Team:EPF-Lausanne/Results
From 2009.igem.org
(→Analysis) |
(→Analysis) |
||
| Line 63: | Line 63: | ||
Here is a small plot of pressure and temperature in function of time | Here is a small plot of pressure and temperature in function of time | ||
[[Image:1st_run.jpg|Run]] | [[Image:1st_run.jpg|Run]] | ||
| + | |||
* <big>RMSD</big> | * <big>RMSD</big> | ||
| Line 69: | Line 70: | ||
[[Image:RMSD_CA_per_res.jpg]] | [[Image:RMSD_CA_per_res.jpg]] | ||
<br> | <br> | ||
| - | |||
<p align="center"><big> <b>RMSD of residue within 3 angström of the FMN</b> </big></p> | <p align="center"><big> <b>RMSD of residue within 3 angström of the FMN</b> </big></p> | ||
| - | |||
[[Image:Resid_3A.jpg]] | [[Image:Resid_3A.jpg]] | ||
<br><br> | <br><br> | ||
| Line 77: | Line 76: | ||
<br><br> | <br><br> | ||
| - | |||
<p align="center"><big> <b>RMSD of residue within 6 angström of the FMN</b> </big></p> | <p align="center"><big> <b>RMSD of residue within 6 angström of the FMN</b> </big></p> | ||
[[Image:Resid_6A.jpg]] | [[Image:Resid_6A.jpg]] | ||
| Line 83: | Line 81: | ||
We can see that the residues that move the most are the residue number: 424, 425, 464, 468 | We can see that the residues that move the most are the residue number: 424, 425, 464, 468 | ||
<br> | <br> | ||
| + | |||
| + | |||
| + | * <big>RMSD of selected atoms compared to initial position along time<big> | ||
| + | Here is a fast graph of the output of the average RMSD of the atoms in function of time. It seems normal. | ||
| + | <br>[[Image:Rmsd.jpg]] | ||
| + | <br><br><br> | ||
| + | |||
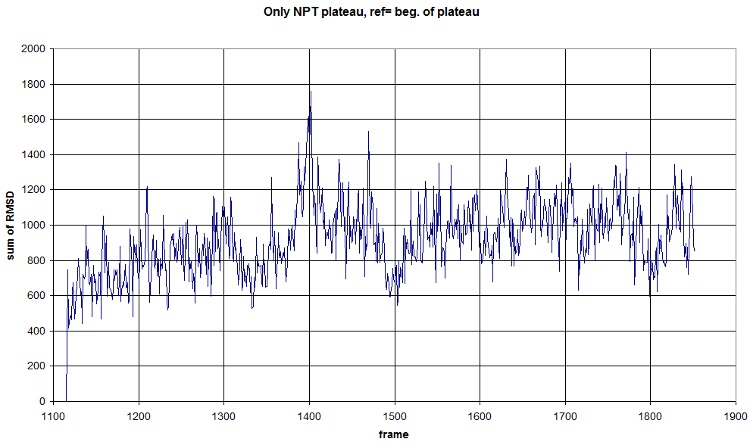
| + | Here is what we got with FIRST_FRAME=1115 REFERENCE_FRAME=1115. Average=921.477, standard deviation=202.1708 | ||
| + | <br>[[Image:RMSD_plateau.jpg]] | ||
| + | <br><br><br> | ||
| + | |||
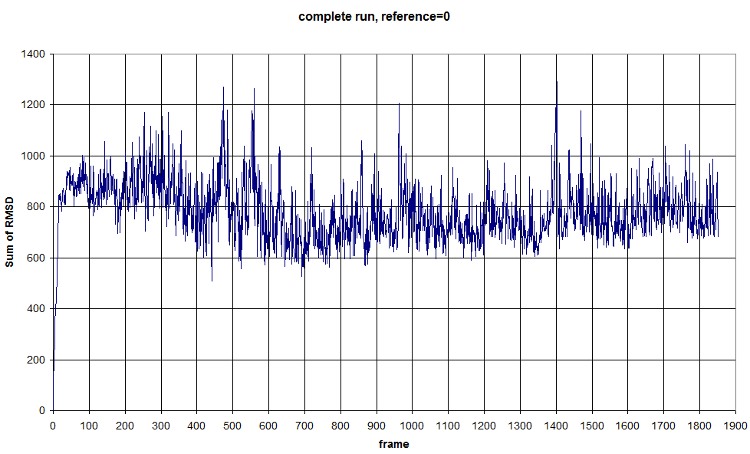
| + | FIRST_FRAME=0 REFERENCE_FRAME=0. The difference of the sum probably comes from the new selection of atoms from the backbone. <b>We should compute an average value to normalize amplitude</b>. (fluctuation is conserved, anyway) Average=781.3913, standard deviation=118.1393 | ||
| + | <br>[[Image:RMSD_COMPLETE_RUN.jpg]] | ||
=Differential analysis= | =Differential analysis= | ||
</div><div CLASS="epfl09bouchon"></div> | </div><div CLASS="epfl09bouchon"></div> | ||
Revision as of 09:58, 21 September 2009
Equilibration of light and dark state
Here is our first movie from the modeling, showing the behavior of the protein in the dark state condition: Dark State
Dark state
After having modified some parameters in the parameter files, here is our second movie, concerning the light state of the protein this time, with the FMN: Light State
Light state
Analysis
- Maxwell-Boltzmann Energy Distribution
We obtain the following histogramm!
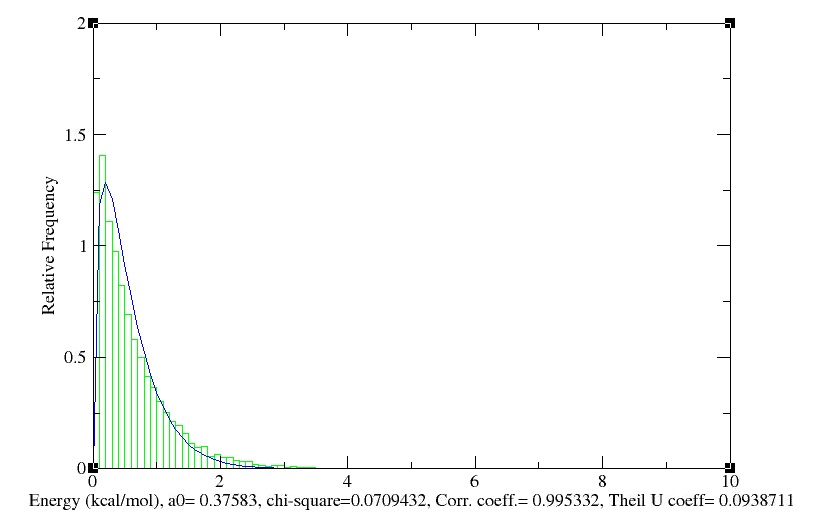
- Temperature
Using EXCEL, we obtain the following graph, which represents the evolution of the temperature in function of time:
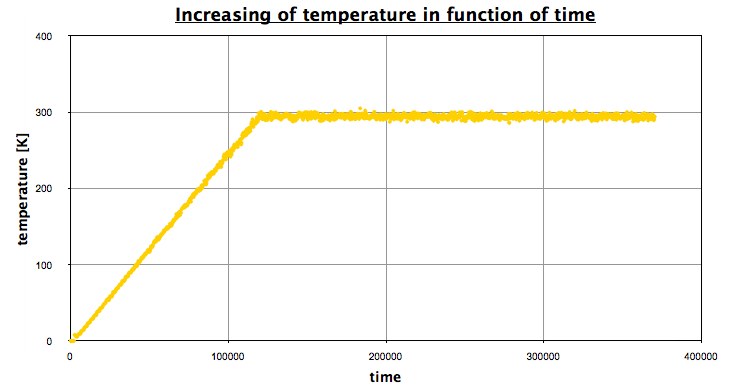
The first part corresponds the the heating, then we let the system reach an equilibrium (NPT state), a NVT portion, and finally a NPT portion again.
- Density
Using EXCEL, we obtain the following graph, which represents the evolution of the density in function of time:
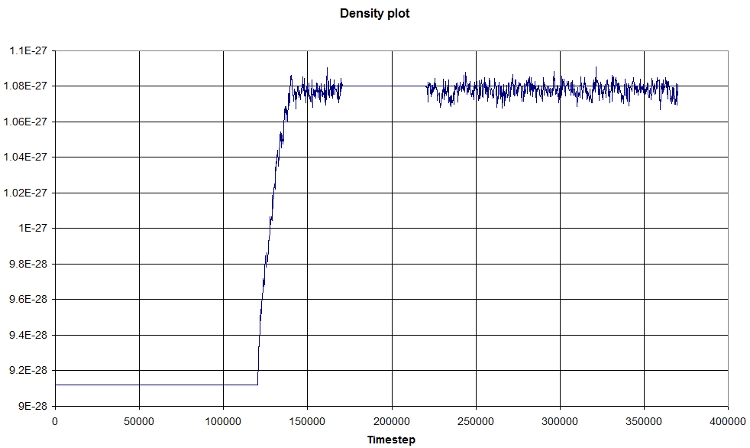
The first part corresponds the the heating, then we let the system reach an equilibrium (NPT state), a NVT portion, and finally a NPT portion again.
- Pressure
Here is a small plot of pressure and temperature in function of time
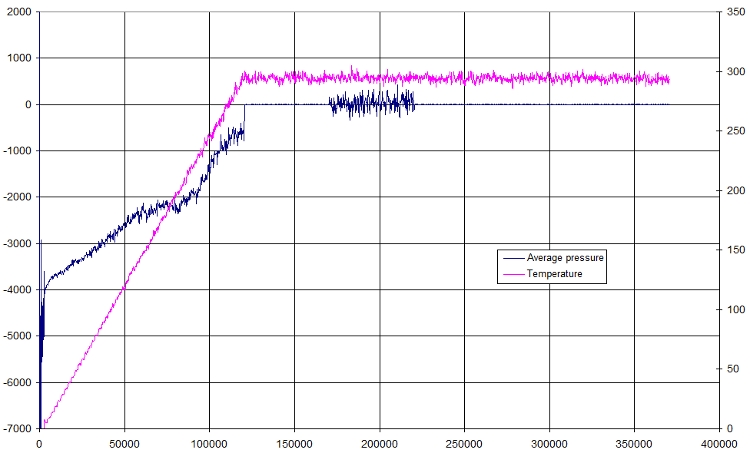
- RMSD
We obtain the following picture:
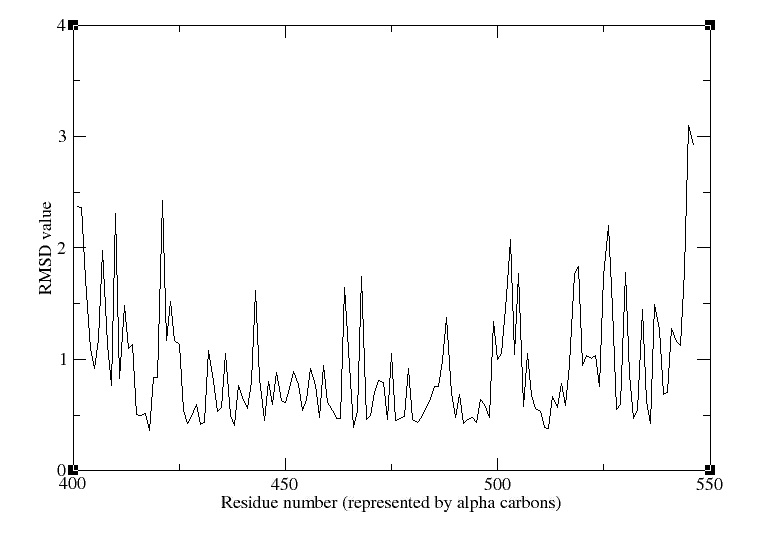
RMSD of residue within 3 angström of the FMN
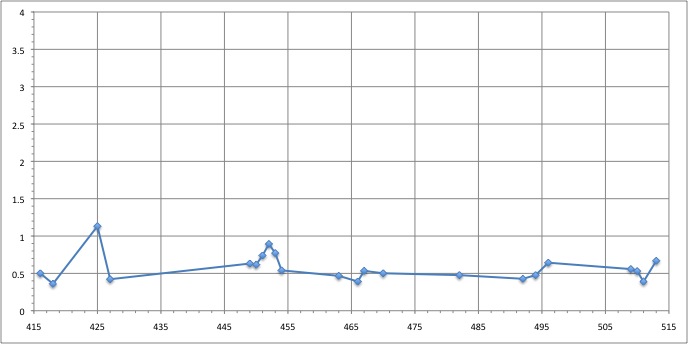
We can see that the residues that move the most are the residue number: 425, 451, 453
RMSD of residue within 6 angström of the FMN
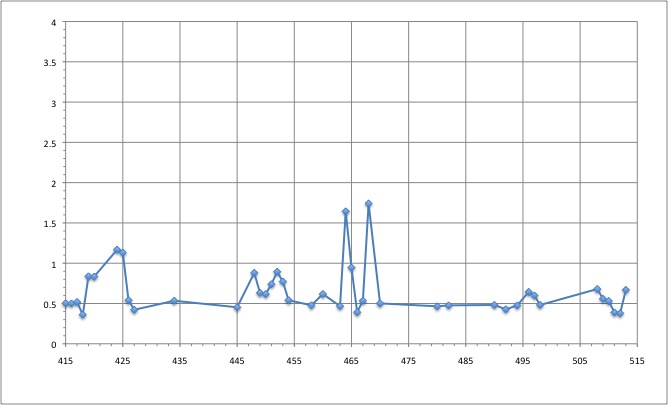
We can see that the residues that move the most are the residue number: 424, 425, 464, 468
- RMSD of selected atoms compared to initial position along time
Here is a fast graph of the output of the average RMSD of the atoms in function of time. It seems normal.
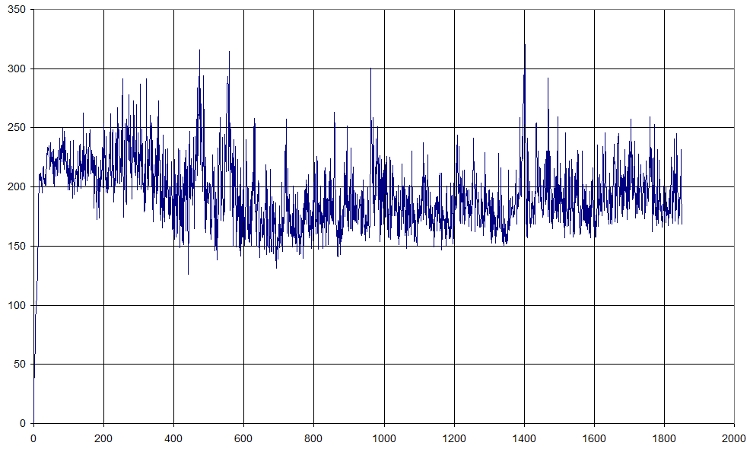
Here is what we got with FIRST_FRAME=1115 REFERENCE_FRAME=1115. Average=921.477, standard deviation=202.1708
FIRST_FRAME=0 REFERENCE_FRAME=0. The difference of the sum probably comes from the new selection of atoms from the backbone. We should compute an average value to normalize amplitude. (fluctuation is conserved, anyway) Average=781.3913, standard deviation=118.1393
Differential analysis
</div>
 "
"
