Team:Newcastle/Metals
From 2009.igem.org
(→Lab Work Done) |
(→Lab Work Done) |
||
| Line 70: | Line 70: | ||
<br> | <br> | ||
==Lab Work Done== | ==Lab Work Done== | ||
| - | {| border="1" | + | {| class=wikitable border="1" |
|- | |- | ||
! colspan="2" | Summary of Lab Sessions for Metal Sequestering | ! colspan="2" | Summary of Lab Sessions for Metal Sequestering | ||
Revision as of 16:19, 21 October 2009
Metal Sequester
Introduction
In order to take up and keep the cadmium from the soil efficiently, we needed to think about a way to cross-link the metal ions to intra-cellular proteins which would end up for the most of it in the spore. If the heavy metal ions were not cross linked to a "sponge protein", it could be lethal to the cell at lower concentration and the cell could end up sporulating too early, or even bursting in the soil, releasing all the heavy metal it has taken up.
From our meeting with Prof. Nigel Robinson on the 18th of March 2009, it was evident that the best way of getting cadmium into the spore is to express a metallothionein, which would 'soak up' the cadmium, which in turn would be trapped within the protein in the spore.
Novelty in this sub-project
To increase the efficiency of our system in sequestering the heavy metal cadmium from the soil, we divised a plan to make sure that most of the metal ions that have been taken in our B.subtilis cell is then rendered bio-unavailable by incorporating it into smtA metallothionein.
SmtA metallothionein protein from E. coli can bind to heavy metals [1,2,3]. They have a tendency to bind to cationic metal ions such as cadmium, copper, arsenic, mercury, silver.
It has been shown that in B.subtilis, in order to express a specific protein in the spore coat, it is possible to make a fusion protein with a spore coat protein called CotC, and a group have successfully expressed antigen proteins which are about the same size as our smtA metallothionein in order to make a vaccine[4]. By fusing CotC spore coat protein from Bacillus subtilis, our smtA metallothionein can be localized to the spore coat, hence we can successfully trap most of the metals ions into the bacterial spores.
It is also fused with GFP reporter gene in order to detect the expression of the fusion protein into the spores using a fluorescence microscopy.
Because we want most of the metals to go into the spore rather than the vegetative cell, we designed our fusion protein so that it is controlled by the native CotC promoter sigK which is activated in sporulation conditions. Therefore, our smtA metal sequester will only be expressed once the cell have made the decision to become a metal container and sporulate, and it will only be expressed in the spore.
BioBrick constructs
Lab Work Strategies
Construction
Synthesised by GeneArt, fragment: 1385 bp DNA
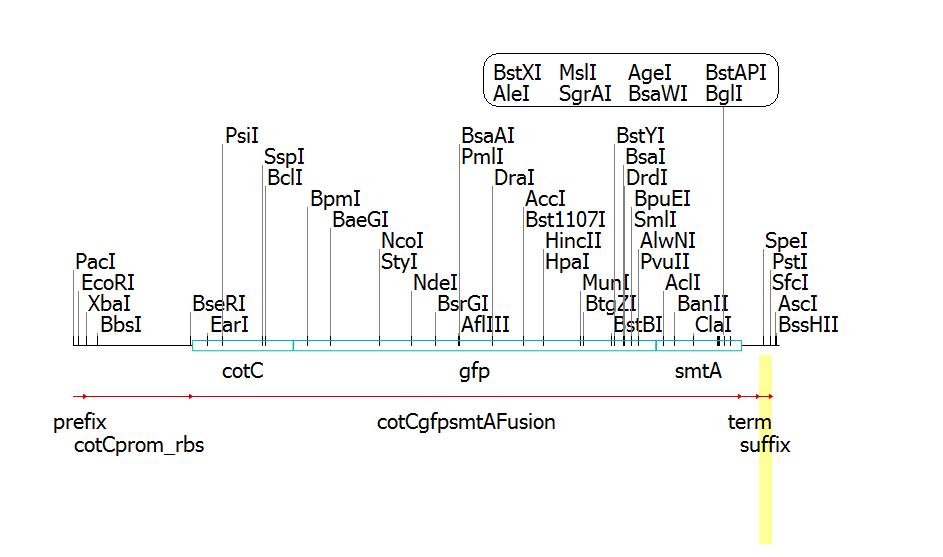
Clone Manager construct and cut map:
Sequencher construct:
Cloning and Integration
GeneArt will clone the vector into a standard biobrick vector. We will send them Plasmid pSB1AT3 with part BBa_J04450 (mCherry).
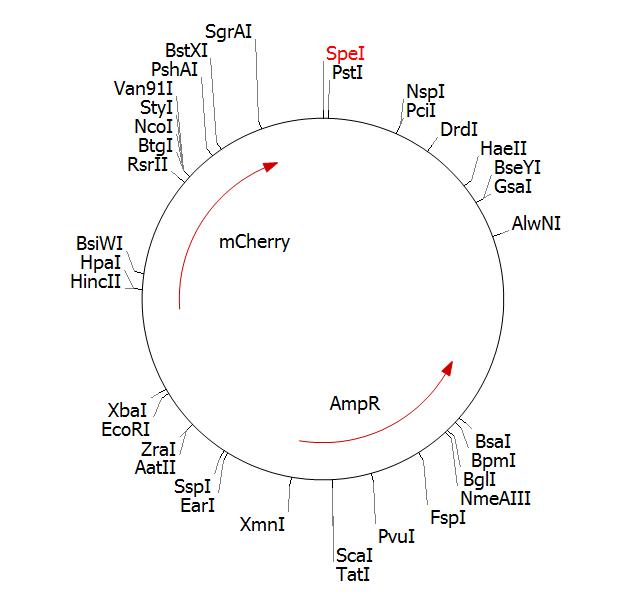
BBa_J04450 in pSB1AT3 Clone Manager plamid map:
Once we receive this fragment cloned in pSBAT3 we will amplify the part by PCR. Primer 1 will incorporate a HinDIII site and primer 2 will incorporate a BamHI. Checked that these enzymes don’t cut the fragment:
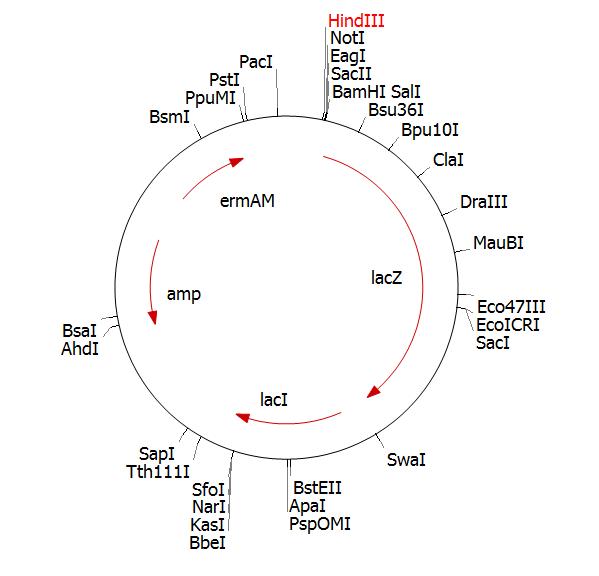
We cut the PCR fragment and clone into pMutin4 cut with the same enzymes (diagram below is pMutin2 but this is essentially the same)
We then integrate into the 168 chromosome using homology between our cotC fusion and the native copy of cotC.
Testing and Characterisation
Once transformed the spores of the mutant will need to be tested for fluorescence. Two obvious methods:
1) Fluorescence microscopy: grow the cells in sporulation medium and look under the microscope for fluorescent spores.
2) Purify spores and measure their fluorescence in a fluorescence plate reader.
3) We would also test whether the strain is able to absorb cadmium in the spores when compared to the wild-type.
Lab Work Done
| Summary of Lab Sessions for Metal Sequestering | |
|---|---|
| | |
| 04/08/09 | Arrival of the non-germination spores. Preparation of the buffer solution required for the treatment of the spores |
| 07/08/09 | Preparation of the lysozyme stock solution required for treatment of the spores |
| 10/08/09 | Re-preparation of the buffer solution required for the treatment of the spores. Pouring of agar plates with the appropriate antibiotics. |
| 11/08/09 | Re-pouring the agar plates with the appropriate antibiotics |
| 12/08/09 | Treatment of the non-germinating cwlD spores using Method A |
| 13/08/09 | Results for the treatment of the cwlD spores using Method A |
| 17/08/09 | Repeat experiment for the treatment of the cwlD spores using Method A |
| 18/08/09 | Successful results for the treatment of the cwlD spores using Method A. Performed treatment for the double-knockout mutants sleB and cwlJ spores using Method A |
| 19/08/09 | Successful results for the treatment of the double-knockout mutants sleB and cwlJ spores using Method A |
| 25/08/09 | Freezing down of the treated non-germinating spores, cwlD, and sleB and cwlJ. |
| 02/09/09 | PCR-ing of gene sleB and cwlJ using primers previously designed and ordered. |
| 03/09/09 | Attempt to PCR-ing of gene sleB and cwlJ using primers previously designed and ordered again. |
| 04/09/09 | Redesign PCR primers |
| 08/09/09 | Cloning of sleB |
References
1- Cretì, P., F. Trinchella, et al. "Heavy metal bioaccumulation and metallothionein content in tissues of the sea bream Sparus aurata from three different fish farming systems." Environmental Monitoring and Assessment.
2- Morby, A. P., J. S. Turner, et al. (1993). SmtB is a metal-dependent repressor of the cyanobacterial metallothionein gene smtA: identification of a Zn inhibited DNA-protein complex. 21: 921-925.
3- Waldron, K. J. and N. J. Robinson (2009). "How do bacterial cells ensure that metalloproteins get the correct metal?" Nat Rev Micro 7(1): 25-35.
4- Mauriello, E. M. F., L. H. Duc, et al. (2004). "Display of heterologous antigens on the Bacillus subtilis spore coat using CotC as a fusion partner." Vaccine 22(9-10): 1177-1187.
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
- Newcastle iGEM Twitter
- [http://www.facebook.com/home.php#/group.php?gid=131709337641 Newcastle on Facebook]
- [http://www.youtube.com/user/newcastle2009igem Newcastle Youtube Channel]
 "
"