Team:Victoria Australia/Project/Introduction
From 2009.igem.org
Introduction
| There has been much talk about the environment in the media today; however the main issue revolves around the accumulation of greenhouse gasses leading to global warming. By developing a biological lighting system this could help in decreasing the burden of greenhouse gas emissions from electrical power supply due to coal-fired power generation. Though our system is very basic, there certainly is potential to fully develop biological lighting systems as a complete in-house lighting system or in lighting street lamps, emergency exit lighting in buildings and even in fun ways such as in trees at Christmas time. To develop this system, we have chosen to examine simple genetic circuits of fluorescing proteins produced from a more difficult cell free chassis due to the distinct advantages of control of the system and greater safeties; which are significant ethical considerations for Synthetic Biology that are potentially addressed using cell free chasses.
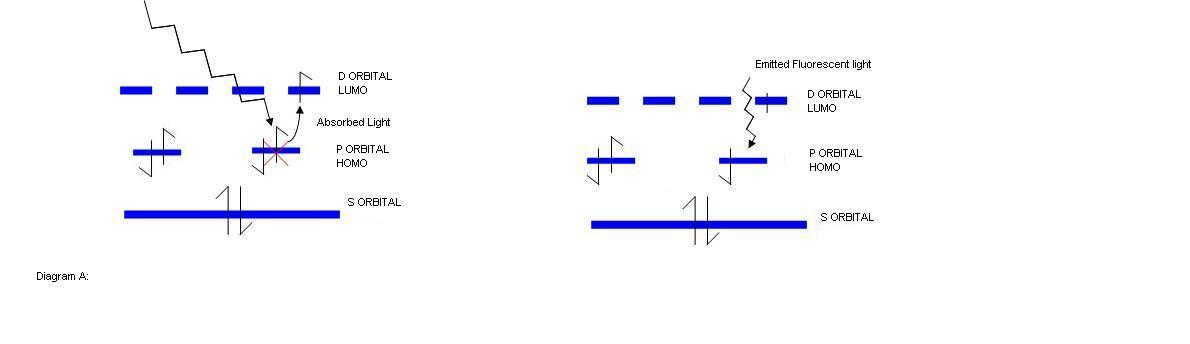
Fluorescent proteins based on Green Fluorescent Protein (GFP) are made of eleven β-strands that make up a three dimensional β-barrel protein which has one internal β-sheet with an β-helix running through the middle. Uniquely, the fluorescence of these proteins is due to the chromophore found in the centre of the β-barrel. The chromophore formation is provided by the inward facing side chains of the β-barrel that induce the formation of a ring compound in the protein’s active site which is found centrally at the tripeptide sequence Ser65-Tyr66-Gly67. Fluorescence is induced when light of an excitation wavelength is captured within the chromophore, causing an outer shell electron to rise to a higher molecular orbital. This is an unstable configuration and results in the slow reversion of the electron to the ground state. In doing so, energy is released as emitted light (see diagram A). For different fluorescent proteins based on GFP, the changing of colours is simply a matter of changing the important residues involved in forming the chromophore or residues that associate with the chromophore.
| Because genomic DNA is methylated, the parent plasmid that is being changed and mutant colour can be resolved since the PCR mutant product will not have methylated DNA, hence digestion with the restriction enzyme Dpn I will remove the template DNA (as it cuts only methylated DNA), leaving the mutated DNA for transformation. Plasmid can therefore be grown, purified and secondarily transformed into a protein expression strain and mutagenesis rapidly screened by fluorescent behaviour.
The fluorescent proteins will be expressed initially in E.coli as it ensures the GFP expresses well and there is a relatively simple translation from E.coli cells to an E.coli-based cell-free chassis. This system of mutagenesis uses defined and simple PCR conditions, simple protocols and rapid steps to “cut to the chase” of producing a new Biobrick with minimal efforts. Essentially it allows us to rapidly perform the PCR-mutagenesis reaction and template digestion which can be directly used in the subsequent transformations and protein synthesis of the blue fluorescent protein. Once it has been tested for function in E.coli, plasmid will be introduced into the analogous E.coli cell free chassis to ensure that everything is working. In the following experiments, an E.coli cell free chassis and a Wheat germ cell free chassis will be used. A cell free chassis is used in the study of biological reactions that occur within the cell without the complex interactions found in the whole cell; hence a cell free expression involves a coupled transcription translation system that is minimal to the production of the desired protein from the genetic circuit. The wheat germ expression is the ultimate end-point as it is analogous to plant waste material which could be used as a viable chassis for expression. The Wheat germ system is also a GRAS system (generally regarded as safe) and due to ethical reasons – especially safety, the public would be more accepting of this chassis than more toxic bacterial systems.
|
 "
"