Team:Newcastle
From 2009.igem.org
https://static.igem.org/mediawiki/2009/9/91/Newcastle3Banner.swf
In 2009 the Newcastle team are tackling environmental issues using Bacillus subtilis. We are a team of eight with wide ranging backgrounds in the fields of Bioinformatics, Computing Science, Chemical Engineering, Genetics and Medical Sciences. This is Newcastle's second year in the iGEM competition; last year our team designed BugBuster, which achieved a Gold Medal.
Project Description
For our BioBricks we are using coding sequences from Bacillus subtilis itself, as well as existing BioBricks from the iGEM parts registry which we intend to further characterise, as well as developing our own BioBricks. We also intend to characterise our own sigmaA promoter library which we believe will be of great use for future iGEM teams.
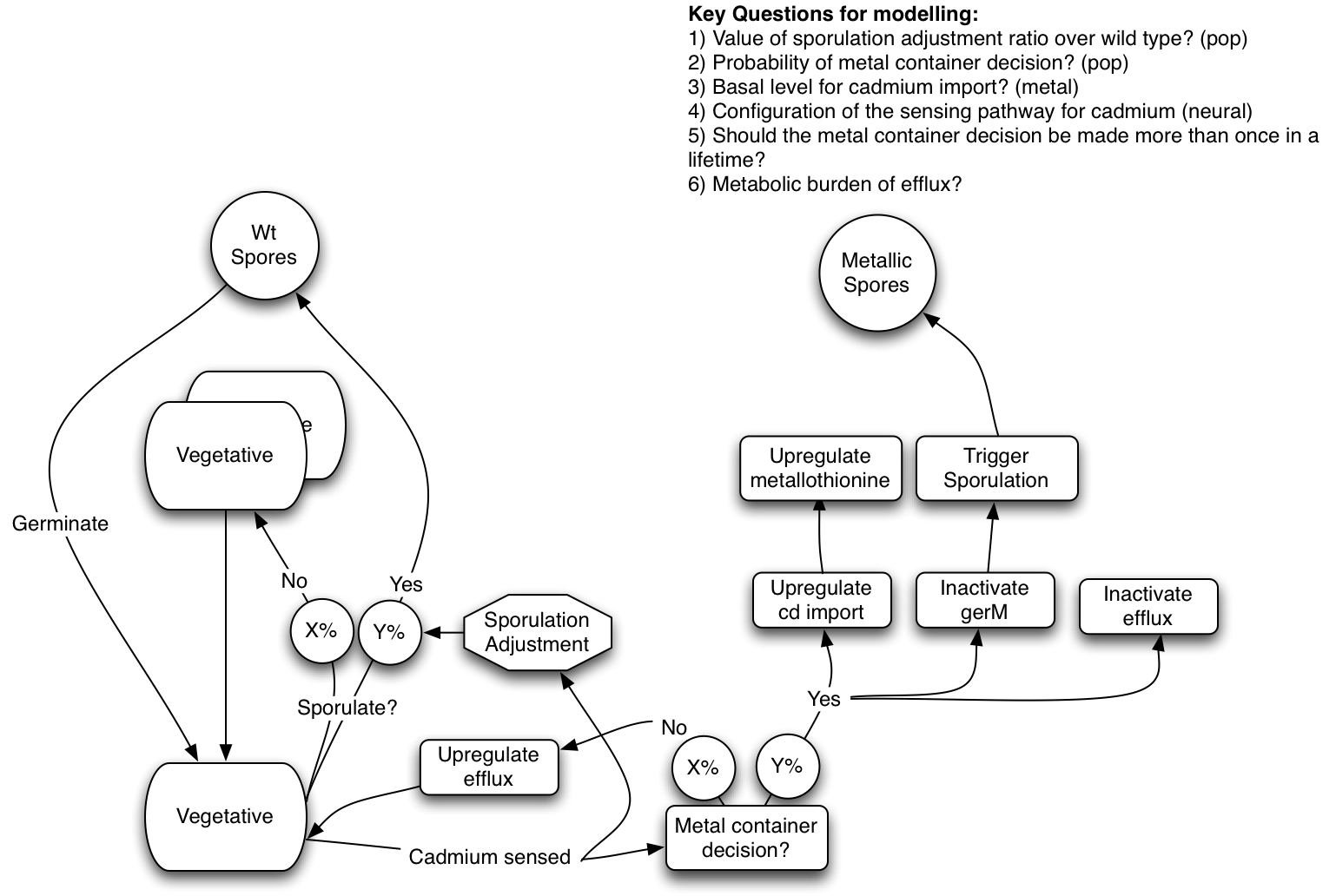
The following diagram gives an overview of what our design hopes to acheive...
Heavy Metal Sensing
Our design is for the bacteria to intake cadmium through the manganese channel MntH, as it has been found that due to the similar size and charge of the ions cadmium leaks through these channels [1].Once inside the cell the cadmium must be sensed in order for the cell to regulate some of the decisions involved in becoming a metal container. The metal sensing proteins we intend to use are yozA (czrA) and arsR which both bind cadmium as well as zinc cobalt and nickel, and arsenic, silver and copper respectively [2].These proteins are repressor proteins which also bind DNA preventing transcription of downstream CDS. The repressor proteins however release the DNA to preferentially bind cadmium [2] allowing transcription to occur. These proteins therefore allow selective sensing of cadmium in the form of an AND gate, a model for which can be seen in our modelling section.
Population Dynamics
As well as sensing cadmium we will attempt to engineer our bacteria’s normal population dynamics, by nudging the natural stochastic sporulation decision in favour of greater sporulation, to account for the spores that will be lost as metal containers. We will also design our own stochastic switch, which will be based on the decision of being a metal container or not. This will be regulated by an invertible segment of DNA using the hin/hix system [3]. In this way our artificial stochastic switch will be a ‘biased heads or tails’ which we can control. This stochastic switch will control expression of our ‘metal sponge’.
Metal Containers
To make our spores metal containers we will be using the metallothionein protein smtA which is a relatively small cysteine rich protein known to ‘soak up’ metals such as cadmium [4]. We will guide this protein to the spore whilst sporulating by creating a fusion with the spore coat protein cotC, which will coat the spore in cadmium bound metallothionein [5]. Finally, to ensure that metal containing spores will not germinate again, releasing their bound cadmium, we will ‘knock-out’ important germination genes using our invertible sequence system.
Novelty
Our design has many components which we believe are novel, such as our control over the sporulation cycle, and our synthetic stochastic switch, both systems we believe are reusable concepts within synthetic biology.
References
- Que, Q. and J.D. Helmann, Manganese homestasis in Bacillus subtilis is regulated by MntR, a bifunctional regulator related to the diphtheria toxin repressor family of proteins. Molecular Microbiology, 2000. 35(6): p. 1454-1468.
- Harvie, D.R., et al., Predicting metals sensed by ArsR-SmtB repressors: Allosteric interference by a non-effector metal. Molecular Microbiology, 2006. 59(4): p. 1341-1356.
- Haynes, K.A., et al., Engineering bacteria to solve the Burnt Pancake Problem. Journal of Biological Engineering, 2008. 2.
- Blindauer, C.A., et al., Multiple bacteria encode metallothioneins and SmtA-like zinc fingers. Molecular Microbiology, 2002. 45(5): p. 1421-1432.
- Mauriello, E.M.F., et al., Display of heterologous antigens on the Bacillus subtilis spore coat using CotC as a fusion partner. Vaccine, 2004. 22(9-10): p. 1177-1187.
External links
- [http://www.ncl.ac.uk Newcastle University]
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
- Newcastle iGEM Twitter
- [http://www.facebook.com/home.php#/group.php?gid=131709337641 Newcastle on Facebook]
- [http://www.youtube.com/user/newcastle2009igem Newcastle Youtube Channel]
 "
"