Team:EPF-Lausanne/LOVTAP Results
From 2009.igem.org
(→Characterization of Read-Out n°1) |
(→Characterization of Read-Out n°1) |
||
| Line 16: | Line 16: | ||
To characterize this biobrick, we cultured the cells in a medium without tryptophane: M9 medium to which we added all other amino acids and thiamine. You can find the protocol for the M9 medium [https://2009.igem.org/wiki/index.php?title=Team:EPF-Lausanne/Protocols/M9medium here]. The purpose of the experiment was to see the difference of RFP expression depending on whether we added tryptophane to the medium or not. | To characterize this biobrick, we cultured the cells in a medium without tryptophane: M9 medium to which we added all other amino acids and thiamine. You can find the protocol for the M9 medium [https://2009.igem.org/wiki/index.php?title=Team:EPF-Lausanne/Protocols/M9medium here]. The purpose of the experiment was to see the difference of RFP expression depending on whether we added tryptophane to the medium or not. | ||
| + | |||
| + | |||
<font size="3">'''Protocol'''</font> | <font size="3">'''Protocol'''</font> | ||
| Line 29: | Line 31: | ||
| - | '''Expected Results''' | + | <font size="3">'''Expected Results'''</font> |
In this experiment, we would expect to see a slight increase in fluorescence for the cells without tryptophane (simply due to cell growth), compared to a decrease in fluorescence for the samples in which we added tryptophane, since tryptophane would enhance the activity of the tryptophane repressor, which in turn would repress the RFP gene via the Trp promoter. Also to be noted, the RFP decay shouldn't be too abrupt, considering that the RFP protein used doesn't contain a degradation tag. | In this experiment, we would expect to see a slight increase in fluorescence for the cells without tryptophane (simply due to cell growth), compared to a decrease in fluorescence for the samples in which we added tryptophane, since tryptophane would enhance the activity of the tryptophane repressor, which in turn would repress the RFP gene via the Trp promoter. Also to be noted, the RFP decay shouldn't be too abrupt, considering that the RFP protein used doesn't contain a degradation tag. | ||
| Line 35: | Line 37: | ||
| - | '''Graphs & Figures''' | + | <font size="3">'''Graphs & Figures'''</font> |
| Line 43: | Line 45: | ||
| - | '''Analysis & Conclusions''' | + | <font size="3">'''Analysis & Conclusions'''</font> |
We can see that there is a decrease in RFP expression when Trp is added to the medium versus a stable expression for the -Trp condition, which corresponds to what we expected. From this we can therefore conclude that our first read-out system is functional, confirming that the sequence for the Trp promoter seems to be correct. | We can see that there is a decrease in RFP expression when Trp is added to the medium versus a stable expression for the -Trp condition, which corresponds to what we expected. From this we can therefore conclude that our first read-out system is functional, confirming that the sequence for the Trp promoter seems to be correct. | ||
Revision as of 19:29, 20 October 2009
Contents |
Results
Characterization of Read-Out n°1
Our first read-out system consists of the Trp promoter followed by the RFP gene. In E. coli, the Trp promoter is situated in front of the Trp operon, which contains the genes necessary for the tryptophane biosynthetic pathway. In brief, genes placed after the Trp promoter should be repressed in the presence of tryptophane.
To characterize this biobrick, we cultured the cells in a medium without tryptophane: M9 medium to which we added all other amino acids and thiamine. You can find the protocol for the M9 medium here. The purpose of the experiment was to see the difference of RFP expression depending on whether we added tryptophane to the medium or not.
Protocol
- Cells containing the read-out system were cultured overnight in M9 medium.
- On the next day, re-inoculated 0.75 mL of cell culture into 25 mL of fresh medium in an Erlenmeyer flask. Incubated for 2 hours at 37°C.
- Normalized the OD of all flasks to 0.06 by adding the appropriate amount of fresh medium.
- Prepared the plate for the measurements with the qPCR machine, 50 ul per well. For the tryptophane, we used a 4.25 g/L solution. For ATC, it was a 500 ng/mL solution in 50% EtOH, and for IPTG, we always added it so as to have a final concentration of 1mM. The conditions we tested were 1/2 Trp, 1 Trp and 3/2 Trp, 1/2 corresponding to 1 ul.
- The plate was then loaded into the qPCR machine, which took measurements of red fluorescence every 3 minutes, giving the graphs you can see below.
Please note that there was a time lag between when we started loading the wells on the plate and when the measurements began, so the induction might already have started.
Expected Results
In this experiment, we would expect to see a slight increase in fluorescence for the cells without tryptophane (simply due to cell growth), compared to a decrease in fluorescence for the samples in which we added tryptophane, since tryptophane would enhance the activity of the tryptophane repressor, which in turn would repress the RFP gene via the Trp promoter. Also to be noted, the RFP decay shouldn't be too abrupt, considering that the RFP protein used doesn't contain a degradation tag.
Graphs & Figures
Analysis & Conclusions
We can see that there is a decrease in RFP expression when Trp is added to the medium versus a stable expression for the -Trp condition, which corresponds to what we expected. From this we can therefore conclude that our first read-out system is functional, confirming that the sequence for the Trp promoter seems to be correct.
Characterization of Read out 2
For RO2 :
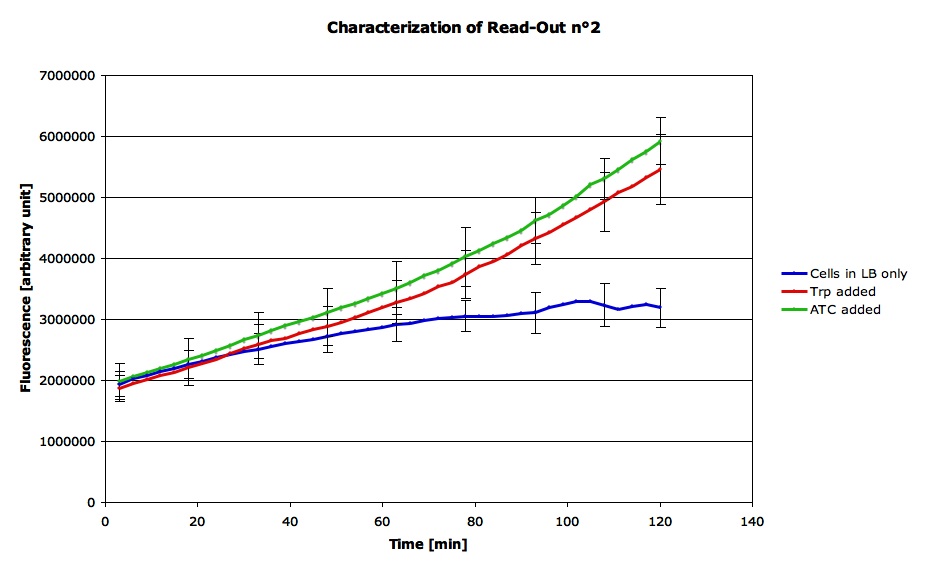
This experiment was done in LB medium. All cells were normalized to an OD's of 0.2 before adding inducers which were put into the liquid culture just before the measurements. The concentration of Tryptophan was 0.9 mg/ml and the concentration of Anhydrous Tetracyclin was of 50 ug/ml.
Here are the result from the machine (qPCR). We did the average signal for each condition as we had 4 measurements for each state :
This confirms that our construct responds both to Trp and ATC and shows the characteristics we expected.Therefore, we can suppose that both the Trp op and the Tet op are functional and that we can use this part in order to see if the LovTAP is functional.
Characterization of the entire system
Now that we have characterized the read out systems, we were able to see if the LovTAP protein would show the same pattern of response when illuminated as when we added Trp to the medium.
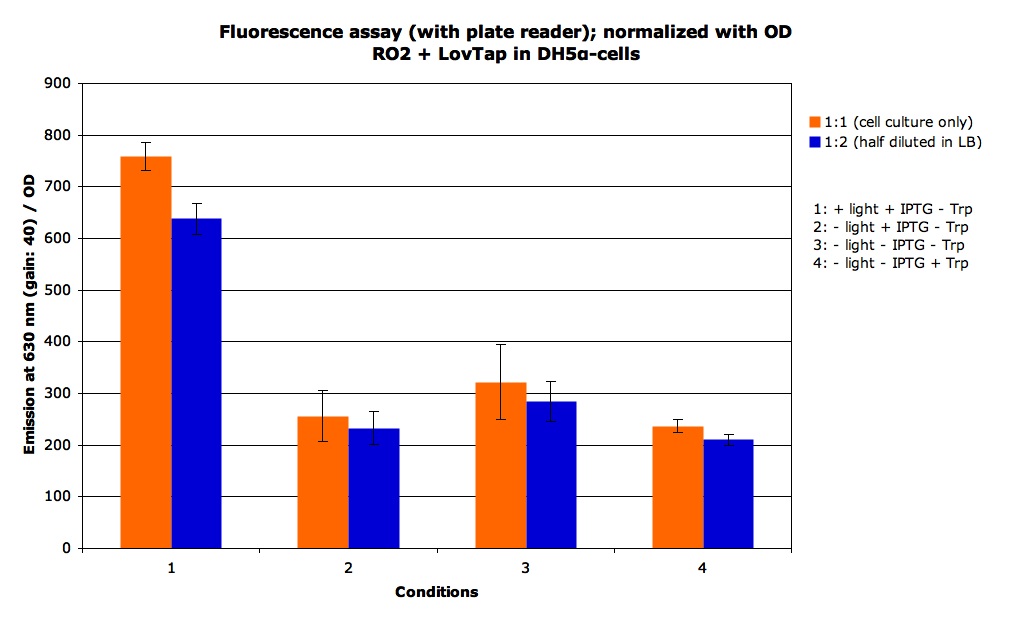
We some static measurement of fluorescence. For the + light condition we exposed the cells to the light for 2 hours, we got the following results
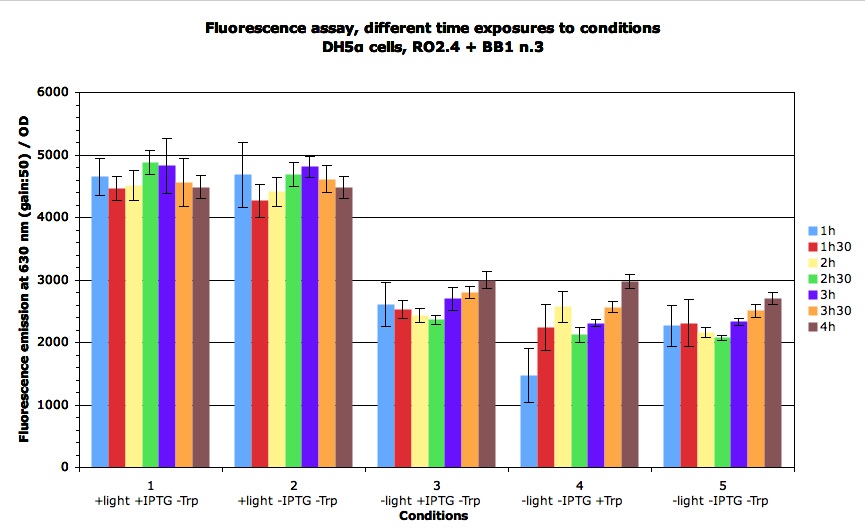
Then we wanted to see the time course of the induction : when is the RFP expressed, how etc.
It appears that this actually happens during the first hour.
Mutations
We used two different mutations in our project.
First we used some E. coli strains that had their Trp promoter mutated.
Then we used some strains we modified to test the results we had in the modeling session.
 "
"