Team:Cambridge/Project/Amplification
From 2009.igem.org
(→Introduction) |
(→Sensitivity Tuners) |
||
| (54 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{Template:Cambridge2}}<!--Do not remove the first and last lines in this page!--> | {{Template:Cambridge2}}<!--Do not remove the first and last lines in this page!--> | ||
| - | = | + | = Sensitivity Tuners = |
| - | + | ==Previous Work== | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
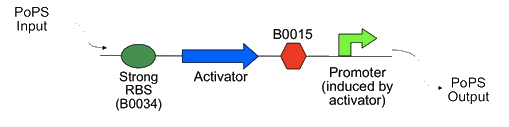
| - | + | Part of the iGEM mission is to build on previous projects. The Cambridge 2007 iGEM team developed a PoPS amplifier system using phage activators and promoters. The system works by using a PoPS input to make an activator protein, as shown in the diagram from their wiki below, which then binds to a promoter and generates a PoPS output. | |
| - | + | ||
| - | + | ||
| - | The Cambridge 2007 iGEM team developed a PoPS amplifier system using phage activators and promoters. The system works by using a PoPS input to make an activator protein, as shown in the diagram from their wiki below, which then binds to a promoter and generates a | + | |
[[Image:amplifier07.jpg]] | [[Image:amplifier07.jpg]] | ||
| - | In order to quantify the ratio between PoPS in and PoPS out, the team built the following construction on the high copy plasmid pSB1A2, with mRFP and GFP as PoPS | + | In order to quantify the ratio between PoPS in and PoPS out, the team built the following construction on the high copy plasmid pSB1A2, with mRFP and GFP as PoPS reporter. They genenerated 15 total combinations of different activators and promoters. |
| + | |||
| + | In addition to analysing the characteristics of the Cambridge 2007 constructs, we looked at [https://2009.igem.org/Team:Cambridge/Modelling models] of alternatives ways to alter the sensitivity of a promoter | ||
[[Image:construction07.jpg]] | [[Image:construction07.jpg]] | ||
| - | They successfully quantified the | + | They successfully quantified the PoPS amplification factors for each activator/promoter combination after arabinose induction. |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | == | + | ==Kit of Parts== |
| - | + | The Cambridge 2009 team analyzed the properties of these constructs, paying particular attention to how adding phage activator and promoter combinations downstream of pBad altered the behaviour of pBad. In particular, it caught our attention that the phage activator and promoter combinations changed the sensitivity of pBad. We sought to thoroughly characterise each combination to generate a useful description of each so that future iGEM teams seeking to build biosensors can easily identify which combination is appropriate, depending on how they desire to manipulate their chosen promoter. Further, we redesigned the 2007 team's constructs, removing the fluorescent reporter genes used for characterisation, as follows: | |
| - | + | [[Image:thresholddevice3.jpg]] = [[Image:converter.jpg]] | |
| - | + | The table summarizes the Sensitivity Tuners we designed and characterised: | |
| - | = | + | {| border="1" |
| + | |+ | ||
| + | ! !! P2 ogr activator !! PSP3 pag activator !! phiR73 delta activator | ||
| + | |- | ||
| + | ! PF promoter | ||
| + | | <partinfo>BBa_K274370</partinfo> || <partinfo>BBa_K274380</partinfo>|| | ||
| + | |- | ||
| + | ! PO promoter | ||
| + | |<partinfo>BBa_K274371</partinfo> | ||
| + | |<partinfo>BBa_K274381</partinfo> | ||
| + | |<partinfo>BBa_K274391</partinfo> | ||
| + | |- | ||
| + | ! PP promoter | ||
| + | | | ||
| + | |<partinfo>BBa_K274382</partinfo> | ||
| + | |<partinfo>BBa_K274392</partinfo> | ||
| + | |- | ||
| + | ! Psid promoter | ||
| + | |<partinfo>BBa_K274374</partinfo> | ||
| + | |<partinfo>BBa_K274384</partinfo> | ||
| + | |<partinfo>BBa_K274394</partinfo> | ||
| + | |- | ||
| + | ! PLL promoter | ||
| + | |<partinfo>BBa_K274375</partinfo> | ||
| + | | | ||
| + | |<partinfo>BBa_K274395</partinfo> | ||
| + | |} | ||
| - | <!--Do not remove the first and last lines in this page!--> | + | <!--Do not remove the first and last lines in this page!-->{{Template:CambridgeBottom}} |
Latest revision as of 22:45, 21 October 2009
Categories :
Project :
-
Overview
Sensitivity Tuner
--- Characterisation
--- Modelling
Colour Generators
--- Carotenoids (Orange/Red)
--- Melanin (Brown)
--- Violacein (Purple/Green)
The Future
Safety
Notebook :
Team Logistics :
Sensitivity Tuners
Previous Work
Part of the iGEM mission is to build on previous projects. The Cambridge 2007 iGEM team developed a PoPS amplifier system using phage activators and promoters. The system works by using a PoPS input to make an activator protein, as shown in the diagram from their wiki below, which then binds to a promoter and generates a PoPS output.
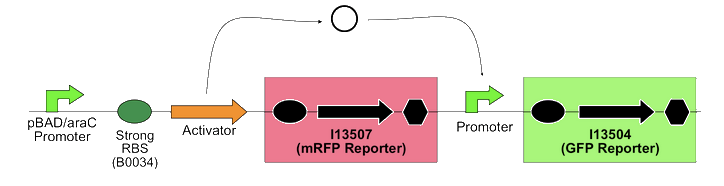
In order to quantify the ratio between PoPS in and PoPS out, the team built the following construction on the high copy plasmid pSB1A2, with mRFP and GFP as PoPS reporter. They genenerated 15 total combinations of different activators and promoters.
In addition to analysing the characteristics of the Cambridge 2007 constructs, we looked at models of alternatives ways to alter the sensitivity of a promoter
They successfully quantified the PoPS amplification factors for each activator/promoter combination after arabinose induction.
Kit of Parts
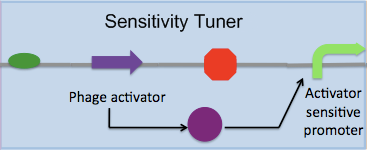
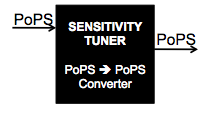
The Cambridge 2009 team analyzed the properties of these constructs, paying particular attention to how adding phage activator and promoter combinations downstream of pBad altered the behaviour of pBad. In particular, it caught our attention that the phage activator and promoter combinations changed the sensitivity of pBad. We sought to thoroughly characterise each combination to generate a useful description of each so that future iGEM teams seeking to build biosensors can easily identify which combination is appropriate, depending on how they desire to manipulate their chosen promoter. Further, we redesigned the 2007 team's constructs, removing the fluorescent reporter genes used for characterisation, as follows:
The table summarizes the Sensitivity Tuners we designed and characterised:
| P2 ogr activator | PSP3 pag activator | phiR73 delta activator | |
|---|---|---|---|
| PF promoter | |||
| PO promoter | |||
| PP promoter | |||
| Psid promoter | |||
| PLL promoter |
 "
"