Team:Kyoto/GSDD/Abstract
From 2009.igem.org
Project1 -- Gene Switch Depending on Duplication --
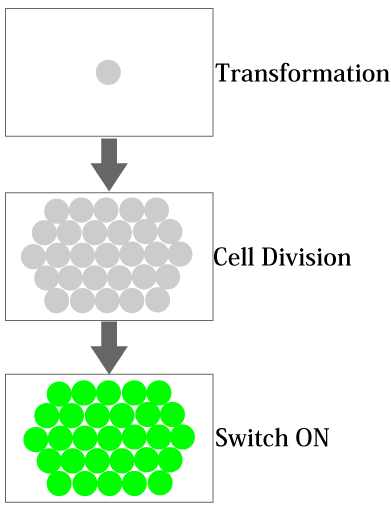
In biotechnology, it is a big challenge to control the temporal gene expression depending on the cellular state. In this GSDD (Gene Switch Depending on Duplication) project, we aimed to design and construct a time controllable gene expression system depending on the number of cell division after transformation. For instance, such a system could be used to engineer the bacteria that penetrate through human cells for therapy; we can remove the bacteria from human body at desired time and cell population. In other words, we could control the life span of bacteria, by expressing a desired gene (e.g., a killer gene) by counting the number of cell division (Figure 1).
To achieve this goal, we came up with the idea that “liner DNA” could be used as “a timer vector “ that expresses a repressor protein depending on the cell division. Liner DNA is replicated incompletely because of “the end replication problem (REFERENCE)”. Therefore, liner DNA becomes shorter and shorter during duplication. After certain period of time, the coding region of a repressor gene will be removed. Thus, the gene (e.g., a killer gene) coded in the other plasmid vector (e.g., a bomb vector) controlled by the repressor will be activated to regulate cell fate (Figure 2).
The problem in this idea is that liner DNA is not stable in cell because of the attack of exonuclease and DNA-repair enzymes. To solve this problem, we engineered the liner DNA by introducing the multiple protein binding sites (e.g., Lac I binding site) at the both end of liner DNA (Figure 2). In this idea, the tight and specific interaction between liner DNA and its binding protein reduces the activity of the exonucleases. Thus, liner DNA becomes shorter and shorter step by step, until the repetitive protein binding sites are lost during duplication. After the removal of the repetitive binding sequences, the exonuclease degrades liner DNA quickly, and therefore the expression of the repressor protein will be repressed. We aimed to construct such a cell division-dependent gene expression system using yeast Saccaromyces cerevisiae. Figure 2 is the outline of this system.
 "
"