Team:Groningen/Project/Transport
From 2009.igem.org
m |
m (→Arsenite uptake via GlpF) |
||
| Line 96: | Line 96: | ||
'''Modelling uptake GlpF''' | '''Modelling uptake GlpF''' | ||
| + | <br> | ||
<html> | <html> | ||
<script type="text/javascript" src="/Team:Groningen/Modelling/Model.js?action=raw"></script> | <script type="text/javascript" src="/Team:Groningen/Modelling/Model.js?action=raw"></script> | ||
| Line 158: | Line 159: | ||
{{GraphHeader}} | {{GraphHeader}} | ||
| - | + | '''Missing information/To Do''' | |
*Expression assesment | *Expression assesment | ||
**Stability | **Stability | ||
| Line 169: | Line 170: | ||
*<del>Q: What does the ars operon of our <i>E. coli</i> look like? Do we have both ArsA and ArsB? (And what about ArsR and ArsD?)</del> A: We only have ArsRBC, see [[Team:Groningen/BLAST|our BLAST results]]. | *<del>Q: What does the ars operon of our <i>E. coli</i> look like? Do we have both ArsA and ArsB? (And what about ArsR and ArsD?)</del> A: We only have ArsRBC, see [[Team:Groningen/BLAST|our BLAST results]]. | ||
| - | + | '''Additional sources''' | |
| + | <br> | ||
* [[Team:Groningen/Literature#Meng2004|Meng2004]] (As(III) and Sb(III) Uptake by GlpF and Efflux by ArsB in Escherichia coli) | * [[Team:Groningen/Literature#Meng2004|Meng2004]] (As(III) and Sb(III) Uptake by GlpF and Efflux by ArsB in Escherichia coli) | ||
* [[Team:Groningen/Literature#Rosen2009|Rosen2009]] (Transport pathways for arsenic and selenium: A minireview) | * [[Team:Groningen/Literature#Rosen2009|Rosen2009]] (Transport pathways for arsenic and selenium: A minireview) | ||
Revision as of 15:24, 9 October 2009
[http://2009.igem.org/Team:Groningen http://2009.igem.org/wiki/images/f/f1/Igemhomelogo.png]
|
|---|
Introduction
We are trying to find suitable systems capable of isolating heavy metals from the environment. There are several different mechanisms to achieve such a thing. We examined 3 kinds:
- Metal transporters, that transport the metal from the environment (i.e. wastewater) to the cytoplasm
- Uncoupled
- Coupled with 'helper' compounds
- Metal binding proteins in the periplasm
We will investigate severals systems, to find which are suitable for the final design. the following systems are under concideration:
- Copper/zinc uptake via HmtA
- Heavy metal uptake coupled to citrate via efCitH bsCitM
- Arsenite uptake via GlpF
- Periplasmic accumulation of heavy metals via Mer Operon.
Copper/zinc uptake via HmtA
HmtA, heavy metal transporter A from Pseudomonas aeruginosa Q9I147 is a P-type ATPase importer. It mediates the uptake of copper (Cu) and zinc (Zn) and is functionally expressed in E.coli ([http://www.ncbi.nlm.nih.gov/pubmed/19264958 Lewinson 2009]). We want to use this membrane protein to accumulate copper and zinc into the cells we believe this active transport is efficient enough to clear the medium from these compounds. NCBI database informations on HmtA: gi|81857196|sp|Q9I147|Q9I147_PSEAE Probable cation-transporting P-type ATPase.
Restriction sites
There are several restriction sites to be modified from the papers bBAD construct. We will create these mutations via PCR.
| Enzyme | Number of Sites |
|---|---|
| EcoRI | 1 |
| XbaI | 0 |
| NotI | 0 |
| SpeI | 0 |
| PstI | 2 |
Missing information/To Do
- Expression assesment
- Stability
- Level
- Functional assesment
- Uptake speed
- Affinity
- Electrolyte potential generating force
- Eliminate BioBrick restriction sites
Heavy metal uptake coupled to citrate via efCitH bsCitM
Citrate uptake coupled to heavy metals enables forcefeeding of the toxic compounds into the cell when citrate is the only carbound source available. This could be a very efficient strategy to accumulate vast ammounts of heavy metals. The current candidates are CitM from Bacilus subtilis and CitH form Enterococcus faecalis.
Missing information/To Do
- Expression assesment
- Stability
- Level
- Functional assesment
- Uptake speed
- Affinity
- Electrolyte potential generating force
- Eliminate BioBrick restriction sites
Additional sources
More information on this topic can be found in:
Bastiaan Krom. Citrate transporters of Bacilus subtilis PhD thesis. http://dissertations.ub.rug.nl/faculties/science/2002/b.p.krom/ Dissertation Groningen
Jessica B. Warner. Regulation and expression of the metal citrate transporter CitM PhD thesis. http://dissertations.ub.rug.nl/faculties/science/2002/j.b.warner/ Dissertation Groningen
Arsenite uptake via GlpF
GlpF
GlpF is an aquaglycerol porin of E.coli which facilitates not only glycerol import, but also arsenic (As) and antimone (Sb) import (Fu, DX, et al.2000, [[Team:Groningen/Literature#Meng, YL, et al.2004|(Meng, YL, et al.2004, (Porquet, A, et al.2007, (Rosen, BR, et al.2009) . It has homologues in other organisms; Fps1p has shown to facilitate arsenic import in yeast and AQP9 is the mammalian homologue [[Team:Groningen/Literature#Porquet, A, et al.2007|(Porquet, A, et al.2007, (Rosen, BR, et al.2009) .
The GlpF aquaglycerol porin is a membrane protein with a symmetric arrangement of four independent GlpF channels. One monomer of this tetramer GlpF porin consists of six transmembrane and two half membrane-spanning α-helices that form a right-handed helical bundle around the channel. The channel has a diameter of ~15Å at the periplasmid end, which constricts towards a diameter of ~3.8Å at the beginning of a 28 Å long selective channel that ends at the cytoplasmic end (Fu, DX, et al.2000).
The GlpF is a stereospecific channel that is thought to be more selective on molecular size than on chemical structure [[Team:Groningen/Literature#Fu, DX, et al.2000|(Fu, DX, et al.2000, (Heller, KB, et al.1980) . It does allow transport of a variance of polyhydric alcohols, glycerol being one of them, and arsenic and antimone [[Team:Groningen/Literature#Fu, DX, et al.2000|(Fu, DX, et al.2000, [[Team:Groningen/Literature#Meng, YL, et al.2004|(Meng, YL, et al.2004, [[Team:Groningen/Literature#Porquet, A, et al.2007|(Porquet, A, et al.2007, [[Team:Groningen/Literature#Rosen, BR, et al.2009|(Rosen, BR, et al.2009, (Heller, KB, et al.1980) . Carbon sugars and ions are shown to be unable to be transported by GlpF (Heller, KB, et al.1980) . At physiological pH arsenic and antimone are not present in their ionic state but rather as As(OH)3 and Sb(OH)3 (Rosen, BR, et al.2009) . These elements show a charge distribution similar to glycerol and a smaller but comparable volume. The structural similarities are thought to be the reason for the possibility of these elements to be transported in the cell by GlpF (Porquet, A, et al.2007) .
If GlpF behaves as a nonsaturable transporter, a transport rate of 1umol of glycerol is transported per minute per mgr of cell protein (Heller, KB, et al.1980) . The transport rate of GlpF for arsenic is estimated to be……
Modelling uptake GlpF
The import of As(III) via GlpF is modelled as a simple import reaction with Michaelis-Menten kinetics, in part because this makes it easy to specify in Simbiology, but also because we only have very high level data. The following allows a comparison with the data acquired from figure 1B from Meng2004.
|
Loading graph...
|
To determine the constants v5 and K5 we performed the following steps:
- Read the wild-type line in figure 1B of Meng2004 by pasting it in a drawing program and aligning/scaling the axes and then manually determining the coordinates of each data point.
- Converted to units of concentration using the data in Meng2004 and [http://gchelpdesk.ualberta.ca/CCDB/cgi-bin/STAT_NEW.cgi the CCDB] (assuming that the cells are resting/non-growing), see our [http://spreadsheets.google.com/pub?key=t4gilzCbEaCFAvpEVWUE_zQ Google Docs spreadsheet]. Here we disregarded the fact that the measurements were made by taking out 0.1mL samples, as this does not change the concentrations. Specifically (note that uptake is in nmol/mg):
- uptaketotal (nmol) = uptake · 8mg · 0.3 The ratio between dry and wet weight is 0.3 (see the [http://gchelpdesk.ualberta.ca/CCDB/cgi-bin/STAT_NEW.cgi CCDB]).i
- As(III)ex (µM=nmol/mL) = (10nmol/mL · 1mL - uptaketotal) / (1.1-0.0073)mL The experiment started with 1mL of a 10µM=10nmol/mL solution of As(III). After adding the cells the total volume of the solution was 1.1mL, and 0.0073mL is an estimate of the total volume of cells in the solution, see below.i
- uptaketotal (nmol) = uptake · 8mg · 0.3
- Fit the Michaelis-Menten equation to find the constants v5 and K5 in Mathematica (see [http://igemgroningen.googlecode.com/svn/trunk/buoyant/Models/Meng2004%20Figure%201B.nb the Mathematica notebook in SVN]) using the method from Goudar1999 (a least squares fit of a closed-form solution of the differential equation).
Missing information/To Do
- Expression assesment
- Stability
- Level
- Functional assesment
- Uptake speed
- Affinity
- Electrolyte potential generating force
- Eliminate BioBrick restriction sites
Q: What does the ars operon of our E. coli look like? Do we have both ArsA and ArsB? (And what about ArsR and ArsD?)A: We only have ArsRBC, see our BLAST results.
Additional sources
- Meng2004 (As(III) and Sb(III) Uptake by GlpF and Efflux by ArsB in Escherichia coli)
- Rosen2009 (Transport pathways for arsenic and selenium: A minireview)
MerR family
Periplasmic accumulation of heavy metals via Mer proteins enables the harvesting of heavy metals from the medium. By binding the cytosolic and periplasmic metals to metallothionein and transporting the metal-protein complex into the periplasm. The MerR family consists of different proteins for one specific metal. (ie. PbrR(Lead), CueR(Copper), ZntR(Zinc), MerR(Mercury), ArsR(Arsenic), CadR(Cadmium))
As the cells die after uptake of Mg (and induction of the Mer transporter) this system is not very well usable for our project. The dead cells will not produce the gas vesicles (it may be used however by having the gas vesicles consitutively expressed), thereby bouyancy may be a problem. Pennella2005, Kao2008
Missing information/To Do
- Expression assesment
- Stability
- Level
- Functional assesment
- Uptake speed
- Affinity
- Electrolyte potential generating force
- Eliminate BioBrick restriction sites
Planning and requirements:
- Modelling:
- Import speed
- Amount
- Max
- Lab:
- HmtA
- Zn/Cu alone
- B-type ATPase (could be use if there is a ATP shortage?)
- CitM (probably not used)
- Divalent ions
- Citrate around
- Citrate can bind metals that are already bound.
- Measurements (both for the "normal" cell and the cell with overexpression of the transporter)
- Transporter, on/off mechanism, up to what concentration (in the cell) does it still have metal uptake.
- Measure concentration of metal. difference between begin and end concentrations of metal outside the cell.
- How fast does it transport metal in/out the cell.
- Set up tests with (initial) extracellular concentrations of about 1/3K (25% of Vmax), K (50% of Vmax), 3K (75% of Vmax) and 10mM (99.7% of Vmax, corresponding to extremely polluted water), and a control with no arsenic. Obviously, more tests is better. In general a desired fraction of Vmax at the initial concentration can be attained by using an initial concentration of x/(1-x) times K.
- Determine "final" (steady-state) concentration of As(III) in the solution and in the cells. (Concentration over time is even better!)
- This means that the total volume of the cells (and the solution) has to be determined. Possibly through looking at the dry weight (without arsenic!).
- By manipulating the equation for the derivative of As(III) in equilibrium, As(III) can be expressed as a function of As(III)ex (given the V and K constants). We can try to fill in the computed V and K constants for GlpF and then use a least squares fit to estimate the V and K constants for ArsB.
- NOTE: Interestingly Kostal2004 already did an experiment like this with cells that overexpressed ArsR. We're looking at analysing these results under the assumption that overexpressing ArsR only gives a constant factor more accumulation (for 1-100µM As(III)), but it would be very nice to do this ourselves for unmodified cells to determine whether this is indeed true (and to determine the factor).
- HmtA
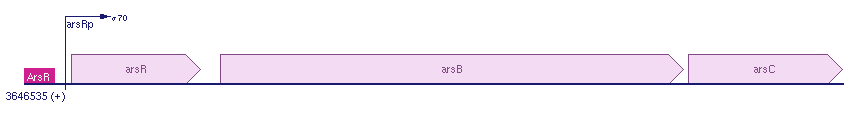
Regulation of the Ars operon
The Ars operon contains three open reading frames. The first is ArsR, second ArsB and the last one is ArsC. Transcription is controlled by the ArsR repressor. All the genes lie in the same frame and there are no others genes between the ArsRBC genes in the E. coli k12. For more information see: [http://biocyc.org/ECOLI/NEW-IMAGE?type=GENE-IN-CHROM-BROWSER&object=EG12235 biocyc].
Missing information/To do
- Expression assesment
- Stability
- Level
- Functional assesment
- Uptake speed
- Affinity
- Electrolyte potential generating force
- Eliminate BioBrick restriction sites
 "
"