyde reduction to ethanol. The latter enzyme, as reported in literature, is also able to work in the opposite direction, oxidizing ethanol to acetaldehyde. The two enzymes come from Zymomonas mobilis ethanologenic bacterium and constitute an essential step of alcoholic fermentation. So, this device contains the minimum set of genes that are required to engineer a heterologous fermentation pathway. The coding sequences of pdc and adhB genes have been optimized for Escherichia coli codon usage. CharacterizationConclusionsBBa_K173007 - aTc inducible device with J23100 promoterDescriptionThis is an aTc sensing device. promoter drives the constitutive production of tetR repressor (), which inhibits tetR promoter () activity. When aTc is added to the medium, it binds tetR and inhibits it. So, the PoPS output is a function of the aTc concentration. A tight regulation is expected for this inducible system because BBa_J23100 is a strong promoter and so tetR repressor shold be produced at extremely high levels. The data below are referred to , which is the measurement system of . Characterization
ConclusionsWe demonstrated that this part works as expected, sensing the aTc concentration provided in the culture medium. The transfer function of this sensor has been characterized in standard units (RPUs) in two different growth media (LB and M9 supplemented with glycerol), as well as the metabolic burden (in terms of doubling time) which affects E. coli bearing this part. On the other hand, we did not expect to have a higher GFP synthesis rate per cell after the exponential growth phase than in the exponential phase itself (as reported in the 3rd plot). COMMENTARE QUI IL TIGHT REGULATION BBa_K173011 - aTc inducible device with J23118 promoterDescriptionThis is an aTc sensing device. promoter drives the constitutive production of tetR repressor (), which inhibits tetR promoter () activity. When aTc is added to the medium, it binds tetR and inhibits it. So, the PoPS output is a function of the aTc concentration. A less tight regulation is expected for this inducible system than in because BBa_J23118 promoter is weaker than BBa_J23100 and so tetR repressor shold be produced at lower levels than in the other sensor. The data below are referred to , which is the measurement system of . Characterization
ConclusionsWe demonstrated that this part works as expected because GFP is produced as an increasing function of the aTc concentration provided in the culture medium. The transfer function of this sensor has been characterized in standard units (RPUs) in two different growth media (LB and M9 supplemented with glycerol), as well as the metabolic burden (in terms of doubling time) which affects E. coli bearing this part. On the other hand, as for BBa_K173007, we did not expect to have a higher GFP synthesis rate per cell after the exponential growth phase than in the exponential phase itself (as reported in the 3rd plot). COMMENTARE QUI LA PARTE DEL LEAKAGE E CONFRONTARLA CON K173007 BBa_K173010 - lactose/IPTG inducible device with J23118 promoterDescriptionThis should work as a lactose/IPTG sensor. promoter drives the constitutive production of lacI repressor (), which inhibits lac promoter () activity. When lactose or IPTG is added to the medium, it binds lacI and inhibits it. So, the PoPS output is a function of lactose/IPTG concentration. Thanks to the hybrid lac promoter (), designed taking the Plambda promoter () and substituting its cI () binding sites with two lacI binding sites, the behaviour of this device is not a function of glucose concentration because the wild type CAP binding sites are not present in this artificial lac promoter. CharacterizationConclusionsRe-built existing partsBBa_K173004/BBa_I732019 - beta-galactosidase protein generatorBBa_K173005/BBa_Q04400 - tetR QPIExisting parts from the RegistryBBa_J23100, BBa_J23101, BBa_J23118 - constitutive promoter family membersDescriptionThese three promoters are from the Anderson Promoter Collection, which is a library of constitutive sigma70 bacterial promoters. The strength of each promoter of the library has already been estimated in saturation growth phase cultures in LB, but here we provide the characterization of BBa_J23100 and BBa_J23118 in standard units (RPUs) in LB medium, in order to add experience and data for these BioBricks. BBa_J23101 is the reference standard promoter, so it has RPU=1 for definition. The data shown below are referred to , and that are the measurement parts of respectively , and . Characterization
ConclusionsRPU estimation of these promoters was not present in the Registry and even the doubling time of these parts was not documented. We added these data in the pages of , and characterized parts, hoping that they can be useful for promoter comparison in standard units. If we consider the promoter ranking, provided in saturation phase in the Anderson Promoter Collection Registry page, the estimated strength in RPU of BBa_J23100 and BBa_J23118 are in accordance with these values: INSERIRE I VALORI E COMMENTARE Note: plasmid is equals to pSB1A2 with a RFP expression system downstream of the cloning site. BBa_F2620 - 3OC6HSL receiver device
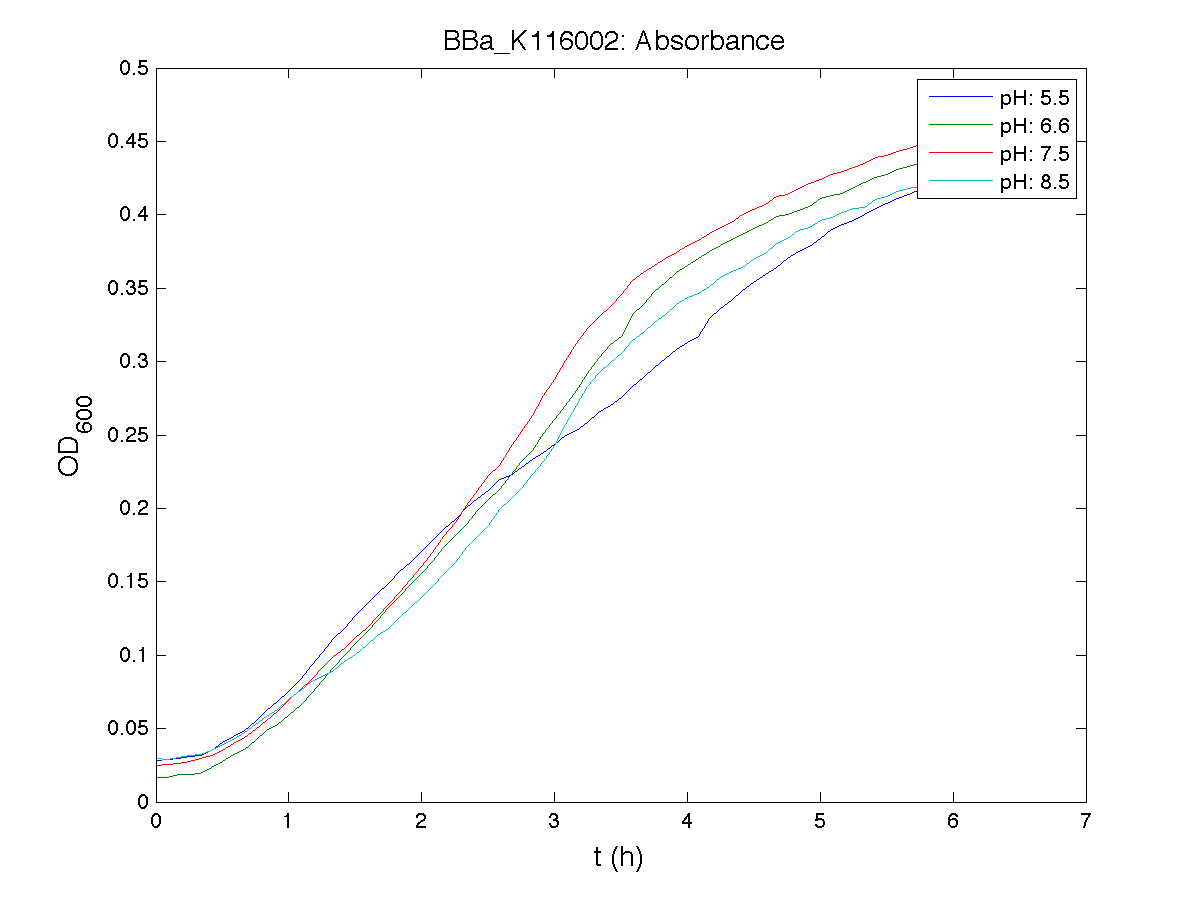
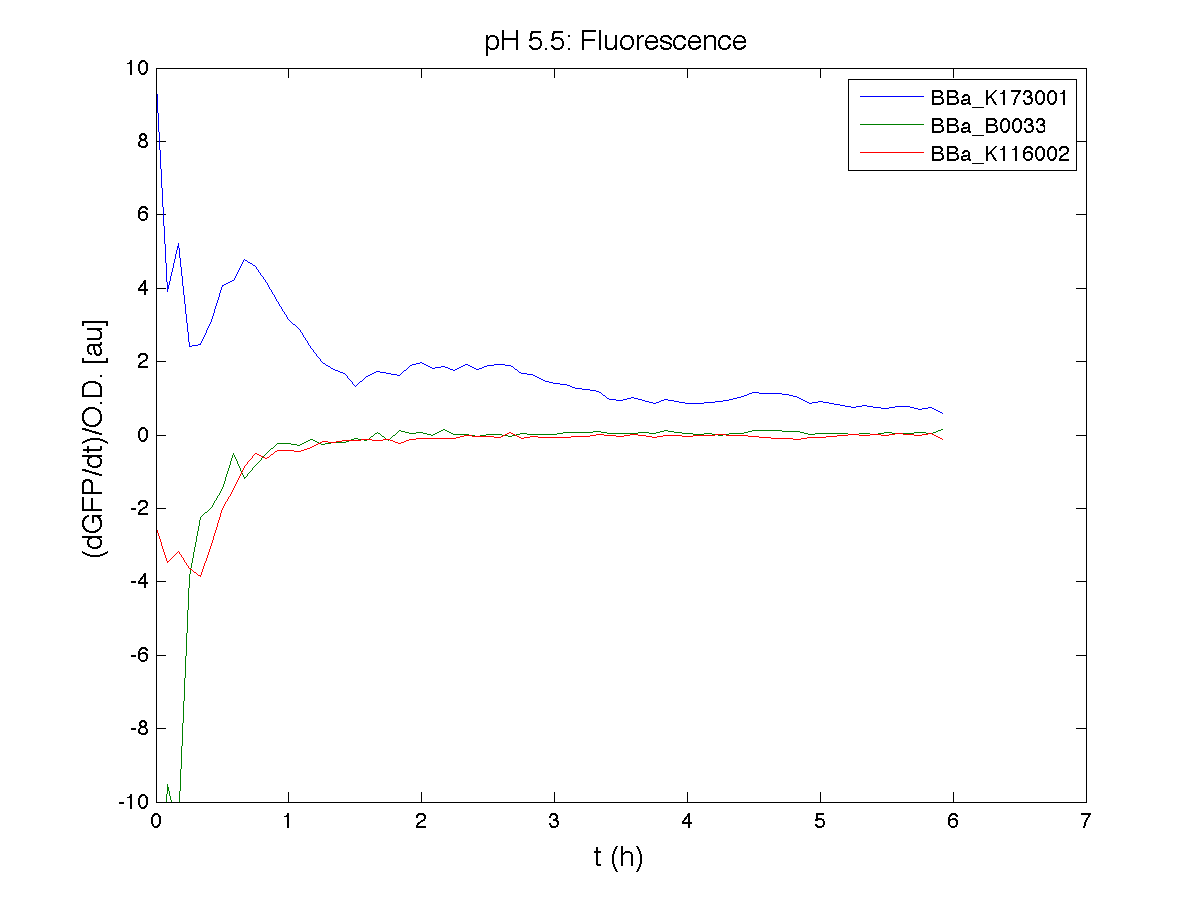
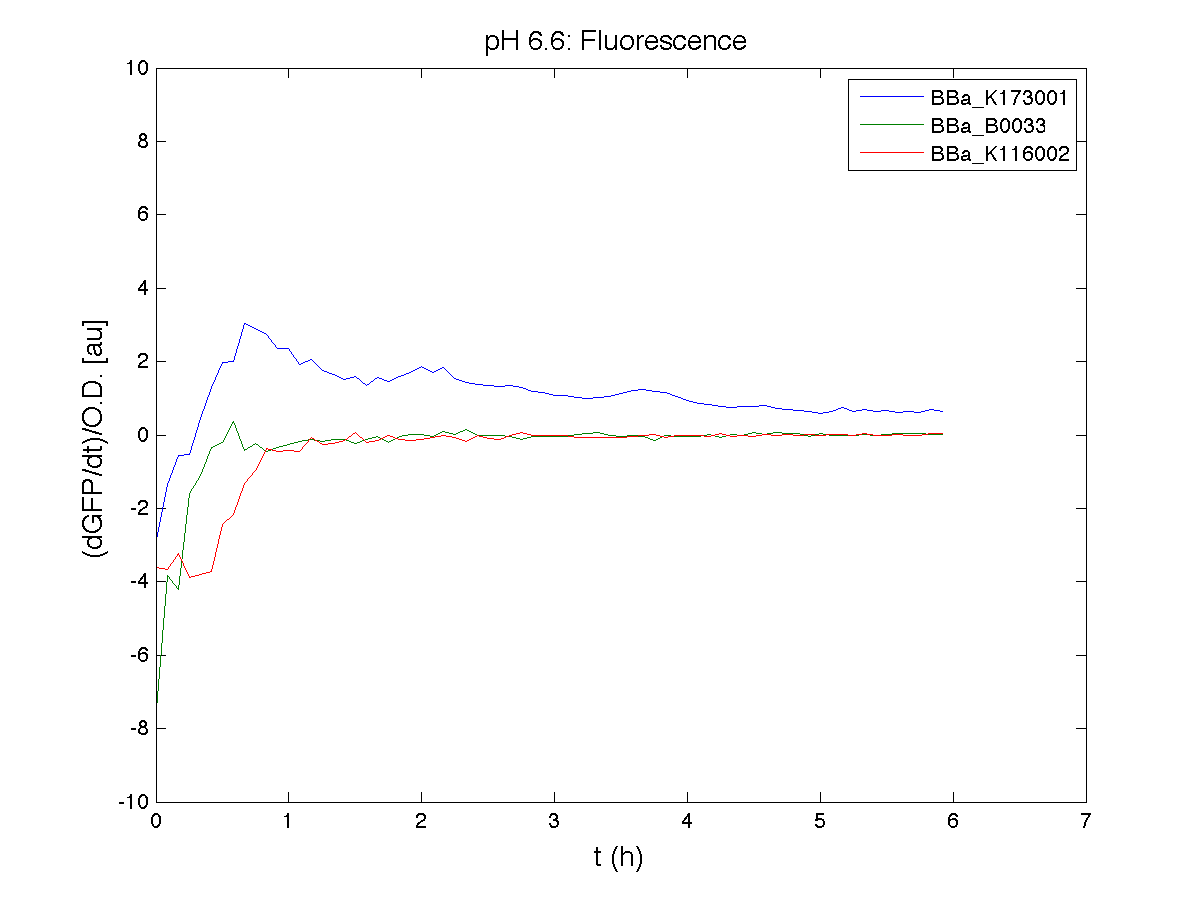
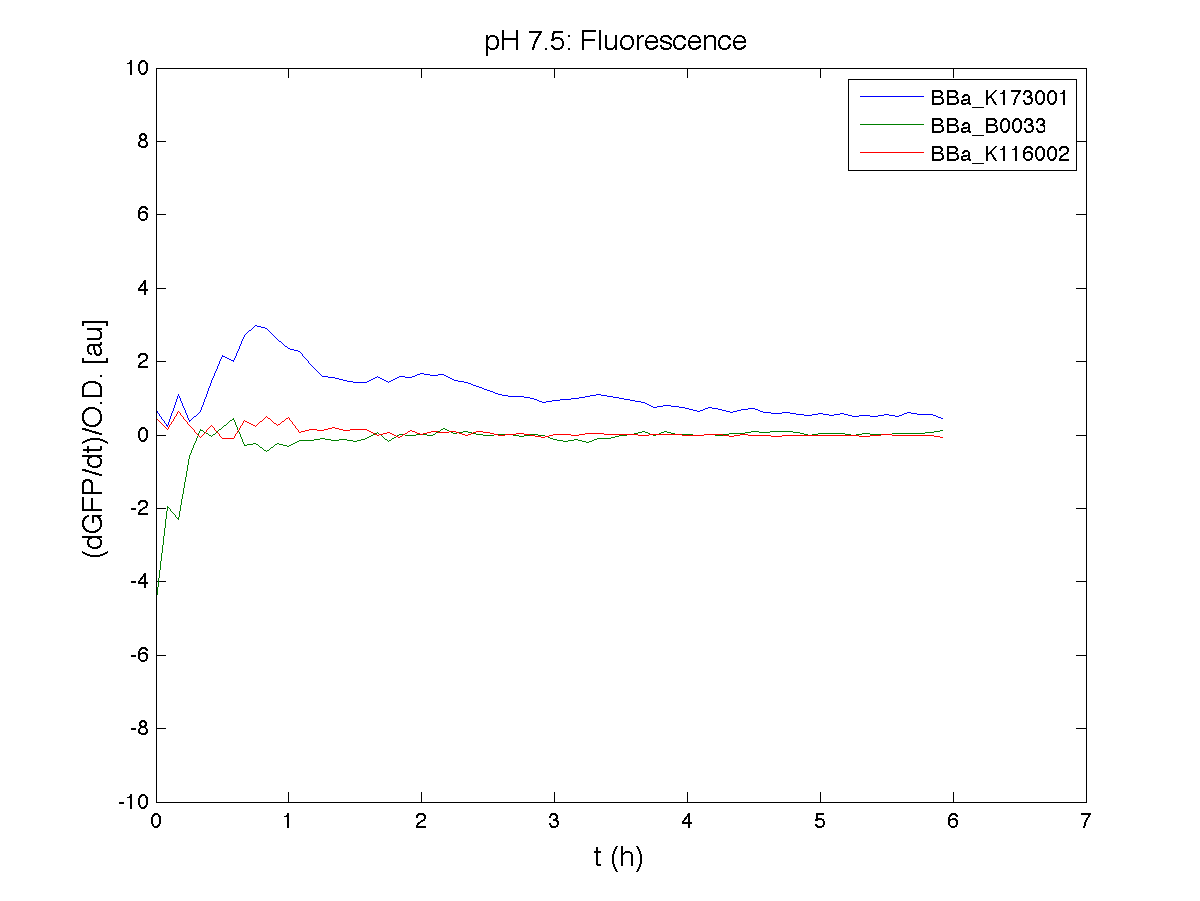
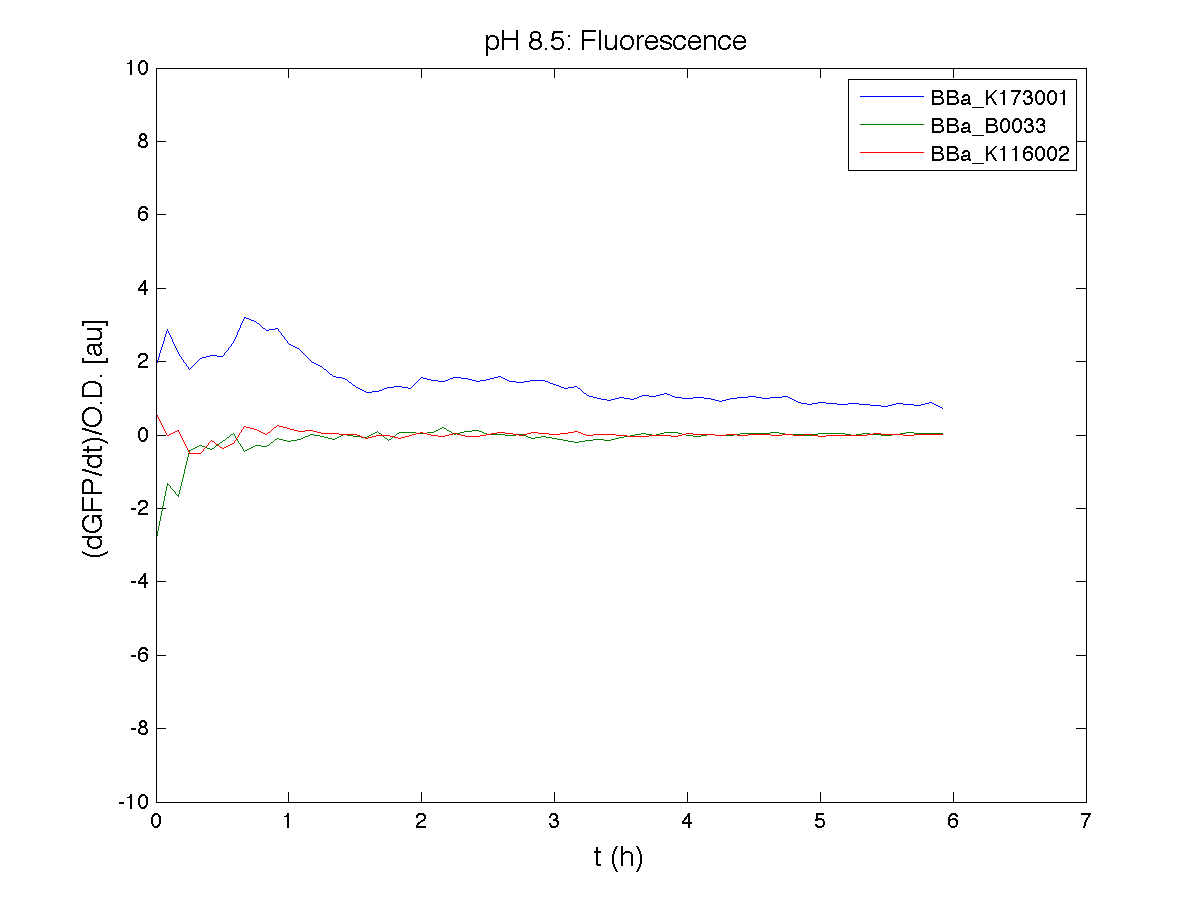
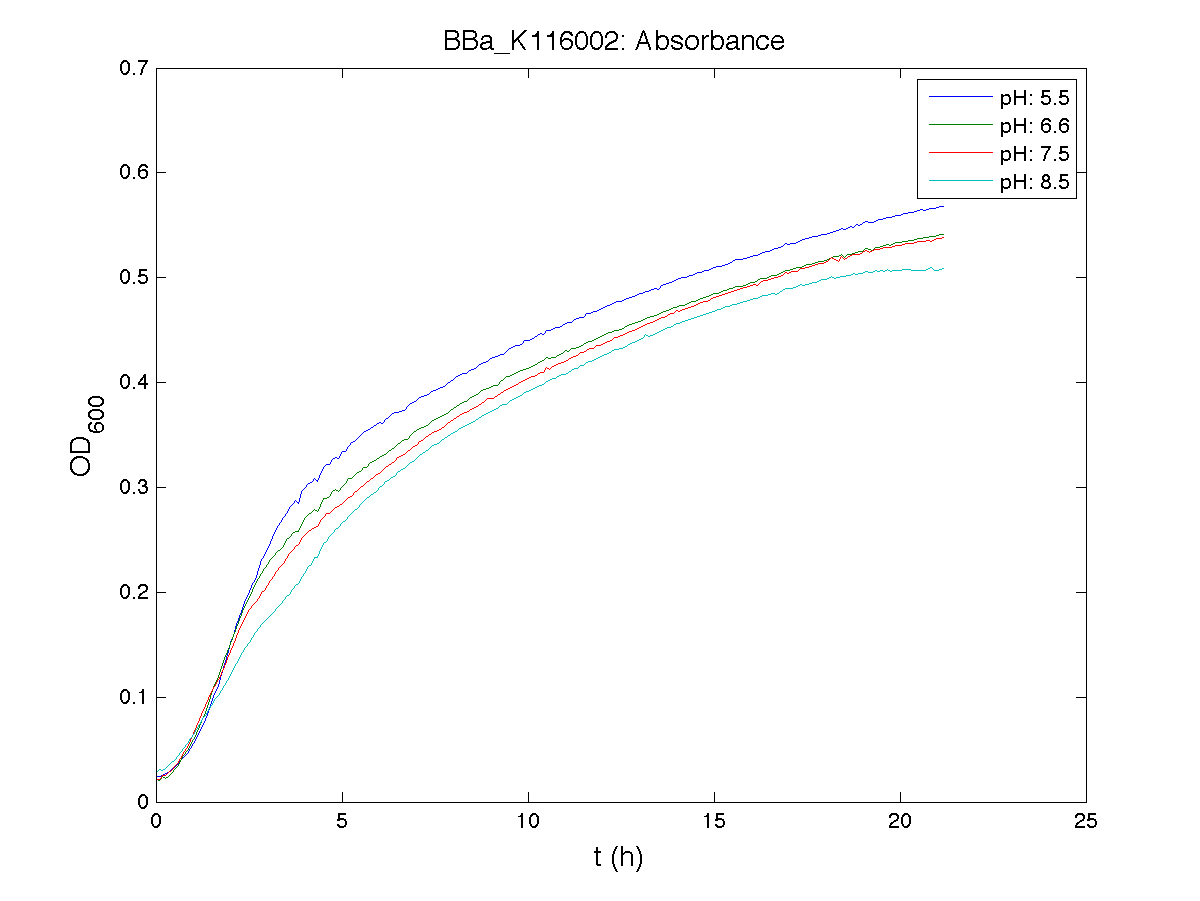
BBa_K116001 - nhaA promoterBBa_K116001 from iGEM 2008 NYMU-Taipei We received this BioBrick from iGEM in September but the bacterial strain that contained the plasmid wasn't declared. So we decided to sequence it as check and transform it into E.coli TOP10. We wanted to perform some experiments to better understand how it works and if can be successfully used. We performed several experiments with different LB medium and we got almost the same results. We used:
Here we show just two experiments to explicate our work. You can download the complete list from this link. Experiment Na+ 0MMotivationIn our opinion the working principle of the antiporter Na+/H+ channel described in [Rachel Karpel et al., Etana Padan et al., N. Dover et al.] makes the nhaA promoter a Na+ sensor and only under certain conditions (presence of Na+) a pH sensor. Methods
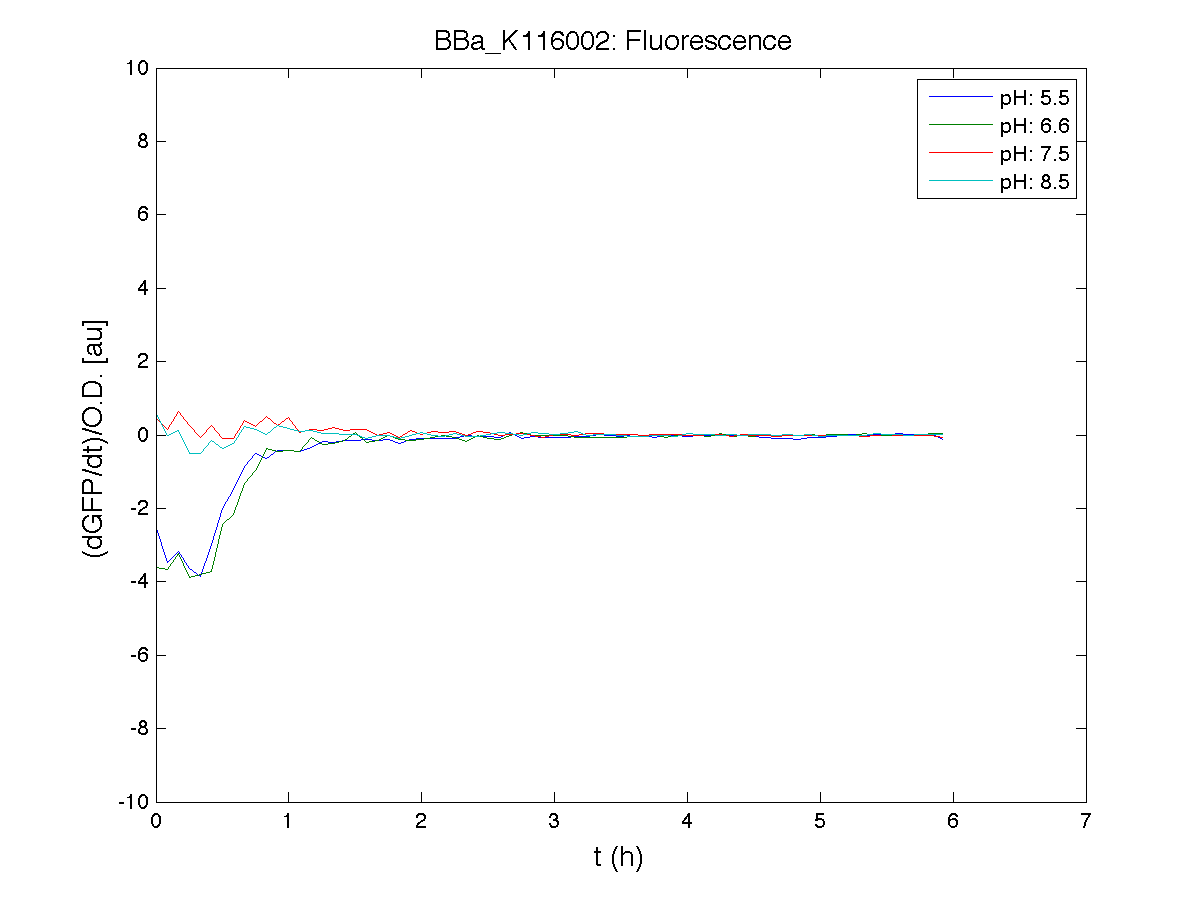
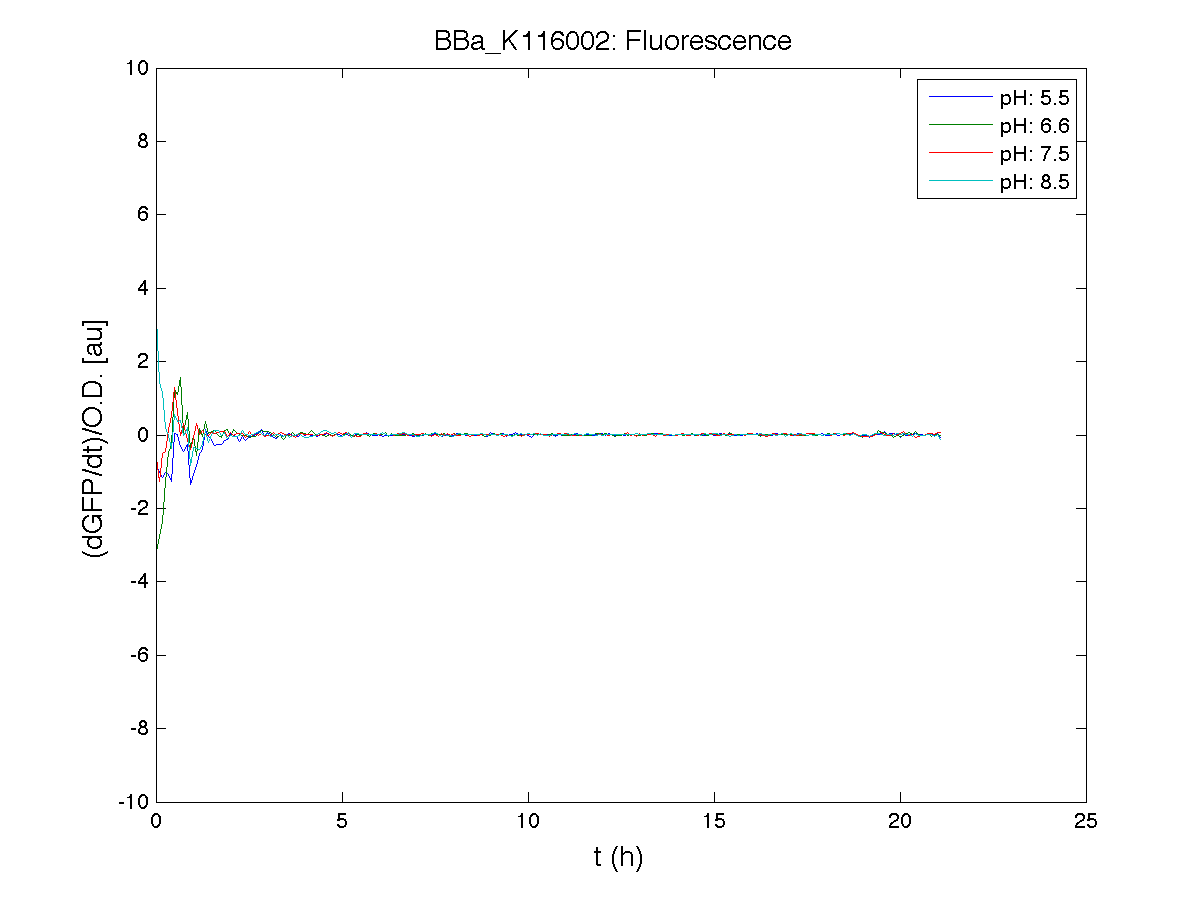
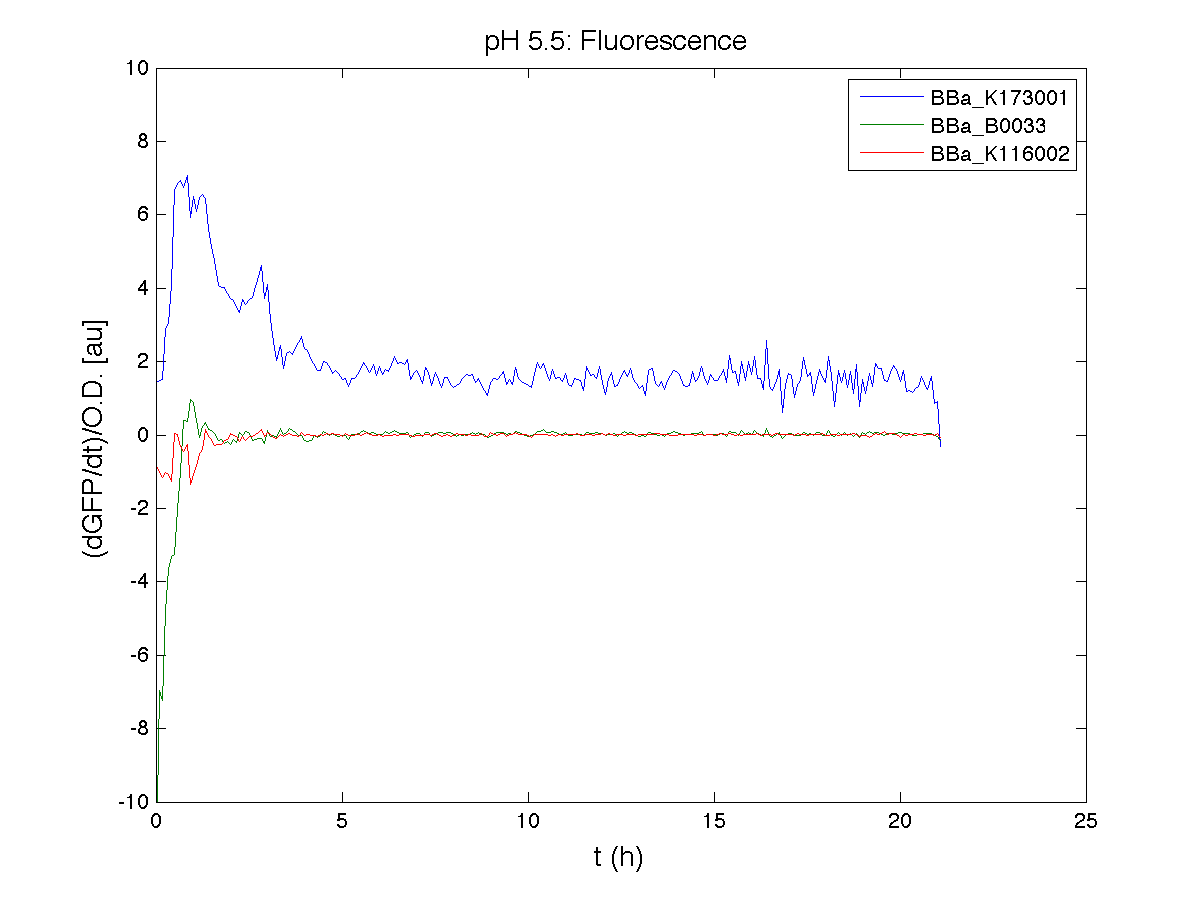
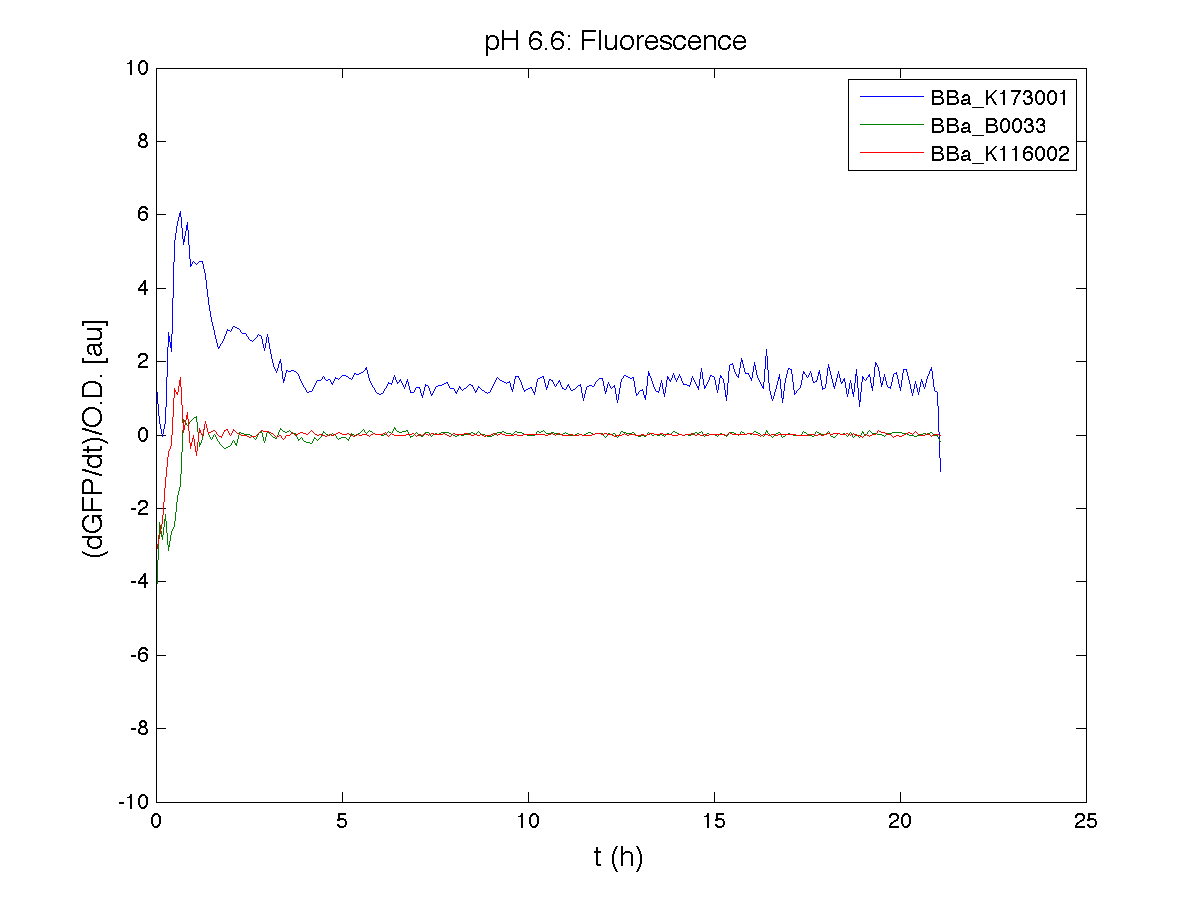
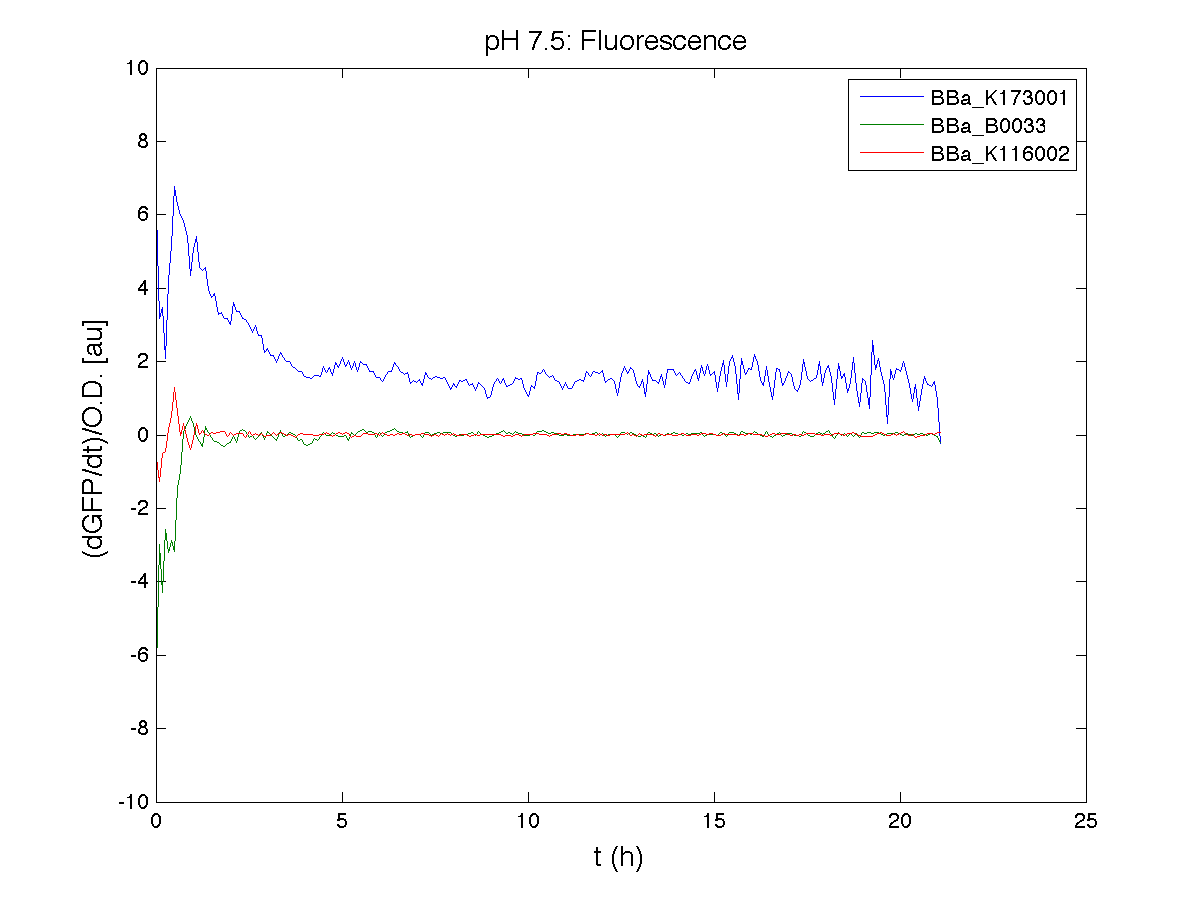
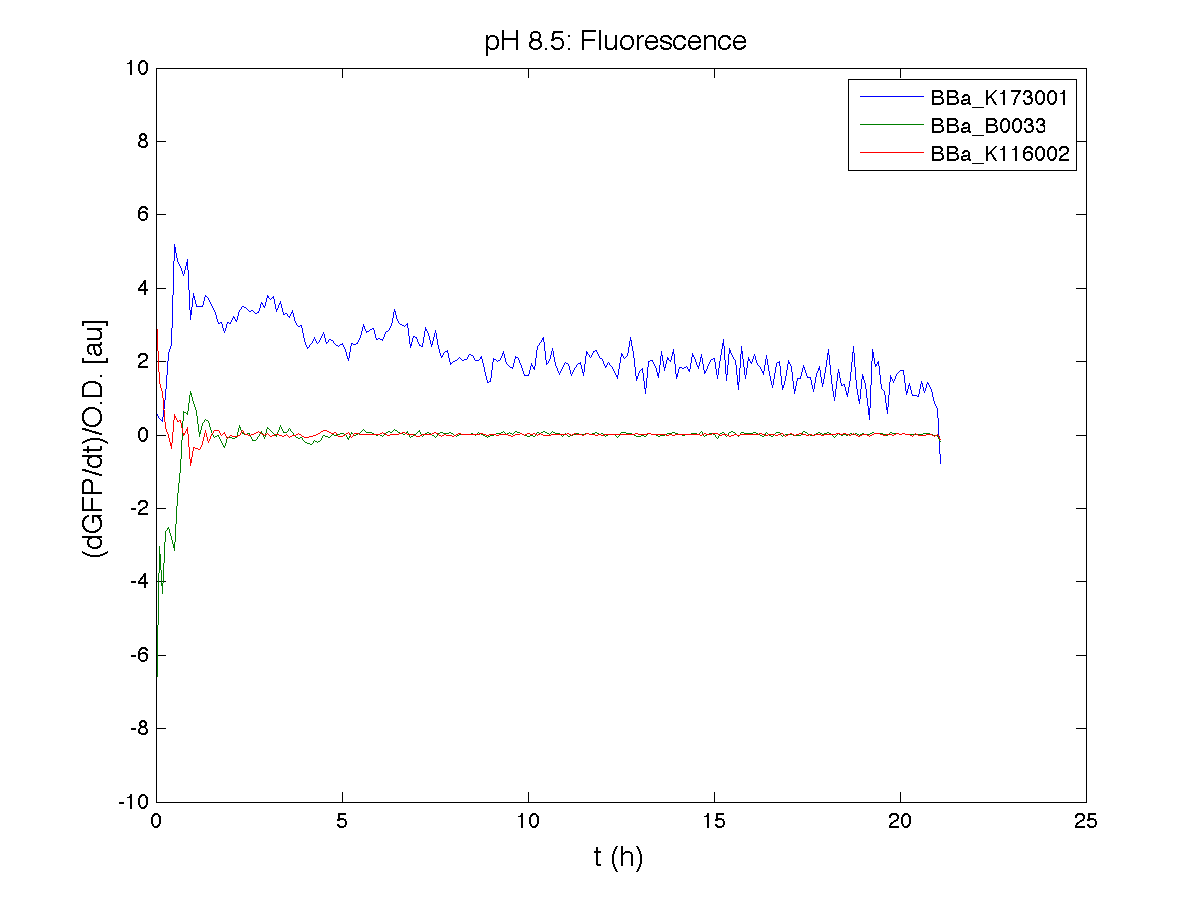
ResultsCommentsAs expected BBa_K116001 didn't produce any GFP. So we can consider it a Na+ sensor and only secondarily a pH sensor. Experiment Na+ 250mMMotivationWe’ll try again to make E.coli producing GFP at the variation of pH. Methods
ResultsCommentsWe didn't expect this. After looking better for a motivation in some articles ([Rachel Karpel et al.]) we think this could be because of the E.coli strain: we use TOP10 while a special strain (delta-pump) without some membrane proteins that regulate E.coli homeostasis is used in other experiments. Final considerationsIn our opinion this sensor (primarily sodium sensor and secondarily pH sensor) needs very particular conditions to work (first of all a specific bacterial strain) we couldn’t reproduce, so we consider it almost unusable. BBa_K112808 - Enterobacteria phage T4 Lysis DeviceBBa_R0011 - Plac hybrid promoterDescriptionThe hybrid lac promoter (BBa_R0011) has been designed taking the Plambda promoter (BBa_R0051) and substituting its cI (BBa_C0051) binding sites with two lacI binding sites. This promoter can be repressed by lacI (BBa_C0012), which can be repressed by lactose or IPTG, providing a lactose/IPTG inducible system. Differently from wild type lac promoter, this part does not have any CAP binding sites, so its behaviour is glucose-independent. Even if lacI is not expressed in this BioBrick, strains bearing a genomic copy of lacI can repress this promoter, which acts as a glucose-independent lactose/IPTG sensor. In the other strains BBa_R0011 acts as a constitutive promoter. Here we provide the characterization of this promoter in E. coli TOP10, which has a lacI genomic copy, constitutively expressed in a weak manner. The data below are referred to , which is the measurement system of . Characterization
Conclusions
Growth conditionsFor every culture we tested:
Data analysisGrowth curvesThe presented growth curves have all been processed as OD600_culture-OD600_broth for each time sample. OD600_broth is the medium in the same conditions as in the culture (e.g. induced with the same inducer concentration as in the culture). Doubling timeThe natural logarithm of the growth curves (processed according to the above section) was computed and the linear phase (corresponding to the bacterial exponential growth phase) was isolated by visual inspection. Then the linear regression was performed in order to estimate the slope of the line m. Finally the doubling time was estimated as d=ln(2)/m [minutes]. In the case of multiple growth curves for a strain, the mean value of the processed curves was computed for each time sample and then this procedure was performed. Relative Promoter Units (RPUs)The RPUs are standard units proposed by Kelly J. et al., 2008, in which the transcriptional strength of a promoter can be measured using a reference standard, just like the ground in electric circuits. RPUs have been computed as: in which:
RPU measurement has the following advantages:
The hypotheses on which RPU theory is based can be found in Kelly J. et al., 2008, as well as all the mathematical steps. From our point of view, the main hypotheses to satisfy are the following:
Materials
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"