Team:Groningen/Modelling/Arsenic
From 2009.igem.org
[http://2009.igem.org/Team:Groningen http://2009.igem.org/wiki/images/f/f1/Igemhomelogo.png]
|
|---|
- Modelling
- DetailedModel
- Characterization
- Downloads
Detailed Model
Based on the quasi-steady-state derivation below we have made the simplified version of our model shown below. The simplification is based on two key assumptions (which are also illustrated below, next to the table "Breakdown of core substances"):
- Binding and unbinding of arsenic to/from the transporters occurs on a much smaller time scale than changes in the concentration of arsenic inside and outside the cell. And similarly, we assume that (un)binding of ArsR to/from the ars promoter is much faster than the production of ArsR (for example).
- The concentration of transporters is insignificant compared to the concentration of arsenic inside and outside the cell.
This leads to the Michaelis-Menten equation for import, but also some more general equations for export using ArsB and accumulation with ArsR (for example, the Hill equation can be recognized in the activity of the ars promoter). We explicitly state relative abundances instead of substituting them into the differential equations. This leads to clearer and more insightful equations and gives more freedom to define complicated, interdependent ratios between substances.
The inexperienced viewer may find the following tables and formulas baffling. We would recommend that one would look at the raw model first to gain an understanding of the basic reactions involved then have a look at the steady-state and the quasi steady-state model. It is not mandatory, but it is probably the the best route to get a better understanding of the model as a whole. Also, perhaps first have a look at Michaelis-Menten kinetics before proceeding.
In contrast to how the quasi-steady-state assumption is normally used we mostly leave the specific states (bound/unbound) of substances intact in the differential equations and explicitly state the relative abundances. This keeps the differential equations shorter and gives more insight into what is actually happening, clearly mapping the "fast" reactions to ratios between substances. This also makes it possible to use quite complicated equations (the Asin and ArsR interdependency is virtually impossible to define using normal methods for example) that would otherwise be unwieldy to handle.
| Reaction | Description | Rate | |
|---|---|---|---|
| Transport | |||
| As(III)exT → As(III)inT | Import of arsenic. | (Vc/Vs) v5† As(III)exT / (K5+As(III)exT) | |
| As(III)inT → As(III)exT | Export of arsenic. | k8 ArsBAs | |
| ars1T → ars1T + ArsBT | Production of ArsB. | βB ars1 | |
| ArsBT → null | Degradation of ArsB | (ln(2)/τB) ArsB | |
| Accumulation | |||
| ars1T → ars1T + ArsRT | From chromosomal operon. | βRN ars1 | |
| proR → proR + ArsRT | Production of ArsR. | βR pro | |
| proM → proM + MBPArsRT | Production of MBPArsR. | βM pro | |
| proF → proF + fMTT | Production of fMT. | βF pro | |
| ArsRT → null | Degradation of ArsR. | (ln(2)/τR) ArsR | |
| MBPArsRT → null | Degradation of MBPArsR. | (ln(2)/τM) MBPArsR | |
| fMTT → null | Degradation of fMT. | (ln(2)/τF) fMT | |
| Gas vesicles | |||
| ars2T → ars2T + GV | Transcription + translation. | βG ars2 | |
| GV → null | Degradation of gas vesicles. | (ln(2)/τG) GV | |
| Name | Description | Derivative to time | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Extracellular | |||||||||||||
| As(III)exT | As(III) in the solution. | (Vc/Vs) k8 ArsBAs - (Vc/Vs) v5† As(III)exT / (K5+As(III)exT) | |||||||||||
| Membrane (all naturally occurring, but we plan to bring GlpF to overexpression) | |||||||||||||
| GlpFT | Importer of As(III) (concentration w.r.t. the exterior of the cell). | (not used directly in model, assumed to be constant) | |||||||||||
| ArsBT | Exporter of As(III) (concentration w.r.t. the interior of the cell). | βB ars1 - (ln(2)/τB) ArsB | |||||||||||
| Intracellular (ars2, pro and GV are introduced) | |||||||||||||
| As(III)inT | As(III) (bound and unbound) in the cell. | v5 As(III)exT / (K5+As(III)exT) - k8 ArsBAs | |||||||||||
| ars1T | ArsR repressed promoters (bound and unbound) naturally occurring in E. coli. | (concentration is constant = 1.6605nM, one per cell) | |||||||||||
| ars2T | ArsR repressed promoters in front of gas vesicle genes. | (concentration is constant = 0-166.05nM) | |||||||||||
| proR | Constitutive promoters in front of arsR. | (concentration is constant = 0-166.05nM) | |||||||||||
| proM | Constitutive promoters in front of mbp-arsR. | (concentration is constant = 0-166.05nM) | |||||||||||
| proF | Constitutive promoters in front of fMT. | (concentration is constant = 0-166.05nM) | |||||||||||
| ArsRT | ArsR in the cell. | βRN ars1 + βR proR - (ln(2)/τR) ArsR | |||||||||||
| MBPArsRT | MBPArsR in the cell. | βM proM - (ln(2)/τM) MBPArsR | |||||||||||
| fMTT | fMT in the cell. | βF proF - (ln(2)/τF) fMT | |||||||||||
| GV | Concentration of gas vesicles. | βG ars2 - (ln(2)/τG) GV | |||||||||||
| |||||||||||||
| Name | Units | Value | Description | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| k8 | 1/s | Reaction rate constant representing how fast ArsB can export arsenic. | |||||||||||
| KRd | M | 6µM | Dissociation constant for ArsR and As(III). Assumed to be about an order of magnitude smaller than KDd = 60µM, the corresponding constant for the similar protein ArsD from Chen1997. | ||||||||||
| KMd | M | 6µM | Dissociation constant for MBPArsR and As(III). We assume this to be roughly equal to KRd. | ||||||||||
| KFd | M | Dissociation constant for fMT and As(III). | |||||||||||
| nf | Hill coefficient for the formation of the complex fMTAs. This is related to the number of arsenic ions that bind to fMT. | ||||||||||||
| KAd | M | 0.33µM | Dissociation constants for ArsR and ars.
| ||||||||||
| v5 | mol/(s·L) | 3.1863µmol/(s·L) | Maximum import rate per liter of cells (see Michaelis-Menten equation). Note that we have purposefully chosen to write the units as mol/(s·L) instead of M/s, to emphasize the fact that the rate is per liter of cells.
| ||||||||||
| K5 | M | 27.718µM | Concentration at which import reaches half its maximum import rate (see Michaelis-Menten equation).
| ||||||||||
| K7 | M | Concentration at which export reaches half its maximum export rate (see Michaelis-Menten equation).
| |||||||||||
| τB, τR, τG, etc. | s | Half-lifes (of ArsB, ArsR and GV, respectively). Degradation rate = ln(2)/τ If you take just the degradation into account you will have the equation dC/dt = -k*C, which leads to C(t) = C(0) e-k t. So if k = ln(2)/τ we get C(t) = C(0) e-ln(2)/τ t = C(0) 2-t/τ. In other words τ is the time it takes for the concentration to half. i | |||||||||||
| βB, βR, etc. | 1/s | Production rates.
| |||||||||||
| Vs | L | Volume of solution (excluding cells). | |||||||||||
| Vc | L | Total volume of cells (in solution) (so Vs+Vc is the total volume). | |||||||||||
| |||||||||||||
The raw model
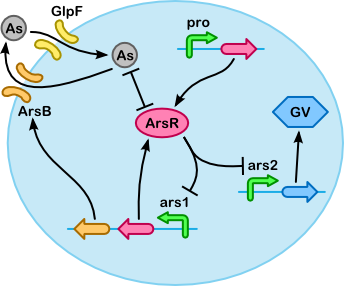
The following table gives all the reactions that take place inside the cell. You can look at the schematic representation of the processes involved to get a good grasp as how every reaction works to the other. Note that proR, ProM and MBPArsR, ProF and Fmt are not displayed in the figure. This has been done for clarity. These reactions are simple constituative promotor reactions. Once you have an insight in the reactions involved you can have a look at the next table.
| Reaction | Description | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Transport In the reactions below you can see the import of arsenic by GlpF and the export of arsenic by ArsB. Only the degradation of ArsB is taken into acount because the ars operon also produces ArsB, as can be seen in the accumulation section. We assume a constant number of GlpF importers. i | ||||||||||||
| As(III)ex + GlpF ↔ GlpFAs | The binding and detachment of arsenic to GlpF on the outside of the cell. | |||||||||||
| GlpFAs → GlpF + As(III) | The release of arsenic on the inside of the cell by GlpF | |||||||||||
| As(III)in + ArsB ↔ ArsBAs | The binding and detachment of arsenic to the Exporter ArsB | |||||||||||
| ArsBAs → ArsB + As(III)ex | The release of the bound arsenic by ArsB on the outside of the cell. | |||||||||||
| ArsB → null | The degradation of Ars B | |||||||||||
| Accumulation In the reactions below you can see the production and degradation of all our accumulation proteins. Two things should be noticed: ArsR represses it's own production and that of the GVP clusters and the ars1 operon does not only produce ArsR but also the exporter ArsB i | ||||||||||||
| As(III)in + ArsR ↔ ArsRAs | The binding and detachment of arsenic to ArsR | |||||||||||
| As(III)in + MBPArsR ↔ MBPArsRAs | The binding and detachment of arsenic to MBPArsR | |||||||||||
| nf As(III)in + fMT ↔ fMTAs | The binding and detachment of arsenic to fMT | |||||||||||
| ars1 + 2 ArsR ↔ ArsRars1 | the repression of the promotor of the ars1 operon by 2 arsR molecules | |||||||||||
| ars2 + 2 ArsR ↔ ArsRars2 | the repression of the promotor of the ars1 operon by 2 arsR molecules | |||||||||||
| ars1 → ars1 + ArsR + ArsB | The transcription and translation of the ars1 operon to produce ArsR and ArsB | |||||||||||
| proR → proR + ArsR | The transcription and translation of the proR operon to produce ArsR | |||||||||||
| proM → proM + MBPArsR | The transcription and translation of the proM operon to produce MBPArsR | |||||||||||
| proF → proF + fMT | The transcription and translation of the proF operon to produce fMT | |||||||||||
| ArsR → null | The degradation of ArsR | |||||||||||
| MBPArsR → null | The degradation of MBPArsR | |||||||||||
| fMT → null | The degradation of fMT | |||||||||||
| Gas vesicles These two reactions give the production and degradation rate of the GVP clusters. Keep in mind that ars2 is repressed by the accumulation protein ArsR. This reaction can be found under accumulation part. i | ||||||||||||
| ars2 → ars2 + GV | The transcription and translation of the ars2 operon to produce GVP clusters wich will make the cell float | |||||||||||
| GV → null | The degradation of GVP | |||||||||||
| ||||||||||||
Here you can find the time derivatives for each substance we derived. The constants are explained in the next teble. After one has a full understanding of all the constants and derivatives and and reactions. One can begin the process of simplifying the model and thus one can have a look at the quasi steady-state model and the steady-state model.
| substance | Description | Derivative to time | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Extracellular | |||||||||||||
| As(III)ex | As(III) in the solution | (d/dt) As(III)ex = - (d/dt) GlpFAs - k6 GlpFAs + (Vc/Vs) k8 ArsBAs | |||||||||||
| Membrane (all naturally occurring, but we plan to bring GlpF to overexpression) | |||||||||||||
| GlpF | concentration w.r.t. the exterior of the cell | (d/dt) GlpF = - (d/dt) GlpFAs | |||||||||||
| GlpFAs | concentration w.r.t. the exterior of the cell | (d/dt) GlpFAs = k5on As(III)ex GlpF - (k5off+k6) GlpFAs | |||||||||||
| ArsB | concentration w.r.t. the interior of the cell | (d/dt) ArsB = - (d/dt) ArsBAs + β4 ars1 - ln(2)/τB ArsB | |||||||||||
| ArsBAs | concentration w.r.t. the interior of the cell | (d/dt) ArsBAs = k7on As(III)in ArsB - (k7off+k8) ArsBAs | |||||||||||
| Intracellular (ars2, pro and GV are introduced) | |||||||||||||
| As(III)in | concentration of As(III) inside the cell | (d/dt) As(III)in = - (d/dt) ArsRAs - (d/dt) MBPArsRAs - nf (d/dt) fMTAs - (d/dt) ArsBAs - k8 ArsBAs + (Vs/Vc) k6 GlpFAs | |||||||||||
| ars1 ars1 stands for the promotor in front of the operon which contains the information for the production of the accumulation protein ArsR and the exporter ArsB. It is selfregulatory in the sence that it produces it's own repressor in the form of ArsR i | concentration of unbound promoters naturally occurring in E. coli | (d/dt) ars1 = - (d/dt) ArsRars1 | |||||||||||
| ars2 ars2 stands for the promotor in front of the operon which contains the information for the production of Gas Vesicles. Unlike ars 1 it is not selfregulatory, but the if everything goes correctly the production of gas vesicles will only start if there arsenic inside the cell i | concentration of unbound promoters in front of gas vesicle genes | (d/dt) ars2 = - (d/dt) ArsRars2 | |||||||||||
| proR | concentration of constitutive promoters in front of arsR | (d/dt)proR = 0 in our model | |||||||||||
| proM | concentration of constitutive promoters in front of mbp-arsR | (d/dt)proM = 0 in our model | |||||||||||
| proF | concentration of constitutive promoters in front of fMT | (d/dt)proF = 0 in our model | |||||||||||
| ArsR ArsR binds to ars to repress production of the genes they regulate, and binds to As(III) to make it less of a problem for the cell. i | concentration of the accumulation protein ArsR | (d/dt) ArsR = βRN ars1 + βR proR - (ln(2)/τR) ArsR - (d/dt) ArsRAs - 2 (d/dt) ArsRars1 - 2 (d/dt) ArsRars2 | |||||||||||
| ArsRAs | the concentration of ArsR bound to As(III) | (d/dt) ArsRAs = kRon ArsR As(III)in - kRoff ArsRAs | |||||||||||
| ArsRars1 | the concentration of ArsR bound to ars1 | (d/dt) ArsRars1 = kAon ArsR² ars1 - kAoff ArsRars1 | |||||||||||
| ArsRars2 | the concentration of ArsR bound to ars2 | (d/dt) ArsRars2 = kAon ArsR² ars2 - kAoff ArsRars2 | |||||||||||
| MBPArsR A fusion of maltose binding protein and ArsR. It is more stable than the normal ArsR variant, but it is no longer able to act as a repressor for the ars promotor. i | a fusion of maltose binding protein and ArsR | (d/dt) MBPArsR = βM proM - (ln(2)/τM) MBPArsR - (d/dt) MBPArsRAs | |||||||||||
| MBPArsRAs | bound to As(III) | (d/dt) MBPArsRAs = kMon MBPArsR As(III)in - kMoff MBPArsRAs | |||||||||||
| fMT It is another binding protein. Unlike it's counterpart it capeble of containing up to five As(III) particles or one As(V) particle i | Arsenic binding metallotein | (d/dt) fMT = βF proF - (ln(2)/τF) fMT - (d/dt) fMTAs | |||||||||||
| fMTAs | bound to multiple As(III) | fMTAs = kFon fMT As(III)innf - kFoff fMTAs | |||||||||||
| ArsRAs | bound to As(III) | ||||||||||||
| GV | concentration of gas vesicles | (d/dt) GV = βG ars2 - ln(2)/τG GV | |||||||||||
| |||||||||||||
The variables above can be related to each other through the following "reactions" (color coding is continued below to show which parts of the differential equations refer to which groups of reactions):
Using the following constants/definitions:
| Name | Units | Description |
|---|---|---|
| kRon, kMon, k5on, etc. | 1/(M·s) | Reaction rate constants for reactions to a complex. |
| kAon | 1/(M²·s) | Reaction rate constants for reactions to a complex. |
| kFon | 1/(Mnf·s) | Reaction rate constants for reactions to a complex. |
| kRoff, kMoff, kFoff, kAoff, k5off, etc. | 1/s | Reaction rate constants for reactions from a complex. |
| k6, k8 | 1/s | Reaction rate constants representing how fast transporters transport their cargo to "the other side". |
| τB, τR, τM, τF, τG | s | Half-lifes (of ArsB, ArsR, MBPArsR, fMT and GV, respectively). Degradation rate = ln(2)/τ If you take just the degradation into account you will have the equation dC/dt = -k*C, which leads to C(t) = C(0) e-k t. So if k = ln(2)/τ we get C(t) = C(0) e-ln(2)/τ t = C(0) 2-t/τ. In other words τ is the time it takes for the concentration to half. i |
| βRN, βR, etc. | 1/s | Production rates.
|
| Vs | L | Volume of solution (excluding cells). |
| Vc | L | Total volume of cells (in solution) (so Vs+Vc is the total volume). |
See Chen1997 for the interplay between ArsR and ArsD (the latter has a role similar to ArsR, but we do not treat it, as it is not present in our system).
Quasi steady state
First of all, we assume the concentration of transporters is quite low compared to the concentration of the transported substances. After all, if this were not the case the transporters would act more like "storage" proteins than transporters (note that this can be even more rigorously justified if, for example, GlpFT<<K5). This leads to:
As(III)exT ≈ As(III)ex As(III)inT ≈ As(III)in + ArsRAs + MBPArsRAs + nf fMTAs
Also, we assume the binding and unbinding of molecules to the transporters occurs on a much finer time-scale than any actual changes to the concentrations inside and outside the cell. Similarly, within the cell we assume diffusion processes are very fast and binding/unbinding of substances is quite fast compared to the production of proteins. This leads us to assume that the following ratios between substances are constantly in equilibrium:
As(III)ex : GlpFAs ≈ As(III)ex : 0 GlpF : GlpFAs ArsB : ArsBAs As(III)in : ArsRAs : MBPArsRAs : nf fMTAs : ArsBAs ≈ As(III)in : ArsRAs : MBPArsRAs : nf fMTAs : 0 ArsR : ArsRAs : 2 ArsRars ars : ArsRars
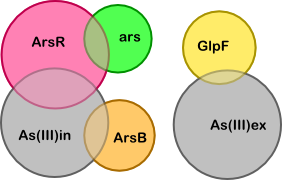
To determine what the unknown ratios are we can set the following derivatives to zero (these are the derivatives of the complexes corresponding to the four overlapping regions in the diagram):
0 = (d/dt) GlpFAs = k5on As(III)ex GlpF - (k5off+k6) GlpFAs 0 = (d/dt) ArsBAs = k7on As(III)in ArsB - (k7off+k8) ArsBAs 0 = (d/dt) ArsRars = kAon ArsR² ars - kAoff ArsRars 0 = (d/dt) ArsRAs = kRon ArsR As(III)in - kRoff ArsRAs 0 = (d/dt) MBPArsRAs = kMon MBPArsR As(III)in - kMoff MBPArsRAs 0 = (d/dt) fMTAs = kFon fMT As(III)in^nf - kFoff fMTAs
The first two derivates let us determine the ratios between bound and unbound transporters:
0 = (d/dt) GlpFAs = k5on As(III)ex GlpF - (k5off+k6) GlpFAs
k5on As(III)ex GlpF = (k5off+k6) GlpFAs
GlpF = (k5off+k6)/k5on GlpFAs / As(III)ex
GlpF = K5 GlpFAs / As(III)ex
GlpF : GlpFAs
K5 GlpFAs / As(III)ex : GlpFAs
K5 : As(III)ex
0 = (d/dt) ArsBAs = k7on As(III)in ArsB - (k7off+k8) ArsBAs
k7on As(III)in ArsB = (k7off+k8) ArsBAs
ArsB = (k7off+k8)/k7on ArsBAs / As(III)in
ArsB = K7 ArsBAs / As(III)in
ArsB : ArsBAs
K7 ArsBAs / As(III)in : ArsBAs
K7 : As(III)in
The next two differential equations can be used to determine the relative abundances of ArsR and ArsRAs, and ars and ArsRars:
0 = (d/dt) ArsRAs = kRon ArsR As(III)in - kRoff ArsRAs
kRon ArsR As(III)in = kRoff ArsRAs
ArsRAs = kRon/kRoff ArsR As(III)in
ArsRAs = ArsR As(III)in / KRd
ArsR : ArsRAs
ArsR : ArsR As(III)in / KRd
KRd : As(III)in
0 = (d/dt) ArsRars = kAon ArsR² ars - kAoff ArsRars
kAon ArsR² ars = kAoff ArsRars
ArsRars = kAon/kAoff ArsR² ars
ArsRars = ArsR² ars / KAd²
ArsR : 2 ArsRars
ArsR : 2 ArsR² ars / KAd²
KAd² : 2 ArsR ars
ars : ArsRars
ars : ArsR² ars / KAd²
KAd² : ArsR²
For MBPArsR and fMT we find:
0 = (d/dt) MBPArsRAs = kMon MBPArsR As(III)in - kMoff MBPArsRAs MBPArsR : MBPArsRAs = KMd : As(III)in 0 = (d/dt) fMTAs = kFon fMT As(III)in^nf - kFoff fMTAs fMT : fMTAs = KFd^nf : As(III)in^nf
And finally the relative abundances of arsenic:
ArsRAs = ArsR As(III)in / KRd As(III)in : ArsRAs : MBPArsRAs : n fMTAs As(III)in : ArsR As(III)in / KRd : MBPArsRT As(III)in / (KMd+As(III)in) : n fMTT As(III)in^nf / (KFd^nf+As(III)in^nf) 1 : ArsR / KRd : MBPArsRT / (KMd+As(III)in) : n fMTT As(III)in^(nf-1) / (KFd^nf+As(III)in^nf)
Summarizing:
GlpF : GlpFAs = K5 : As(III)ex ArsB : ArsBAs = K7 : As(III)in As(III)in : ArsRAs : MBPArsRAs : n fMTAs ≈ 1 : ArsR / KRd : MBPArsRT / (KMd+As(III)in) : n fMTT As(III)in^(nf-1) / (KFd^nf+As(III)in^nf) ars : ArsRars = KAd² : ArsR² ArsR : ArsRAs : 2 ArsRars ≈ 1 : As(III)in / KRd : 2 ArsR ars / KAd² MBPArsR : MBPArsRAs = KMd : As(III)in fMT : fMTAs = KFd^nf : As(III)in^nf
Now we can look at the differential equations for the totals of ArsB (so ArsBT=ArsB+ArsBAs), ArsR, As(III)in and As(III)ex (GlpFT and arsT are assumed to be constant):
(d/dt) As(III)exT = (d/dt) As(III)ex + (d/dt) GlpFAs
= - (d/dt) GlpFAs - k6 GlpFAs + (Vc/Vs) k8 ArsBAs + (d/dt) GlpFAs
= (Vc/Vs) k8 ArsBAs - k6 GlpFAs
= (Vc/Vs) k8 ArsBAs - (Vc/Vs) v5 GlpFAs / GlpFT
= (Vc/Vs) k8 ArsBAs - (Vc/Vs) v5 As(III)ex / (K5+As(III)ex)
= (Vc/Vs) k8 ArsBAs - (Vc/Vs) v5 As(III)exT / (K5+As(III)exT)
(d/dt) ArsBT = (d/dt) ArsB + (d/dt) ArsBAs
= - (d/dt) ArsBAs + βB ars1 - ln(2)/τB ArsB + (d/dt) ArsBAs
= βB ars1 - ln(2)/τB ArsB
(d/dt) As(III)inT = -(Vs/Vc) (d/dt) As(III)exT
= v5 As(III)exT / (K5+As(III)exT) - k8 ArsBT As(III)in / (K7+As(III)in)
(d/dt) ArsRT = (d/dt) ArsR + (d/dt) ArsRAs + 2 (d/dt) ArsRars
= βRN ars1 + βR proR - (ln(2)/τR) ArsR - (d/dt) ArsRAs - 2 (d/dt) ArsRars + (d/dt) ArsRAs + 2 (d/dt) ArsRars
= βRN ars1 + βR proR - (ln(2)/τR) ArsR
(d/dt) MBPArsRT = (d/dt) MBPArsR + (d/dt) MBPArsRAs
= βM proM - (ln(2)/τM) MBPArsR
(d/dt) fMTT = (d/dt) fMT + (d/dt) fMTAs
= βF proF - (ln(2)/τF) fMT
Steady state
By looking at the steady state of the system we can say something about its long-term behaviour. This also makes it easier to analyze relations between variables. To derive the steady state solution we take the quasi steady state solution and simplify it further by setting additional derivatives to zero:
0 = (d/dt) ArsBT = βB ars1 - ln(2)/τB ArsB 0 = (d/dt) As(III)inT = v5 As(III)exT / (K5+As(III)exT) - k8 ArsBAs 0 = (d/dt) ArsRT = βRN ars1 + βR pro - (ln(2)/τR) ArsR 0 = (d/dt) MBPArsRT = βM proM - (ln(2)/τM) MBPArsR 0 = (d/dt) fMTT = βF proF - (ln(2)/τF) fMT 0 = (d/dt) GV = βG ars2 - ln(2)/τG GV
This directly leads to:
0 = βB ars1 - ln(2)/τB ArsB
ArsB = βB (τB/ln(2)) ars1
ArsB = βB (τB/ln(2)) ars1T KAd²/(KAd²+ArsR²)
0 = βM proM - (ln(2)/τM) MBPArsR
MBPArsR = βM (τM/ln(2)) proM
0 = βF proF - (ln(2)/τF) fMT
fMT = βF (τF/ln(2)) proF
0 = βG ars2 - ln(2)/τG GV
GV = βG (τB/ln(2)) ars2
GV = βG (τB/ln(2)) ars2T KAd²/(KAd²+ArsR²)
For the intra- and extracellular concentrations we can find the following equation, giving a maximum for As(III)in of K7 v5/(k8 ArsB) (as As(III)exT cannot be negative)x/(c-x) is non-negative and non-decreasing for x∈[0,c⟩.
0 = v5 As(III)exT / (K5+As(III)exT) - k8 ArsBAs
0 = v5 As(III)exT / (K5+As(III)exT) - k8 ArsB As(III)in / K7
0 = v5 As(III)exT - k8 ArsB As(III)in / K7 (K5+As(III)exT)
0 = v5 As(III)exT - k8 ArsB As(III)in As(III)exT / K7 - k8 ArsB As(III)in K5 / K7
0 = As(III)exT (v5 - k8 ArsB As(III)in / K7) - k8 ArsB As(III)in K5 / K7
As(III)exT = k8 ArsB As(III)in K5 / (v5 K7 - k8 ArsB As(III)in)
As(III)exT = K5 As(III)in / (K7 v5/(k8 ArsB) - As(III)in)
As we can safely assume arsenic neither disappears into nothingness nor appears from nothingness, we can use this to derive (As(III)T is the total amount of arsenic):
As(III)inT = As(III)in (1 + ArsR/KRd + MBPArsR/KMd + fMT As(III)in^(nf-1)/KFd^nf) As(III)T = Vs As(III)exT + Vc As(III)inT 0 = Vs As(III)exT + Vc As(III)inT - As(III)T 0 = Vs K5 As(III)in / (K7 v5/(k8 ArsB) - As(III)in) + Vc As(III)in (1 + ArsR/KRd + MBPArsR/KMd + fMT As(III)in^(nf-1)/KFd^nf) - As(III)T
As the function on the right-hand side is non-decreasing for As(III)in∈[0,K7 v5/(k8 ArsB)⟩ it at most has one zero on this interval (and it has one, as it starts at a negative value and gets arbitrarily large as As(III)in approaches the end of its range). So this zero can safely be found using any number of numerical methods.
Finally, for ArsR we can find the following third-order equation:
0 = βRN ars1 + βR pro - (ln(2)/τR) ArsR 0 = βRN ars1T KAd²/(KAd²+ArsR²) + βR pro - (ln(2)/τR) ArsR 0 = βRN ars1T KAd² + βR pro (KAd²+ArsR²) - (ln(2)/τR) ArsR (KAd²+ArsR²) 0 = βRN ars1T KAd² + βR pro KAd² + βR pro ArsR² - (ln(2)/τR) ArsR KAd² - (ln(2)/τR) ArsR³ 0 = (βRN ars1T + βR pro) KAd² - (ln(2)/τR) KAd² ArsR + βR pro ArsR² - (ln(2)/τR) ArsR³ 0 = (βRN ars1T + βR pro) (τR/ln(2)) KAd² - KAd² ArsR + βR (τR/ln(2)) pro ArsR² - ArsR³
According to Mathematica's solution of Reduce[eq && KAd > 0 && arsT >= 0 && pro >= 0 && β1 > 0 && β3 > 0 && τR > 0, ArsR, Reals] (where eq is the equation shown above) there is only one real solution (examining the discriminant of eq confirms this), so we can solve the equation safely using Newton's (or Halley's) method.
 "
"