Team:EPF-Lausanne/Results
From 2009.igem.org
m (→Fusion of the LOV domain and the trpR DNA-binding domain) |
m (→Fusion of the LOV domain and the trpR DNA-binding domain) |
||
| Line 41: | Line 41: | ||
<br> | <br> | ||
* @MET11: we clearly see the the j-alpha helix is not aligned with the helix of the trpR. It is also much longer than in the LovTAP. As we know that the change in the chromophore induced a change in the the j-alpha helix relatively to the beta-sheet of the LOV, we can imagine the j-alpha helix is not well positioned. | * @MET11: we clearly see the the j-alpha helix is not aligned with the helix of the trpR. It is also much longer than in the LovTAP. As we know that the change in the chromophore induced a change in the the j-alpha helix relatively to the beta-sheet of the LOV, we can imagine the j-alpha helix is not well positioned. | ||
| - | < | + | <br><center> |
| - | <object width="425" height="344"><param name="movie" value="http://www.youtube.com/v/1LlLiJkuUzc&hl=fr&fs=1&"></param><param name="allowFullScreen" value="true"></param><param name="allowscriptaccess" value="always"></param><embed src="http://www.youtube.com/v/1LlLiJkuUzc&hl=fr&fs=1&" type="application/x-shockwave-flash" allowscriptaccess="always" allowfullscreen="true" width="425" height="344"></embed></object></center>< | + | <object width="425" height="344"><param name="movie" value="http://www.youtube.com/v/1LlLiJkuUzc&hl=fr&fs=1&"></param><param name="allowFullScreen" value="true"></param><param name="allowscriptaccess" value="always"></param><embed src="http://www.youtube.com/v/1LlLiJkuUzc&hl=fr&fs=1&" type="application/x-shockwave-flash" allowscriptaccess="always" allowfullscreen="true" width="425" height="344"></embed></object></center><br> |
* @ALA12: same remarks as for the previous. Furthermore, it is clear that LOV is in interaction with bound DNA. | * @ALA12: same remarks as for the previous. Furthermore, it is clear that LOV is in interaction with bound DNA. | ||
| - | < | + | <br><center><object width="425" height="344"><param name="movie" value="http://www.youtube.com/v/FQ-xXnKAYEE&hl=fr&fs=1&"></param><param name="allowFullScreen" value="true"></param><param name="allowscriptaccess" value="always"></param><embed src="http://www.youtube.com/v/FQ-xXnKAYEE&hl=fr&fs=1&" type="application/x-shockwave-flash" allowscriptaccess="always" allowfullscreen="true" width="425" height="344"></embed></object></center></br> |
Revision as of 15:39, 17 October 2009
Contents |
Fusion of the LOV domain and the trpR DNA-binding domain
The first step in our computational study of the LOV domain was to fuse the 2 domains of interest in VMD. We were then able to visualize the different proteins tried by Sosnick. The working protein, that we call LovTAP is the result of the fusion at PHE22 of trpR and can be seen on the next video. The general LOV domain is in yellow. Please note the chromophore called Flavin (FMN) in red in the center of LOV2. The trpR dna binding domain is in orange and DNA in gray.
- @MET11
- @ALA12
- @GLU13
- ...
- @PHE22
- ...
- @LEU25
Clear here to view the other fusions
Equilibration of light and dark state
Here is our first movie from the modeling, showing the behavior of the protein in the dark state condition: Dark State
Dark state
After having modified some parameters in the parameter files, here is our second movie, concerning the light state of the protein this time, with the FMN: Light State
Light state
Analysis
- Maxwell-Boltzmann Energy Distribution
We obtain the following histogramm!
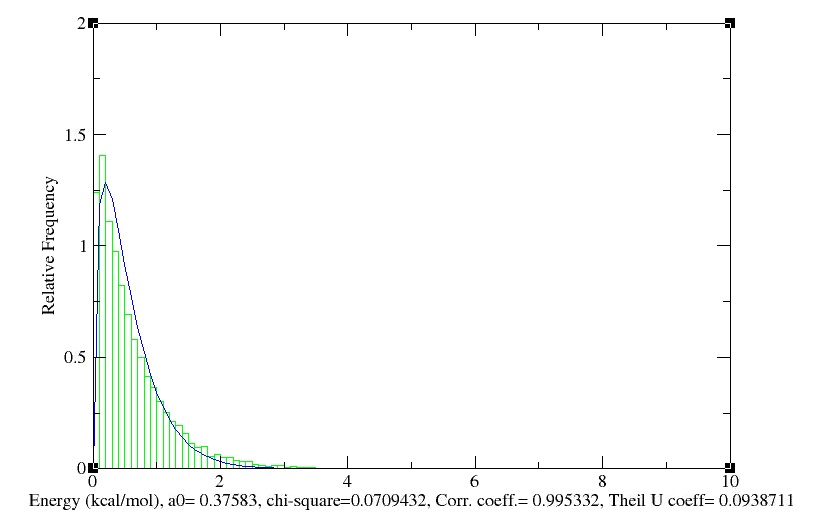
- Temperature
Using EXCEL, we obtain the following graph, which represents the evolution of the temperature in function of time:
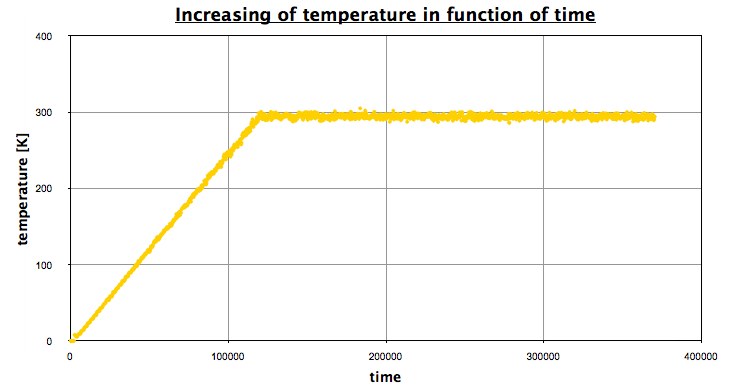
The first part corresponds the the heating, then we let the system reach an equilibrium (NPT state), a NVT portion, and finally a NPT portion again.
- Density
Using EXCEL, we obtain the following graph, which represents the evolution of the density in function of time:
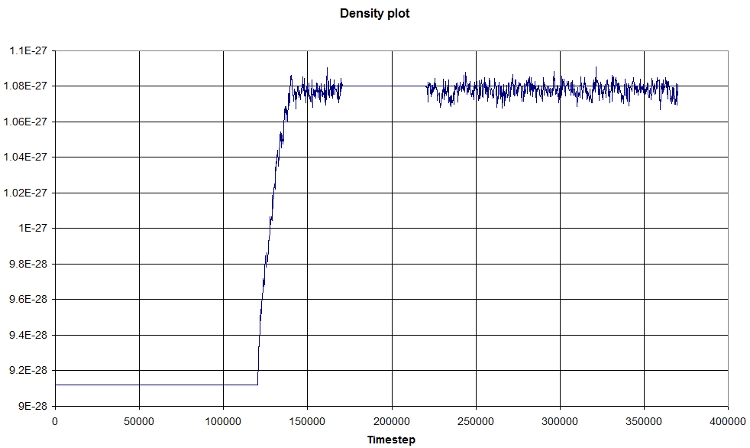
The first part corresponds the the heating, then we let the system reach an equilibrium (NPT state), a NVT portion, and finally a NPT portion again.
- Pressure
Here is a small plot of pressure and temperature in function of time
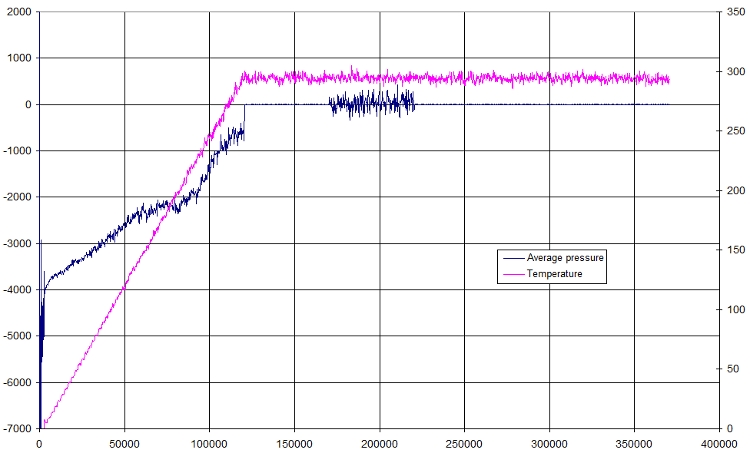
- RMSD
We obtain the following picture:
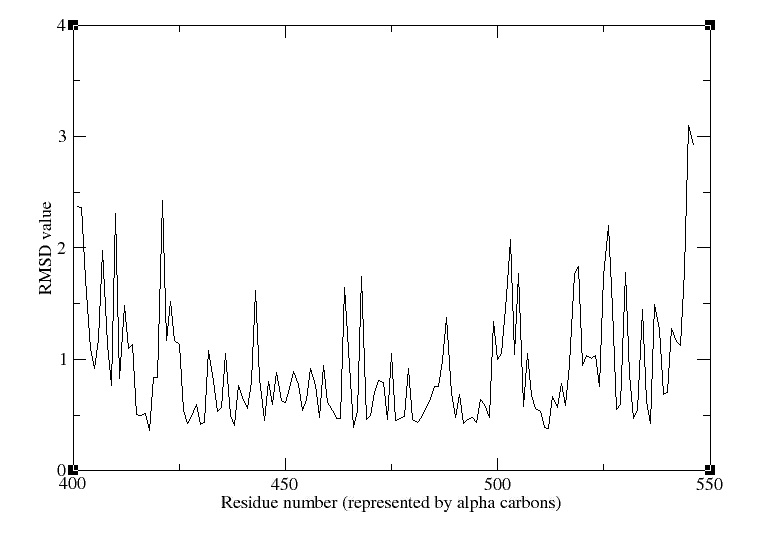
RMSD of residue within 3 angström of the FMN
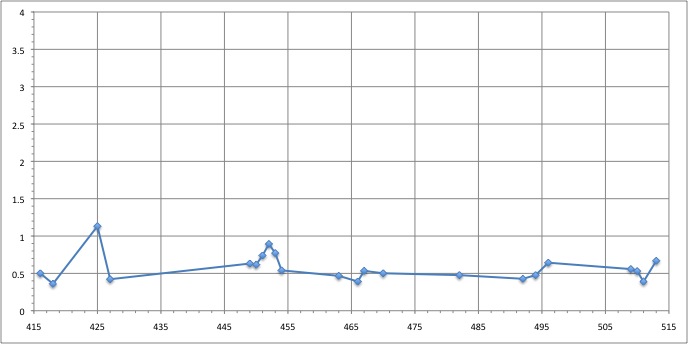
We can see that the residues that move the most are the residue number: 425, 451, 453
RMSD of residue within 6 angström of the FMN
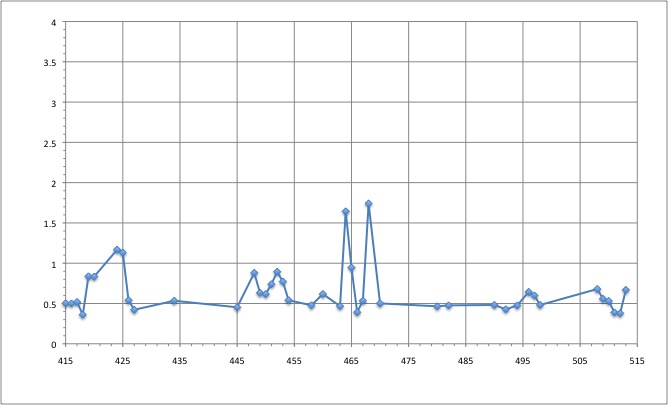
We can see that the residues that move the most are the residue number: 424, 425, 464, 468
- RMSD of selected atoms compared to initial position along time
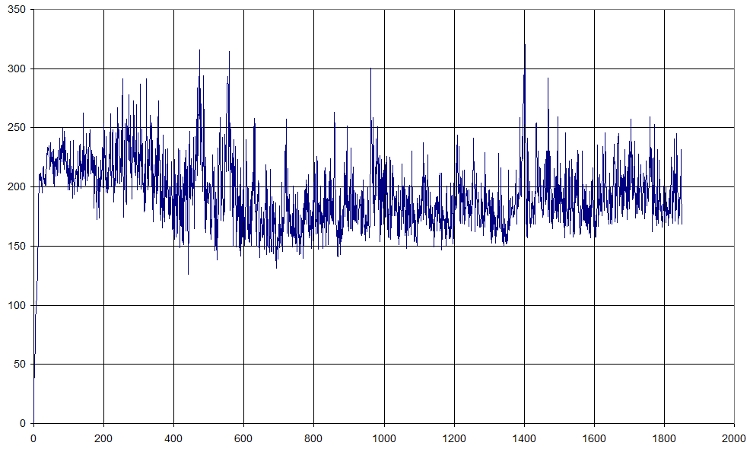
Here is a fast graph of the output of the average RMSD of the atoms in function of time. It seems normal.
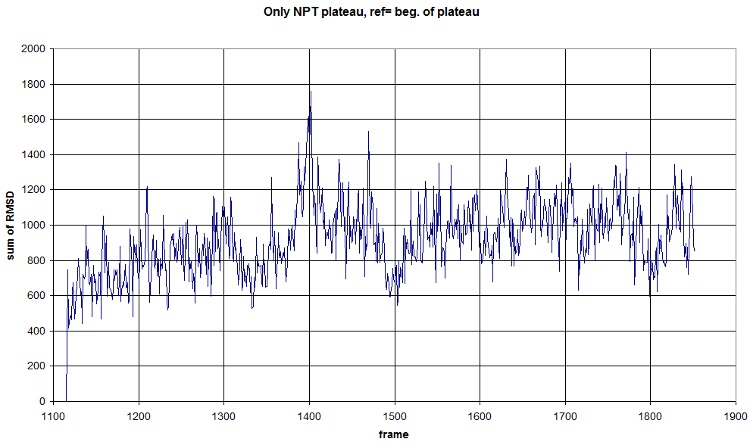
Here is what we got with FIRST_FRAME=1115 REFERENCE_FRAME=1115. Average=921.477, standard deviation=202.1708
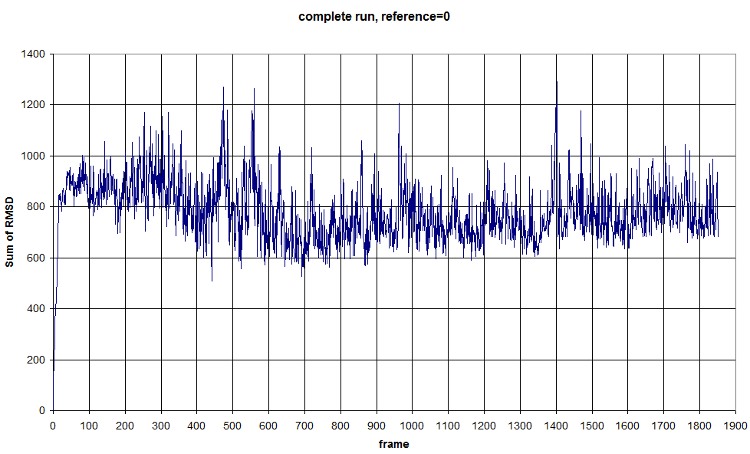
FIRST_FRAME=0 REFERENCE_FRAME=0. The difference of the sum probably comes from the new selection of atoms from the backbone. We should compute an average value to normalize amplitude. (fluctuation is conserved, anyway) Average=781.3913, standard deviation=118.1393
- Salt bridges
Here is a plot for one of the bridges. We have to look for the max distance for a salt bridge.
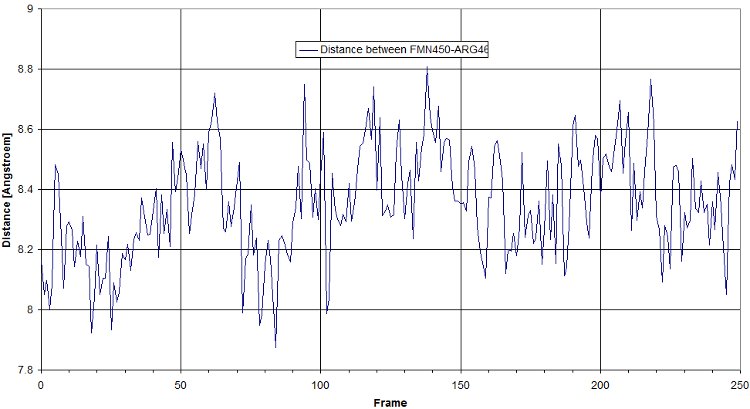
- RMSF
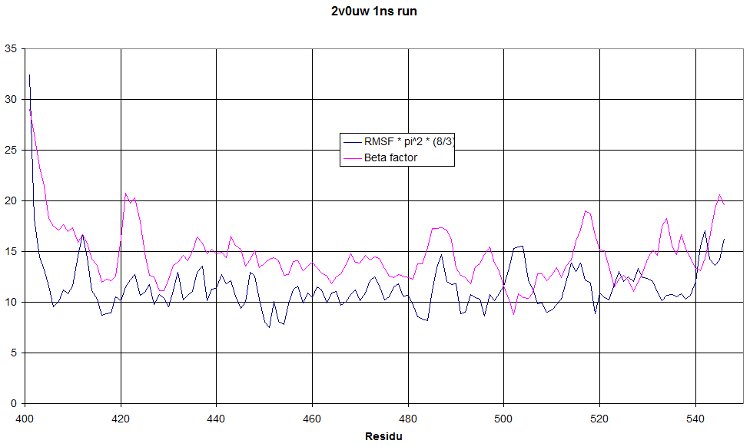
This is a 1 nanosecond NPT run at 300°K. We hope to see a RMSF curve identical to the beta factor. It should only be shifted higher because of the increased temperature. But having a similar tendency would mean our simulation show oscillations similar to what was observed during crystallography. This is really a quite nice validation of our run!
Differential analysis
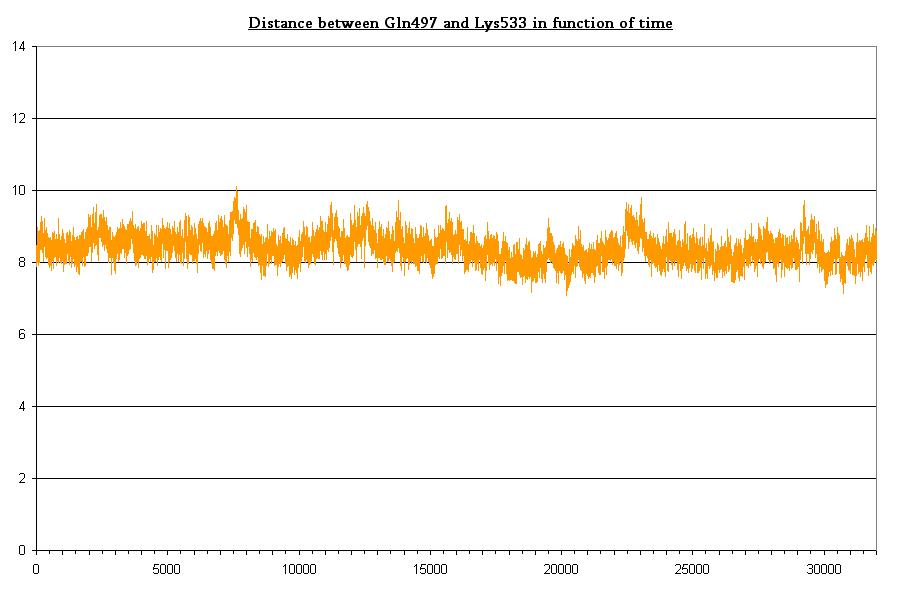
Some useful distances
- Bond between Gln497 and Lys533 in dark state
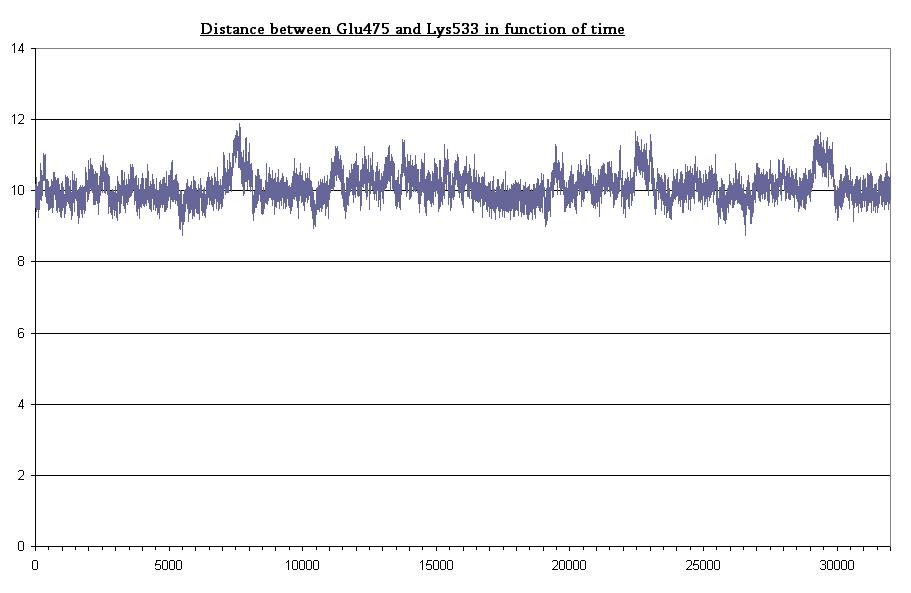
- Bond between Gln475 and Lys533 in dark state
 "
"