Team:TUDelft/Modeling Cascade
From 2009.igem.org
(Difference between revisions)
(→Stability) |
|||
| Line 100: | Line 100: | ||
|- | |- | ||
|rowspan="2"| | |rowspan="2"| | ||
| - | <gallery> | + | <gallery widths="240px" perrow="2" heights="188px"> |
| - | + | ||
| - | Image: | + | Image:3d Clamp vs alpha4 regions.png| Maximum Transcription Rate of λp vs Translation Rate α4 (of GFP mRNA). |
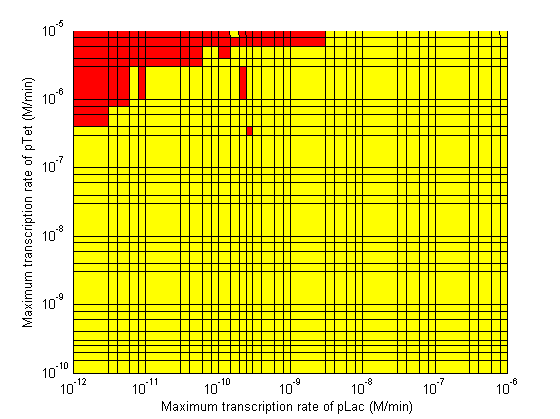
| - | Image: | + | Image:3d CpLac vs Clamp regions.png| Maximum Transcription Rate of pLac vs Maximum Transcription Rate of λp. |
| - | + | Image:3d CpLac vs CpTet regions.png| Maximum Transcription Rate of pLac vs Maximum Transcription Rate of pTet. | |
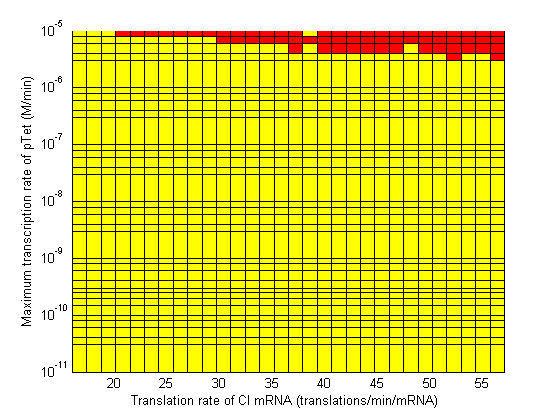
| - | Image: | + | Image:3d CpTet vs alpha2 regions.png| Maximum Transcription Rate of pTet vs Translation Rate α2 (of CI mRNA). |
| - | Image: | + | Image:3d CpTet vs Clamp regions.png| Maximum Transcription Rate of pTet vs Maximum Transcription Rate of λp. |
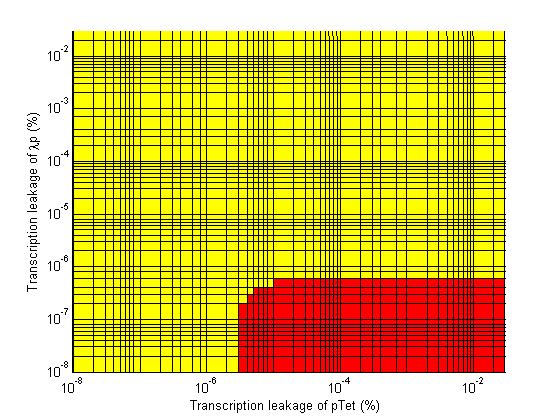
| + | Image:3d leakage a2 vs a4 regions.png| Transcription Leakage of λp vs Transcription Leakage of pTet. | ||
</gallery> | </gallery> | ||
Revision as of 14:52, 1 October 2009
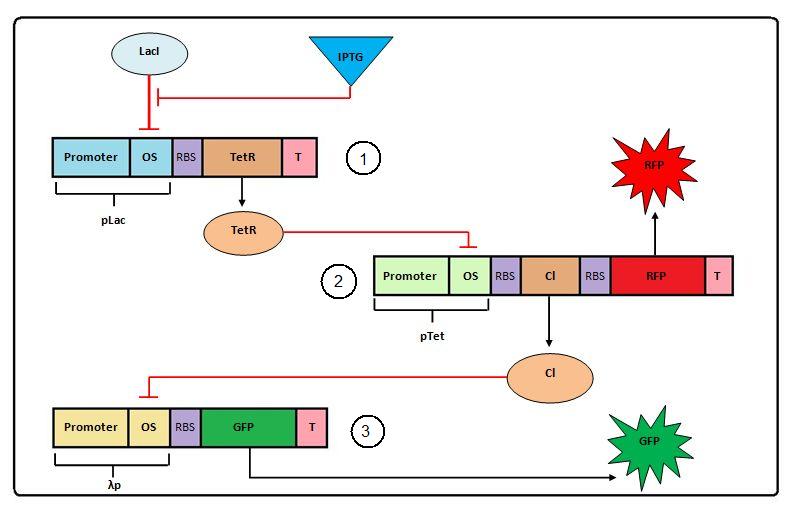
Modeling the Transcriptional Cascade
A full description of the Transcriptional Cascade can be found here.
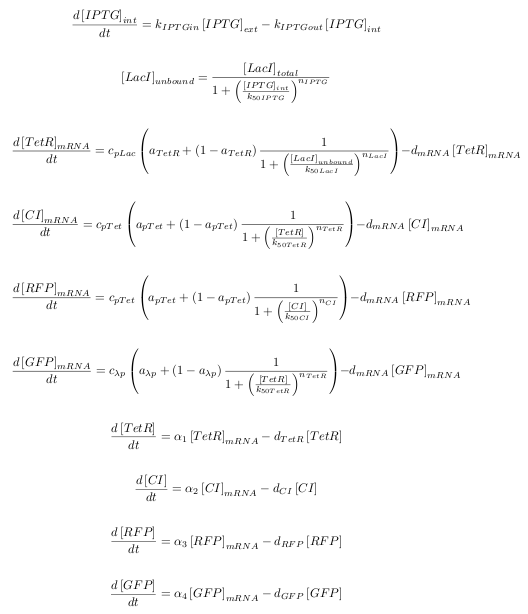
ODEs
The kinetic equations were written out in a Matlab script for both transcription and translation.
| Symbol | Definition |
| kIPTGin, kIPTGout | rate constants |
| k50IPTG, k50LacI, k50TetR, k50CI | dissociation constants |
| dmRNA | mRNA degradation rate |
| dTetR, dCI, dRFP, dGFP | protein degradation rates |
| apLac, apTet, aλp | transcription leakage (%) |
| cpLac, cpTet, cλp | maximum transcription rates |
| α1, α2, α3, α4 | translation rates |
| nIPTG, nLacI, nTetR, nCI | Hill coefficients |
| [X]mRNA | concentration of X mRNA |
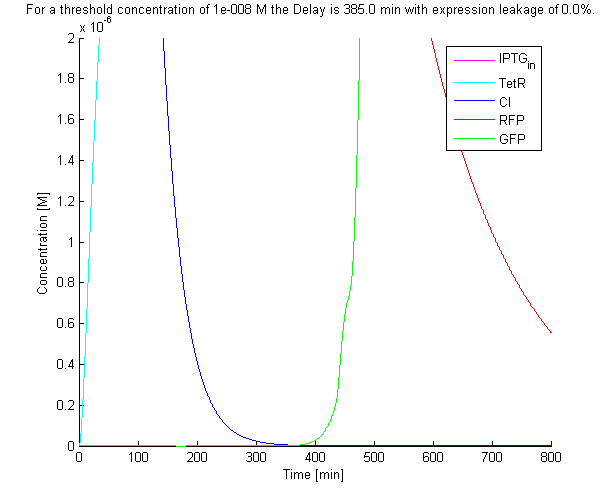
The default solution of the system of ODEs can be seen below:
Sensitivity
| Parameter | Normalized Sensitivity |
| kIPTGin, kIPTGout | |
| k50IPTG, k50LacI, k50TetR, k50CI | |
| dmRNA | |
| dTetR, dCI, dRFP, dGFP | |
| apLac, apTet, aλp | |
| cpLac, cpTet, cλp | |
| α1, α2, α3, α4 | |
| nIPTG, nLacI, nTetR, nCI | |
| [X]mRNA |
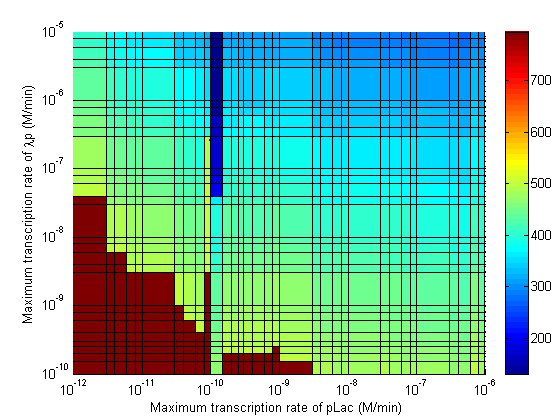
Parameter Sweeps
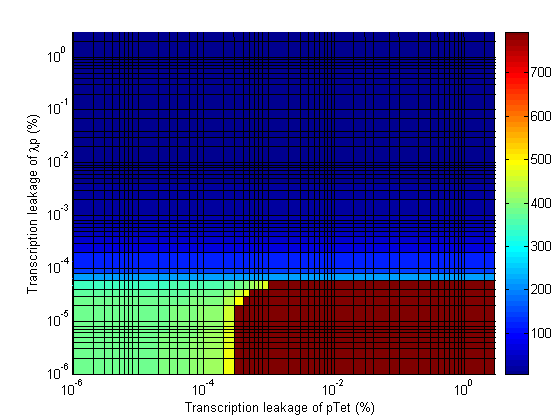
In the following plots the delay time of the cascade is shown as a function of two different parameters. A delay time off 500 is used to represent an infinite delay (maroon colour). The delay is considered to be over once the concentration of the final product (in this case GFP) goes above 10nM.
|
|
Stability
Jacobian
|
|
Design Recommendations
Based on the results of the simulations, a series of recommendations were given to the delay team to aid them in choosing parts which would maximize the delay time.
- Significant transcription leakages greatly shorten the delay time. Attempt to minimize leakages. Leakage of λp is a far bigger problem than pTet leakage.
- Use a weak promoter and a weak RBS on the last stage (λp) of the cascade.
- A weak pLac promoters is favorable.
- A strong pTet promoter is favorable.
- A strong RBS on CI gene is favorable.
- A weak RBS on TetR gene is favorable.
- A weak RBS on the endonuclease is favorable although a strong RBS can be used for the GFP gene.
- When choosing RBS and promoter strengths avoid the red areas on the stability plots.
 "
"