Team:Cambridge/Notebook/Week4
From 2009.igem.org
Categories :
Project :
-
Overview
Sensitivity Tuner
--- Characterisation
--- Modelling
Colour Generators
--- Carotenoids (Orange/Red)
--- Melanin (Brown)
--- Violacein (Purple/Green)
The Future
Safety
Notebook :
Team Logistics :
Week 4 - Development
Design and transformation of cells with activator/promoter constructs - started assembling carotenoid constructs - MelA plan added to the registry - troubleshooting of restriction digests - decision made to synthesise the vio gene
Monday
Wet Work
Melanin
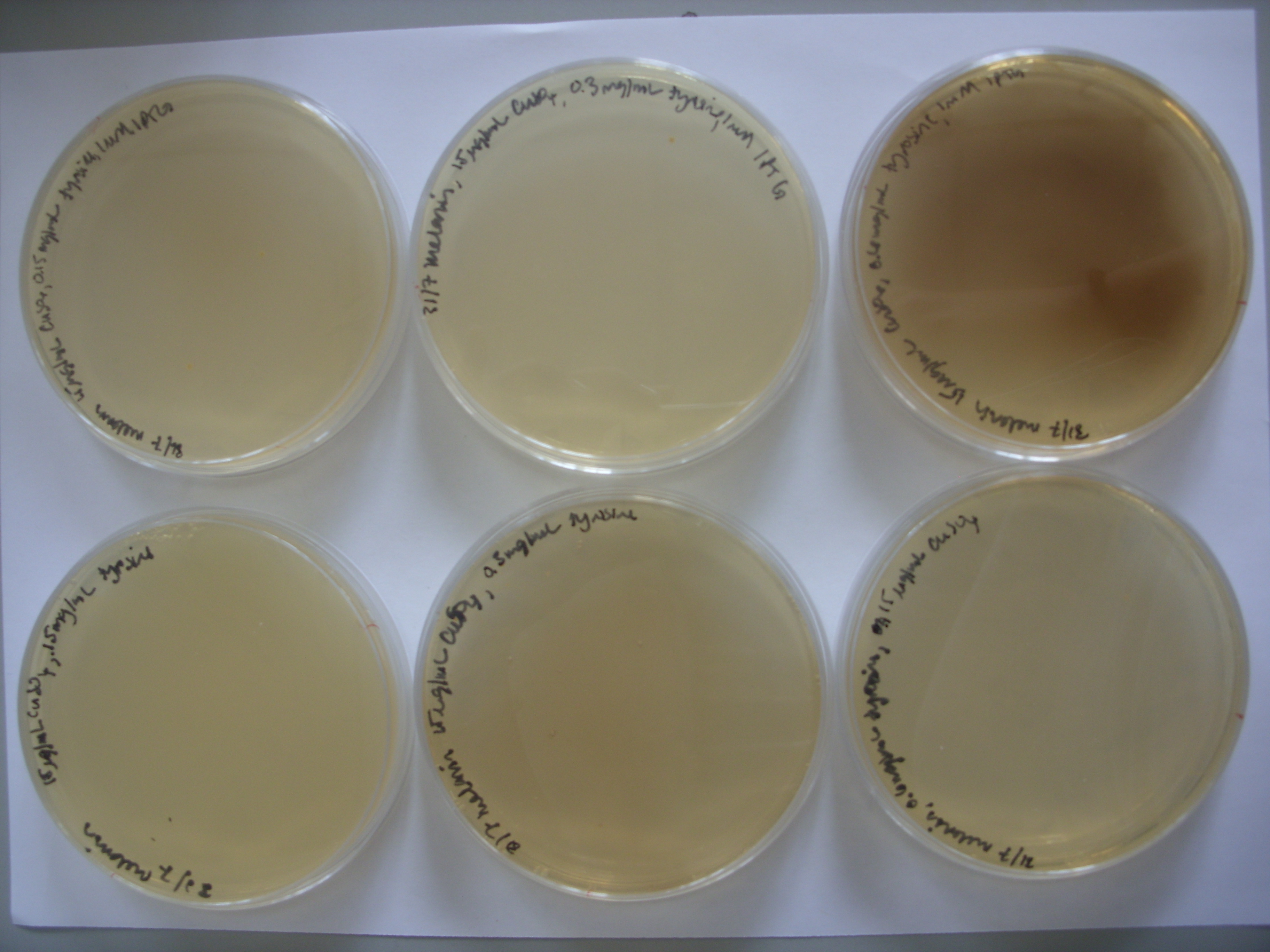
- Results from weekend plates:
From left to right, top row: 1mM IPTG with 0.075 mg/mL tyrosine, 1mM IPTG and 0.3 mg/mL tyrosine, and then 1mM IPTG and 0.6 mg/mL tyrosine From left to right, bottom row: 0.075 mg/mL tyrosine, 0.3 mg/mL tyrosine, and then 0.6 mg/mL tyrosine.
It appears that the bacteria were much too high a concentration; they formed lawns. As we saw on the first plates, pigment production is reduced at high bacterial concentrations. However, it is encouraging that the darkest plate is the one with the greatest tyrosine concentration and IPTG.
To follow up, the experiment was repeated with 1/100 and 1/1000 dilutions of bacterial culture on the following plates:
100ug/ml Ampicillin, 15 ug/ml CuSO4, 0.3 mg/mL tyrosine
100ug/ml Ampicillin, 15 ug/ml CuSO4, 0.6 mg/mL tyrosine
100ug/ml Ampicillin, 7.5 ug/ml CuSO4, 0.075 mg/mL tyrosine, 0.5mM IPTG
100ug/ml Ampicillin, 15 ug/ml CuSO4, 0.3 mg/mL tyrosine, 1mM IPTG
100ug/ml Ampicillin, 15 ug/ml CuSO4, 0.6 mg/mL tyrosine, 1mM IPTG
- Unfortunately, the overnight cultures for the plate readers were contaminated, so a single colony was once again incubated overnight in 10 mL of 100ug/ml Ampicillin, 15 ug/ml CuSO4, 0.3 mg/mL tyrosine, 1mM IPTG in preparation for pigment characterisation using the plate reader.
Violacein
As described in the resource cited below, violacein can be extracted using butanol. A single colony was incubated overnight in LB with trimethoprim. We hope to achieve pigment production in liquid cultures and then extract pigment later in the week.
P.R. August, T.H. Grossman, C. Minor, M.P. Draper, I.A. MacNeil, J.M. Pemberton, K.M. Call, d. Holt, and M. S. Osbourne, Sequence Analysis and Functional Characterization of the Violacein Biosynthetic Pathway from Chromobacterium violaceum, J. Mol. Microbiol. Biotechnol. (2000) 2(4): 513-519. [http://www.horizonpress.com/jmmb/v2/v2n4/26.pdf]
Amplification
- Chose 4 activator / promoter combinations to continue work with:
- P2 activator with Psid promoter (I746374)
- P2 activator with PLL promoter (I746375)
- PSP3 activator with PF promoter (I746380)
- phiR73 with PO promoter (I746391)
- Of the above, on plates all showed leaky expression of GFP and RFP, suggesting that last weeks transformation was successful.
- In preparation for a plate reader assay tomorrow, incubated single colonies overnight of the above constructions as well as the following controls
- I3540 (pBad and GFP) - taken from 2007 glycerol stocks
- I3620 (pBad and RFP) - taken from 2007 glycerol stocks
- untransformed Arabinose strain colony
- Plate reader assay will have triple replicates of all of the above constructs induced with arabinose at various concentrations.
Progress on carotenoids (Monday)
We attempt to start assembling individual Biobricks (K118014/CrtE, K118006/CrtB and K118014/CrtI) into a composite construct (CrtEBI), which is expected to produce reddish/pink lycopene.
We have access to Fermentas FastDigest enzymes. XbaI, SpeI and PstI were used in pair to carry out double digest. In our preliminary trial, the following combinations were used:
- SpeI and PstI on K118006/CrtB
- XbaI and PstI on K118014/CrtI
- XbaI and PstI on K152005/CrtEBIY
- SpeI and PstI on R0011/Constitutive Promoter
First lane on the left was loaded with 2-log DNA ladder. Last lane on the right was loaded with undigested K152005/CrtEBIY as negative control.
In each case, 1 ug of DNA sample was used, with 1 ul of each enzymes, 2 ul of 10X FastDigest buffer, and appropriate amount of water to make up 20 ul final volume. The protocol suggested keeping at 37 oC for 5 minutes for fast digestion, then heat inactivation at 80 oC for 20 minutes to stop enzymatic reaction.
Restriction digestion products were loaded into agarose gel with GelGreen stain and run at 80 V (constant) for 45 minutes.
Dry Work
Promoter response to input concentration
- The pBAD promoter currently used in the amplification system is sensitive to different concentrations of arabinose, although this effect could be considered limited, so this is probably not the best promoter to use to get a sequence of outputs at different concentrations [].
Codon Optimisation for B.Subtilis and E.Coli
A comparison table for codon usage in the two species was made. For each amino acid, a codon was classed as Low, Medium or High use depending on the relative occurence in the genome. The intention is to create an optimised table for both species. A more rigorous and possibly more generally useful approach is under development.
BioBricks
The MelA gene and natural rbs was added to the registry as a BioBrick part. This part will be constructed, ligated into a vector, and sent to the registry as soon as the primers arrive.
Tuesday
Wet Work
Amplification
Transformed the following biobricks into the arabinose strain bacteria, plated overnight
- I746313 - pBad with activator P2 ogr and mRFP (kan resistance, low copy plasmid)
- I746314- pBad with activator PSP3 pag and mRFP (kan resistance, low copy plasmid)
- I746315- pBad with activator phiR73 delta and mRFP (kan resistance, low copy plasmid)
- I746320- PF promoter followed by GFP (high copy plasmid)
- I746321- PO promoter followed by GFP (high copy plasmid)
- I746324- Psid promoter followed by GFP (high copy plasmid)
- I746325- PLL promoter followed by GFP (high copy plasmid)
- I746370 - (failed over the weekend) P2 ogr inducing PF promoter
Also overnight, ran the plate reader to take fluorescence readings for GFP and RFP as well as OD readings of the following cultures induced with arabinose at 10mM, 1mM, 100 uM, 10 uM, and 1uM (as well as no arabinose)
- I746374
- I746375
- I746380
- I746391
Melanin
The plates varying tyrosine and IPTG induction for the bacteria transformed with the melanin plasmid were consistent with what was expected; the most pigment was produced by bacteria on plates with highest tyrosine concentration and with IPTG induction.
Progress on carotenoid (Tuesday)
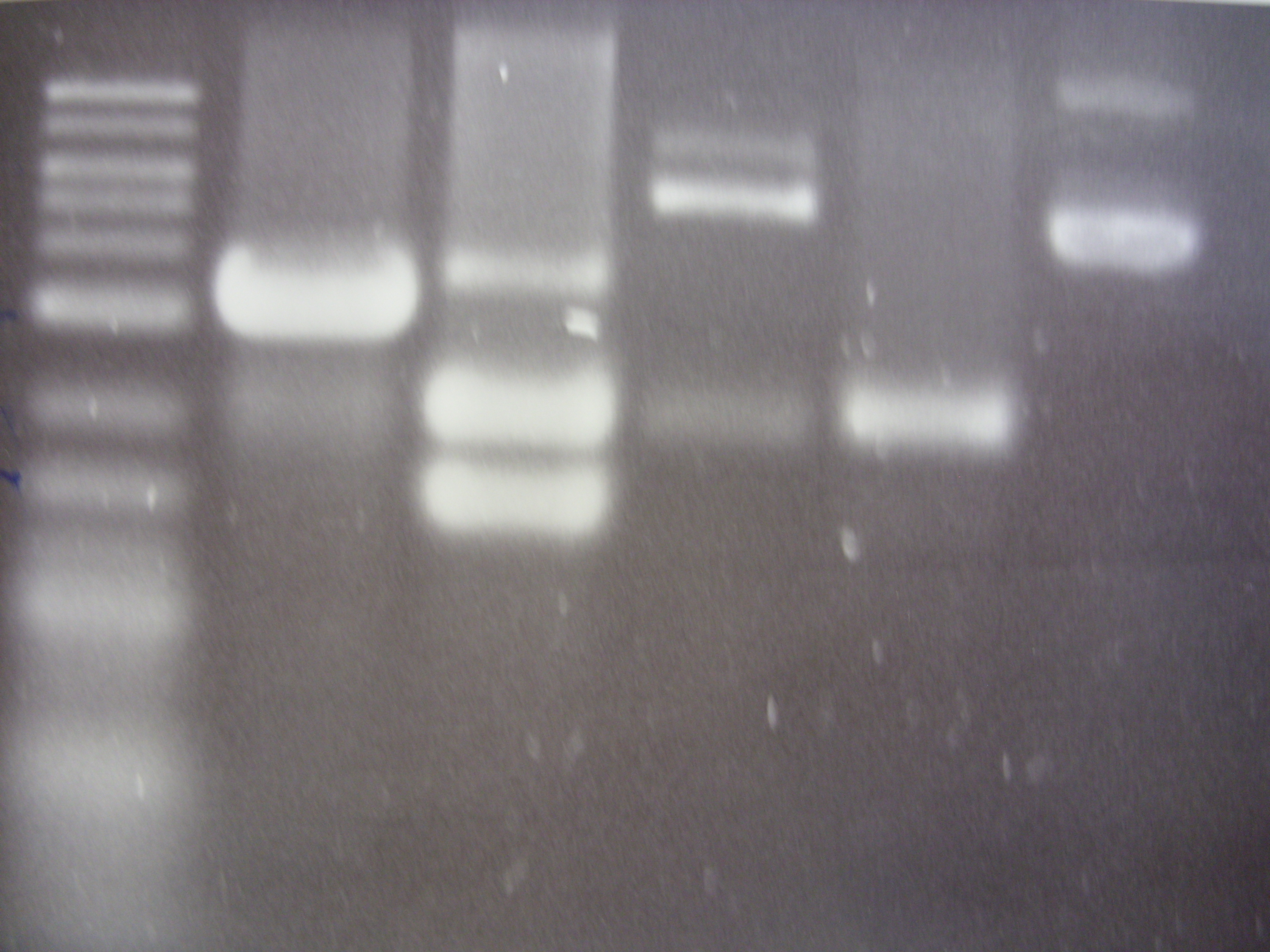
The restriction digest products of yesterday were run on 1% agarose gel with GelGreen stains for 45 minutes. 1 ug of DNA were used for each sample, together with ladder and uncut plasmid (as negative control). However, no bands were visible under UV or Blue light illumination.
To find out the cause of the problem, we put the agarose gel in EtBr solution for 1 hour staining (post-gel staining). The bands then showed up under UV illumination. This suggested that the problem might reside in GelGreen. We might have used the wrong illumination frequency, or GelGreen added was insufficient (protocol recommended 1 ul GelGreen for every 1 ml of agarose gel). Consequently, we will run an agarose gel tomorrow and then do post-gel staining with 3X GelGreen to check if the problem persists.
Results of restriction digest are shown below:
Lane 1: 2-log DNA ladder Lane 2: SpeI and PstI on K118006/CrtB (insert: 949bp, backbone: 2079bp, total: 3028bp) Lane 3: XbaI and PstI on K118014/CrtI (insert: 1498bp, backbone: 2079bp, total: 3577bp) Lane 4: XbaI and PstI on K152005/CrtEBIY (insert: 5412bp, backbone: 2079bp, total: 7491bp) Lane 5: SpeI and PstI on R0011/Constitutive Promoter (insert: 55bp, backbone: 2079bp, total: 2134bp) Lane 6: uncut K152005/CrtEBIY (negative control) (total: 7491bp)
The common bands at 2097bp are the plasmid backbone of pSB1A2. Other than that, each lane (except DNA ladder and negative control) shows at least 2 more bands: the gene insert, and the singly-cut / uncut plasmid. More importantly, the amount of singly-cut/uncut plasmid is significant, indicating that 5 minutes are probably insufficient for FastDigest. We then run restriction digest experiments at 5 minutes, 10 minutes, and 15 minutes, each with either equal amounts of XbaI and PstI, or 2:1 ratio of PstI to XbaI (this ratio is recommended on website of Fermentas for double digestion using PstI and XbaI). Restriction products will be run on agarose gel for analysis tomorrow.
To make sure that transformation colonies growing on Week 1 plates (Amp) are still viable and contain our plasmid insert, 8 colonies were picked from each Week 1 plate and transfered to freshly made Amp plates. They will be ready for mini-prep on Thursday.
Dry Work
Melanin
Received four of the six primers required to make melA into a BioBrick. The primers recieved were B/C/D and E, the four needed to cut out the restriction sites. A and F were slightly longer plasmids and so should arrive tomorrow.
Violacein
We decided to get the violacein gene synthesised in four sections, as the whole operon is too large to be economically feasible for one synthesis run. Gene designer from DNA2.0 was used to prepare the genes to be sent away for synthesis.
Wednesday
Wet Work
Amplification
Last nights plate reader results were very interesting and beg further analysis. However, it was discovered that the plate reader was at room temperature rather than 37 degrees. We'll wait to analyze the data until we have data from tonight's plate reader run.
All of last nights transformations worked except I746324- Psid promoter followed by GFP (high copy plasmid) - will try transforming it again tonight.
Cultured single colonies overnight of the following in preparation for the plate reader on Thursday night (may not actually stick to our original plan of plate reader assays, so we may not need these):
- I746313 - pBad with activator P2 ogr and mRFP (kan resistance, low copy plasmid)
- I746314- pBad with activator PSP3 pag and mRFP (kan resistance, low copy plasmid)
- I746315- pBad with activator phiR73 delta and mRFP (kan resistance, low copy plasmid)
Overnight, ran the plate reader to take fluorescence readings for GFP and RFP as well as OD readings of the following cultures induced with arabinose at 500 uM, 100 uM, 50 uM, 10 uM, 1 uM, 0.5 uM, 0,1 uM, and no arabinose
- I13540 (pBad and GFP)
- I13520 (pBad and RFP)
- I746391 (phiR73 with PO promoter) (repeated in order to assess the validity of last night's plate reader run)
MelA BioBrick
Made up the primers in preparation for tomorrows PCRs (shown in the plan below). Primers were made up to 100uM and then aliquoted into working stocks of 10uM. Primers were resuspended into sterile single distilled water and then stored at -20 degrees.
Pigment Characterisation
Attempts to trigger MelA and Violacein pigment production in culture proved unsuccessful; cell extracts were analyzed with the plate reader, but showed that no pigment was extracted. We will investigate other ways to characterize both pigments, and are considering plate-culture extraction for the melanin pigment.
Progress on carotenoids (Wednesday)
Restriction digest results from yesterday indicated that 5 minutes were probably insufficient for FastDigest. Today we wish to investigate the optimal time for restriction.
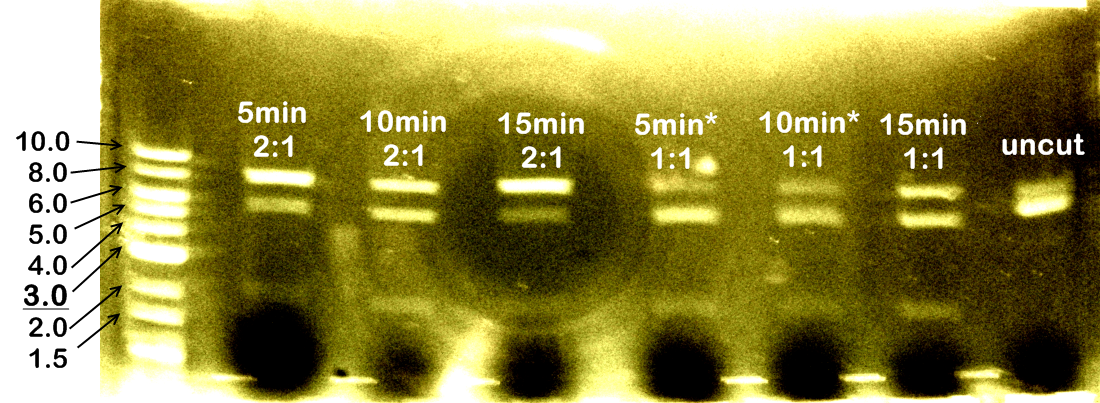
We carried out test restriction on K152005/CrtEBIY (insert 5412bp, backbone 2079bp, total 7491bp), using XbaI and PstI. We decided to investigate 2 variables: time for restriction (5 minutes, 10 minutes, and 15 minutes) and amount of enzymes used (PstI to XbaI ratio: 1:1 or 2:1 -- the protocol for "FastDigest" with 2 enzymes suggested 1:1 ratio, while protocol for "DoubleDigest" suggested 2:1 ratio for PstI to XbaI). The results were as follows:
(*"5min 1:1" and "10min 1:1" had larger volume of water than the other tubes, so the concentrations of DNA in them were slightly diluted).
The results indicated that increasing time for restriction digest could improve the yield of doubly cut products (compare "5min 2:1" and "10min 2:1"). And it seemed "1:1" ratio was ideal for restriction (compare "15min 2:1" and "15min 1:1").
Dry Work
MelA Biobrick
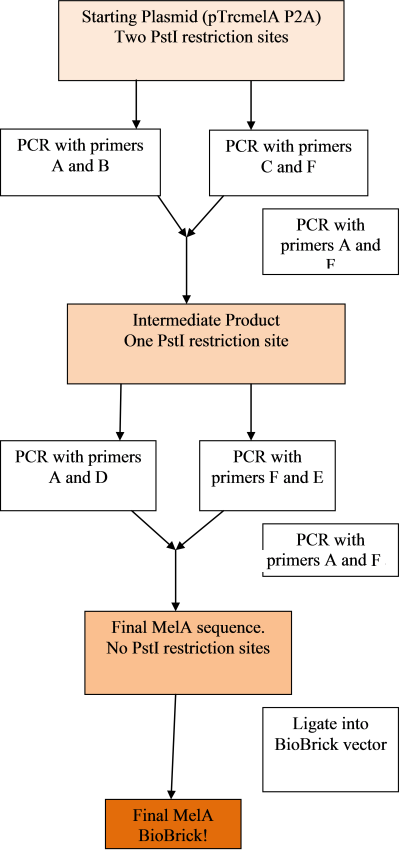
The remaining two plasmids were delivered, so a plan was made for creating the melanin biobrick and removing the PstI sites:
The PCR rounds are designed to cut the melA gene away from it's current surrounding plasmid, and change the two PstI restriction sites from CTGGAC to CTGCAA. This should not affect the coding ability of the gene. The primers will also attach the BioBrick prefix and suffix to the gene, allowing it to be easily ligated into the vector.
Violacein synthesis
As the violacein operon is too large to synthesise in one section, four seperate parts were taken (each containing one gene) to be sunthesised seperately. After synthesis they will be spliced back together to produce pigment, before isolating the genes properly to create the coloured logic gates.
Amplifier and the Big Idea
Having looked at the results of the amplification experiments, we considered further development. We wish to create a system that takes an input and at given threshold levels, turns on an output, our pigment. Potential systems have been put forward:
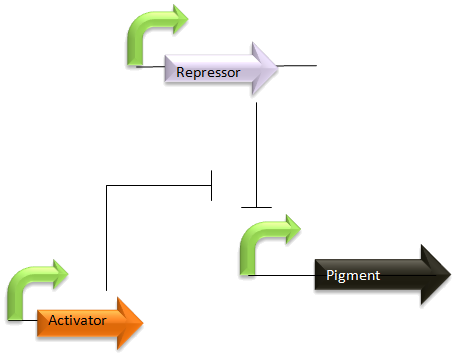
The repressor is constitutively expressed at a level determined by its controlling promoter. It prevents production of a pigment until sufficient activator is present that it overcomes the inhibition, perhaps through a competitive inhibition mechanism. A given input concentration producing a specific amount of activator will be unable to switch on the pigment until a level matching the repressor is reached. A range of strains with different repressor levels could be used to create a 'ladder' of thresholds for the switching on of different pigments.
Placing the amplifier system after this, perhaps with a positive feedback system (the activator driving its own promoter) would permit not only amplification but also allows for a latch so that pigment production, once stimulated, remains on. This would be important in an enviromental sensor to maximise an output that was clearly visual.
It was also suggested by a very intelligent engineer (he didn't write that) that placing the amplifier before the threshold system could be used to change the output for a given input by changing the effective input to the system. With one amplifier a 50um input, say, would produce brown, but with a different amplifier in place the sensitivity of the system would be altered and this input would produce violet. This permits a tuning of the system.
Research into implementation is underway.
Thursday
Wet Work
Amplification - Threshold Device
Transformed J69591 (promoter standard, 1 RPU) into arabinose strain, plated overnight.
Melanin Primers
The primers were made up to 100uM and then aliquoted into working samples of 10uM in single distilled water. All primers were stored in the freezer.
Dry Work
Ampiflication - Threshold Device
The data from the plate reader indicates that we could use our amplifiers as a means to create strains with different sensitivities. In the case of I746391, it appears to act as a digital switch, turning on GFP expression at an arabinose concentration that is lower than the arabinose concentration necessary to trigger significant RFP expression. This shows that only a small amount of activator is necessary to activate GFP expression, and hopefully different activator/promoter constructs will have a different switching threshold. This means that we already have two different sensitivities to arabinose: pBAd has a lower sensitivity to arabinose concentration and produces a linear response, while I746391 has a higher sensitivity to arabinose sensitivity and produces a digital response. We could replace fluorescence with a biobricked pigment (once we've built one) as the output. We hope to characterize each of Cambridge 2007's amplifier constructs to determine their arabinose sensitivities, and we also hope to characterize output in terms of RPUs. J69591, the standard promoter, is on pSB3k3, a low copy plasmid. We would compare the activator constructs on high copy plasmids to J69591 on pSB3k3, and then move the constructs to pSB3k3 and compare again.
The Big Idea
We've decided that we are building an arabinose sensor; different concentrations of arabinose trigger different pigment output. However, it would be brilliant if we could use an existing biosensor and use pigment as an output to show the compatibility of our output system. Thus we are looking into the arsenic biosensor project (Edinburgh 2006).
Friday
Wet Work
Threshold Device
Miniprep of J69591
Dry Work
Vio operon synthesis
As DNA2.0 very generously agreed to synthesise the whole violacein operon for us, we used the GeneDesigner program (also provided by DNA2.0) to design the operon that we wanted to synthesise (pictorial version of the sequence is shown below).
This could then by ligated into a registry vector to produce another coloured output BioBrick. The genes could also later be seperated, in particular the vioC which could then be put under a different promoter to create the logic gate system, while the remaining three genes would create yet another colour for our input system.
Data Analysis
Data from the plate reader runs was imported into Octave. The data was stored in a multi-dimensional array, with format (updated on Monday 10/8/09):
well(celltype,repeat,measurement,conc,time)
Where
- celltype is i_20, i_40 or i_91, our three biobricks. Here the 'i' stands for index
- repeat is 1, 2 or 3
- measurement is i_od, i_gfp or i_rfp
- concentration is 1, 2, 3, 4, 5, 6, 7 or 8, corresponding to 0, 0.1, 0.5, 1, 10, 50, 100 and 500μM Arabinose respectively
- time has indices running through the vector for time, 't'
This data format allows easy comparisons of responses between different runs.
Afternoon Lab Meeting
Pigments:
- Vio operon needs to be changed and designed by us ... good idea to include inducable promoter (as suggested by DNA2.0) and optimise codon usage/remove restriction sites. *Characterisation - we were having problems triggering melanin and violacein pigment production in liquid culture. Suggestions include going ahead with solid media extraction, and to trigger pigment production in liqiud culture to grow in media with more stable forms of ampicillin to ensure no overcrowding due to lack of selection, grow them longer and make sure they get enough oxygen.
- Look into different hosts with mutli-drug efflux pump components knocked out to attempt to contain melanin with in colonies so it doesn't diffuse on the plate.
- Make up strains and stocks of EVERYTHING and organise storage
- retransform violacein and melanin into Top10, make glycerol stocks
- also, transform violacein and melanin into MG1655 strain to see if we get better pigment expression
- Evidence suggests that the efflux of melanin from bacterial cells may be controlled by multi-drug transporters endogenous to e. coli [http://www3.interscience.wiley.com/cgi-bin/fulltext/119078282/HTMLSTART]
- The acrEF transporter referenced in the paper targets indoles for explusion from the cell - melanin (or di-hydroxyindole) could thus be a potential substrate for this transporter, which shows a broad range of export products
- The acrAB transporter is also implicated in the efflux of a large variety of molecules, from dyes to antiobiotics[http://www.pubmedcentral.nih.gov/picrender.fcgi?artid=177656&blobtype=pdf]
- Knockouts of these transporters may contain melanin within the cells, preventing diffusion into the surrounding media and allowing a pixelated production of pigment
- Looking to the future, pyocyanin diffusion out of the cells (which may prove to be a problem from inspection of Pseudomonas expression of the pigment) may also be limited by one of these deletions in a multi-drug transporter
- Suitable strains containing the knockouts were later requested from Philip Oliver of the Cambridge Genetics Dept.
Protocols:
- We will have a day where we go over all our protocols to make sure that they are working
- PCR - check with all working parts and do proper controls
- Restriction digests- use fermentas for 1 hour rather than the 5mins suggested. Double digest = use equal amounts of enzymes.
- Miniprep - use more DNA.
Threshold DEvice:
- Need a latch mechanism
- Make sure there is a 'not there' signal i.e negative result is different from a non-working positive response.
- design with arsenic sensor rather than pBad.
 "
"