Team:Newcastle/Labwork/15 September 2009
From 2009.igem.org
(→Chassis team) |
Babyneurone (Talk | contribs) (→Introduction) |
||
| (18 intermediate revisions not shown) | |||
| Line 5: | Line 5: | ||
[[Image:Team Newcastle 2009 iGEM ProbationaryP-Sign.PNG|50px|right]] | [[Image:Team Newcastle 2009 iGEM ProbationaryP-Sign.PNG|50px|right]] | ||
=Formal Lab Session - 15th September 2009= | =Formal Lab Session - 15th September 2009= | ||
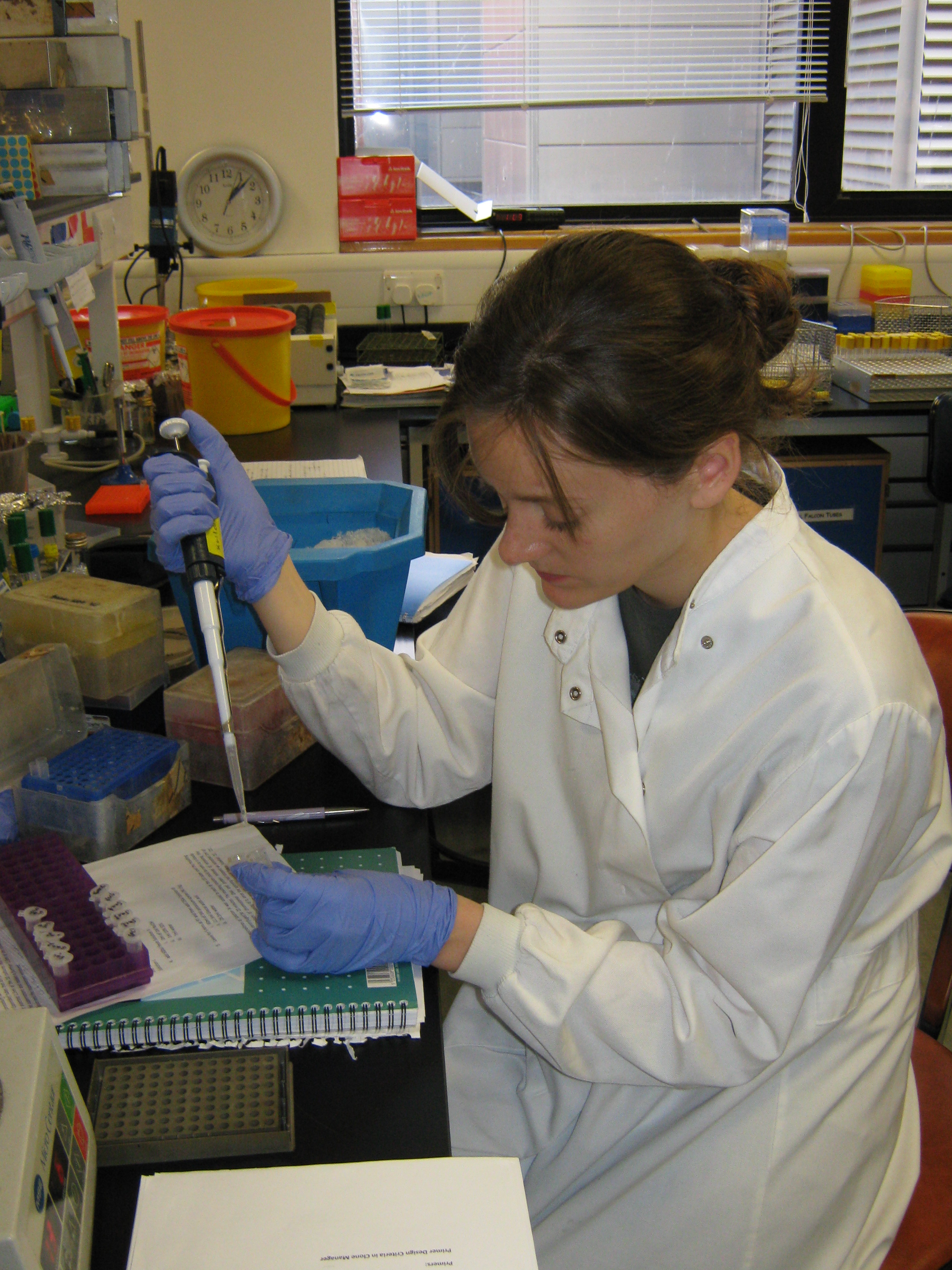
| - | + | [[Image:Team Newcastle 2009 iGEM 15-09-09 IMG 1168.JPG|350px|center]] | |
| + | <br> | ||
| + | =<font color="Orange"><u>Overview</u></font>= | ||
| + | <font color="Orange"> | ||
| + | *[[#Stochastic Switch Team|Stochastic Switch Team]] '''- ''' | ||
| + | <br> | ||
| + | *[[#Metal Sensor Team|Metal Sensor Team]] '''- ''' | ||
| + | <br> | ||
| + | *[[#Sporulation Tuning/Chassis Team|Sporulation Tuning/Chassis Team]] '''- ''' | ||
| + | <br> | ||
| + | </font> | ||
| + | <br> | ||
==<u>Stochastic Switch Team</u>== | ==<u>Stochastic Switch Team</u>== | ||
| + | [[Image:Team Newcastle 2009 iGEM 15-09-09 IMG 1169.JPG|200px|right]] | ||
Today we set up another restriction disest of the KinA pMKRQ synthesised brick using NheI and EcoRI we ran this on an 0.8% gel however we think that this may be too dilute and that I why we are not seeing the two bands ogf the digest which are a similar size. | Today we set up another restriction disest of the KinA pMKRQ synthesised brick using NheI and EcoRI we ran this on an 0.8% gel however we think that this may be too dilute and that I why we are not seeing the two bands ogf the digest which are a similar size. | ||
| Line 17: | Line 29: | ||
Yesterday we did midipreps of Goksels successful sspb plasmid: this was cut with XbaI and PstI ready for ligation into the ara brick (once it's ligated!). | Yesterday we did midipreps of Goksels successful sspb plasmid: this was cut with XbaI and PstI ready for ligation into the ara brick (once it's ligated!). | ||
<br> | <br> | ||
| - | ==<u>Metal | + | |
| + | ==<u>Metal Sensor Team</u>== | ||
===Introduction=== | ===Introduction=== | ||
| + | |||
Yesterday's lab session saw the Metal Sensing team ligate the ''cadA'' promoter region with the linearised ''BBa_J33206'' BioBrick (BioBrick containing ''arsR'' gene and binding site but with promoter absent). The team also attempted to ligate ''cotC'' and linearised ''pMUTIN4''. Both of these ligated products were used to transform ''DH5-alpha'' ''E. coli'' cells. | Yesterday's lab session saw the Metal Sensing team ligate the ''cadA'' promoter region with the linearised ''BBa_J33206'' BioBrick (BioBrick containing ''arsR'' gene and binding site but with promoter absent). The team also attempted to ligate ''cotC'' and linearised ''pMUTIN4''. Both of these ligated products were used to transform ''DH5-alpha'' ''E. coli'' cells. | ||
| Line 39: | Line 53: | ||
===Procedure=== | ===Procedure=== | ||
| + | |||
====Preparation for ''cotC'' transformant mini-preps==== | ====Preparation for ''cotC'' transformant mini-preps==== | ||
To prepare the ''cotC'' BioBricks for midi-preps, 12 colonies were picked from the LB + amp plate containing 200ul of transformant cells. These colonies, under aspetic conditions, were then used to inoculate 12 tubes of LB + ampicillin. The tubes were transferred to the orbital incubator for overnight growth at 37ºC. | To prepare the ''cotC'' BioBricks for midi-preps, 12 colonies were picked from the LB + amp plate containing 200ul of transformant cells. These colonies, under aspetic conditions, were then used to inoculate 12 tubes of LB + ampicillin. The tubes were transferred to the orbital incubator for overnight growth at 37ºC. | ||
| Line 100: | Line 115: | ||
====Ligation of ''cadA'' promoter and ''arsR'' BioBrick==== | ====Ligation of ''cadA'' promoter and ''arsR'' BioBrick==== | ||
| + | |||
====Transformation of ''E. coli'' with ligated product==== | ====Transformation of ''E. coli'' with ligated product==== | ||
Competent ''DH5-alpha E. coli'' cells were then transformed with the ligated ''cotC'' in ''pMUTIN4'' BioBrick. The transformations were carried out according to [https://2009.igem.org/Team:Newcastle/Project/Labwork/PhilsProtocols#Transforming_DNA_.22Phil_Style.22 Dr. Aldridge's transformation protocol] and there were only a few changes made: | Competent ''DH5-alpha E. coli'' cells were then transformed with the ligated ''cotC'' in ''pMUTIN4'' BioBrick. The transformations were carried out according to [https://2009.igem.org/Team:Newcastle/Project/Labwork/PhilsProtocols#Transforming_DNA_.22Phil_Style.22 Dr. Aldridge's transformation protocol] and there were only a few changes made: | ||
| Line 108: | Line 124: | ||
Once the transformation process was completed the cells were plated onto two LB + chloramphenicol plates - one plate containing 200ul of transformant cells in LB and the other plate containing 500ul of cells in LB. | Once the transformation process was completed the cells were plated onto two LB + chloramphenicol plates - one plate containing 200ul of transformant cells in LB and the other plate containing 500ul of cells in LB. | ||
| - | ==<u> Chassis | + | ==<u>Sporulation Tuning/Chassis Team</u>== |
===Introduction=== | ===Introduction=== | ||
| + | [[Image:Team Newcastle 2009 iGEM IMG 1161.JPG|200px|right]] | ||
* It has been proved that we got 9 strains which contain right sleB clone. Yesterday, we cultured 50ml cells for Midi Prep. We'll perform a Midi Prep today to get large amount of plasmid with sleB biobrick. | * It has been proved that we got 9 strains which contain right sleB clone. Yesterday, we cultured 50ml cells for Midi Prep. We'll perform a Midi Prep today to get large amount of plasmid with sleB biobrick. | ||
* Also we had get the cwlJ fragment successful, to get the cwlJ biobrick, we'll do the same procedure as we did to sleB. | * Also we had get the cwlJ fragment successful, to get the cwlJ biobrick, we'll do the same procedure as we did to sleB. | ||
* To get a double clone of cwlJ:sleB, we need digest plasmid pSB1AT3:sleB and ligate with cwlJ fragment. | * To get a double clone of cwlJ:sleB, we need digest plasmid pSB1AT3:sleB and ligate with cwlJ fragment. | ||
| - | |||
| - | |||
<br> | <br> | ||
===Experiment procedure=== | ===Experiment procedure=== | ||
| - | ==== | + | ==== Midi Prep ==== |
| - | + | * Before we carried out Midi Prep, we take out 1ml of cell mix and put them into 1.5ml eppendorf tube to keep in fridge. | |
| + | * 50ml of cells were used for Midi Prep. | ||
| + | * Midi prep processed by following the standard protocal of Plasmid Midi Prep Kit(Sigma). | ||
| - | ==== | + | ==== Ligate cwlJ with pSB1AT3 and transform into E.coli==== |
| + | * Ligation procedure was same as the procedure of sleB ligation in day 08/09/09. Ligase and buffer were came from Fast Ligation and Transformation Kit(Fermentas). | ||
| + | Both cwlJ and pSB1AT3 were cut by EcoRI and speI | ||
| + | |||
| + | * Transformation process was carried out by the same procedure in day 09/09/09. | ||
| + | ==== Digest pSB1AT3:sleB plasmid for double clone ==== | ||
| + | * Prepare the digest reaction. | ||
| + | dd H2O 7ul | ||
| + | 10X fast digest buffer 7ul | ||
| + | Fast EcoRI 3ul | ||
| + | Fast speI 3ul | ||
| + | pSB1AT3:sleB plasmid 50ul | ||
| + | ------------------------------ | ||
| + | 70ul | ||
| + | * Incubate 1 hour at 37 degree. | ||
| + | * Store at -20C. | ||
===Conclusion=== | ===Conclusion=== | ||
| - | + | * The concentration of our Midi Prep plasmid pSB1AT3:sleB is 109.3ng/ul. | |
| - | + | ||
| - | + | ||
===Futher plan=== | ===Futher plan=== | ||
| - | + | * Ligate cwlJ with pSB1AT3:sleB and transform the ligated plasmid into E.coli. | |
{{:Team:Newcastle/Project/Labwork/CalTemplate}} | {{:Team:Newcastle/Project/Labwork/CalTemplate}} | ||
{{:Team:Newcastle/Footer}} | {{:Team:Newcastle/Footer}} | ||
{{:Team:Newcastle/Right}} | {{:Team:Newcastle/Right}} | ||
Latest revision as of 12:17, 20 August 2010
Formal Lab Session - 15th September 2009
Overview
Stochastic Switch Team
Today we set up another restriction disest of the KinA pMKRQ synthesised brick using NheI and EcoRI we ran this on an 0.8% gel however we think that this may be too dilute and that I why we are not seeing the two bands ogf the digest which are a similar size.
We also set up another PCR of the ara fragment as a back up plan for ligation, in case the last ligation did not work as well as we had planned.
We also mini prepped and digested the sac transformations again, and we found that of the 6 samples we had, a few of them had worked!
Just to confirm however we set up 12 more miniprep cultures again for both sac and ara.
Yesterday we did midipreps of Goksels successful sspb plasmid: this was cut with XbaI and PstI ready for ligation into the ara brick (once it's ligated!).
Metal Sensor Team
Introduction
Yesterday's lab session saw the Metal Sensing team ligate the cadA promoter region with the linearised BBa_J33206 BioBrick (BioBrick containing arsR gene and binding site but with promoter absent). The team also attempted to ligate cotC and linearised pMUTIN4. Both of these ligated products were used to transform DH5-alpha E. coli cells.
Today's work will involve the team looking for transformant colonies on the plates and processing them based on observations. If there are colonies, 12 will be picked and used to inoculate 12 tubes of 5ml LB (+ antibiotic) for mini-preps. If unsuccessful, another attempt at ligation and then transformation will follow. The team will also carry out digests of the 5 midi-preps we had done earlier to prove that they have the correct DNA.
Practical Outline
Here are the tasks the Metal Sensing team need to get done by the end of the day:
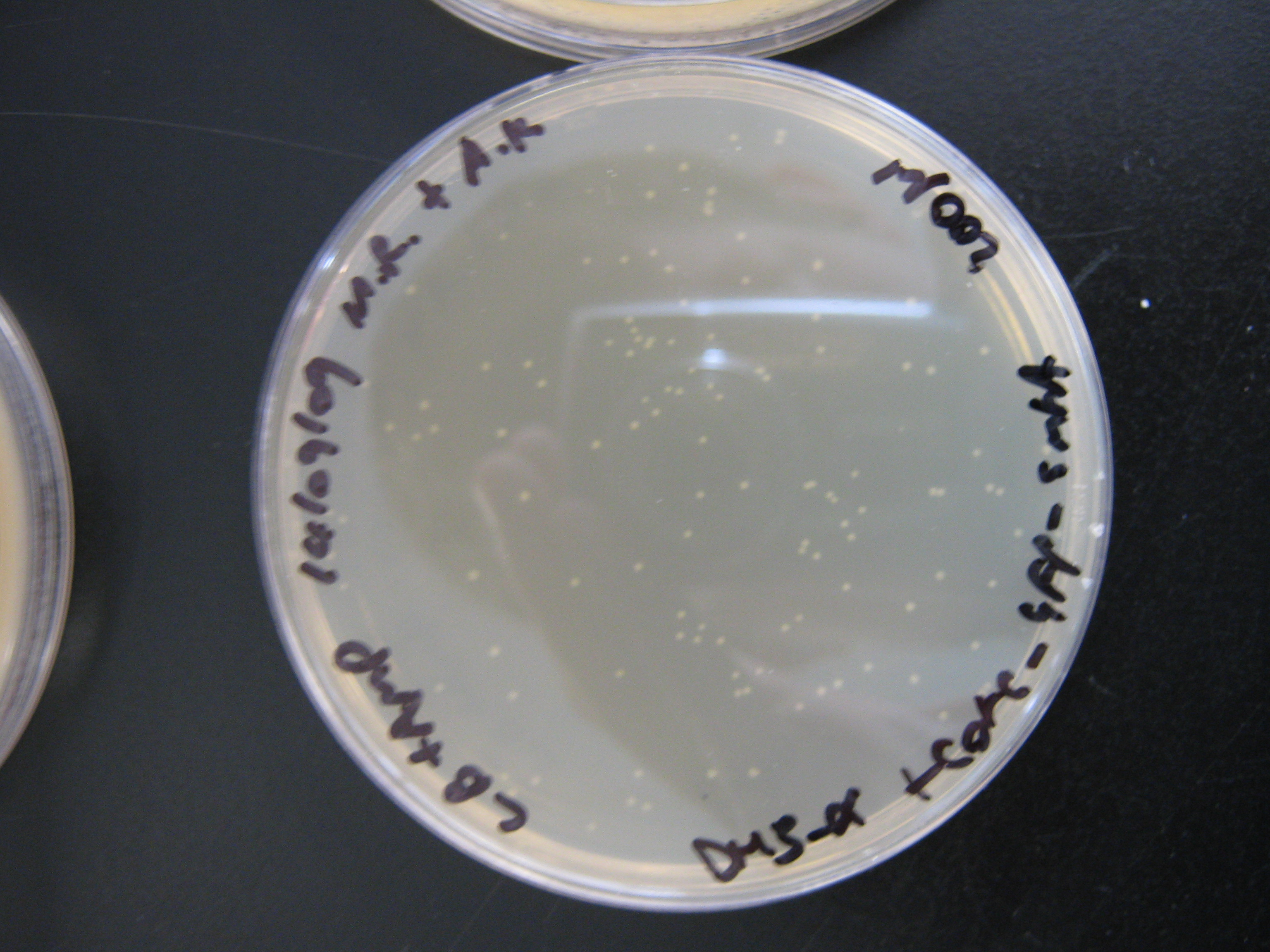
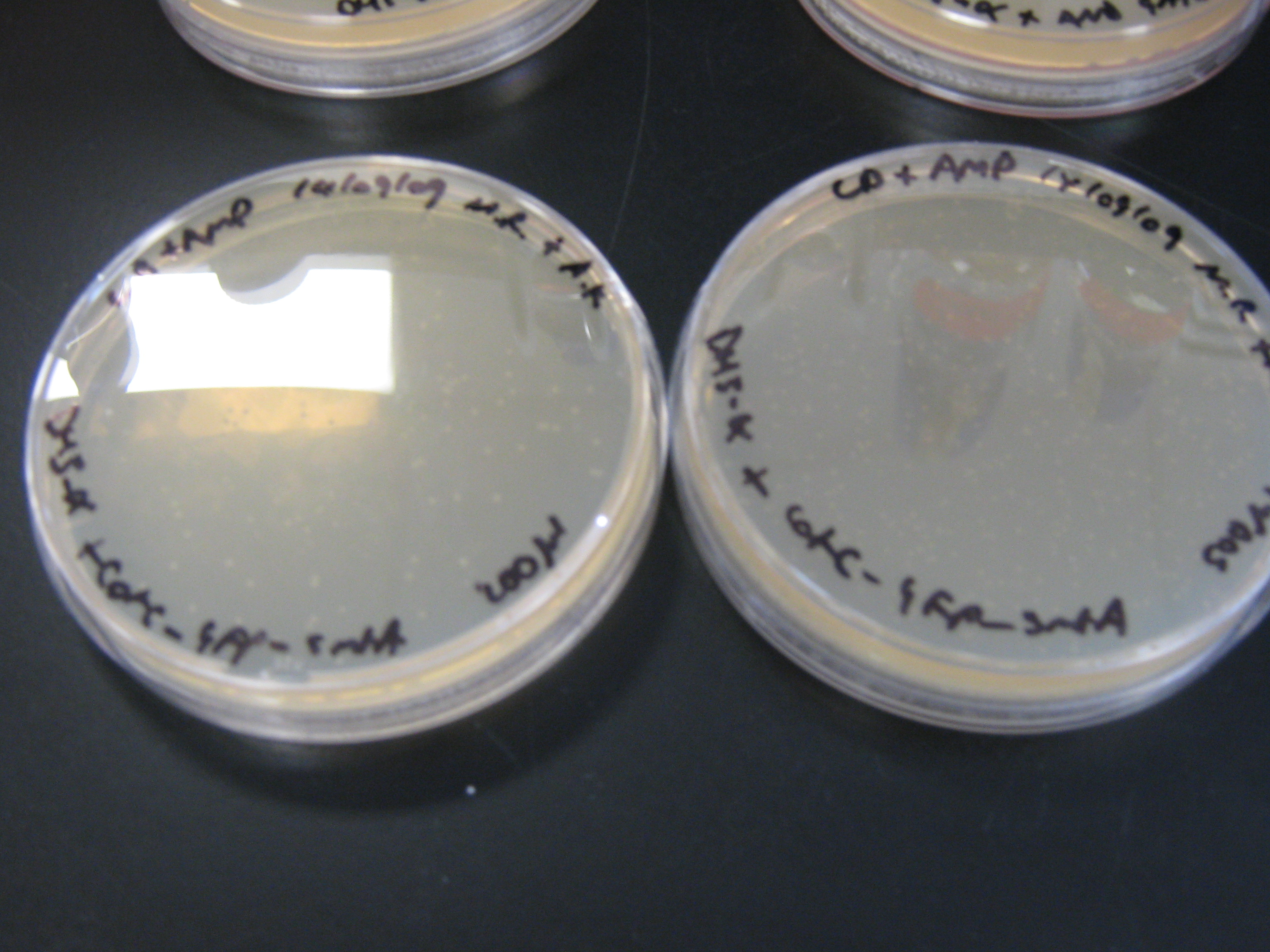
- Check plates for cotC-GFP-smtA transformants and arsR-cadA AND gate transformants
- If successful prepare for mini-preps by picking 12 colonies and using them to inoculate 12 tubes of 5ml LB
- If unsuccessful, reattempt ligations and transformations in E. coli cells.
- Digest the 5 midi-prep samples (cotC-GFP-smtA, kinA, pGFP-rrnB, pMUTIN4 and pSB1AT3) to ensure the DNA is correct.
- Run these digests on gel to analyse.
Observations
It can be seen from the photograph above that there are colonies present on the cotC-GFP-smtA transformant LB + amp plates suggesting that the ligations and transformations were a success. This cannot be said for the cadmium sensor 'transformant' cells which did not yield any colonies on the LB + amp plates.
Procedure
Preparation for cotC transformant mini-preps
To prepare the cotC BioBricks for midi-preps, 12 colonies were picked from the LB + amp plate containing 200ul of transformant cells. These colonies, under aspetic conditions, were then used to inoculate 12 tubes of LB + ampicillin. The tubes were transferred to the orbital incubator for overnight growth at 37ºC.
Digesting midi-prep samples
The digests on the cotC, kinA, pGFP-rrnB, pMUTIN4, and pSB1AT3 midi-prep DNA samples were carried out as follows:
| cotC | kinA | pGFP-rrnB | pMUTIN4 | pSB1AT3 | ||
|---|---|---|---|---|---|---|
| DNA (ul) | 10 | 10 | 10 | 10 | 10 | |
| Buffer (ul) | 2 | 2 | 2 | 2 | 2 | |
| EcoRI (ul) | 1 | 1 | 1 | 0 | 1 | |
| PstI (ul) | 1 | 1 | 0 | 0 | 1 | |
| BamHI (ul) | 0 | 0 | 0 | 1 | 0 | |
| Water | 6 | 6 | 7 | 7 | 6 |
The digest reactions were allowed to run for 1 hour at 37ºC (waterbath).
9ul of each of the samples were taken and placed into 5 Eppendorf tubes. 1ul of loading buffer was added to all samples and mixed. The DNA samples were then loaded into agarose gel and placed under DNA gel electrophoresis for about 1 hour. However the electrophoresis failed because the gel was too thin. Another attempt will be made at a later date.
Ligation of cadA promoter and arsR BioBrick
Transformation of E. coli with ligated product
Competent DH5-alpha E. coli cells were then transformed with the ligated cotC in pMUTIN4 BioBrick. The transformations were carried out according to Dr. Aldridge's transformation protocol and there were only a few changes made:
- Step 2 - the waiting time was 15 minutes instead of 30 minutes
- Step 5 - the waiting/incubation time was also 15 minutes instead of 30 minutes.
Once the transformation process was completed the cells were plated onto two LB + chloramphenicol plates - one plate containing 200ul of transformant cells in LB and the other plate containing 500ul of cells in LB.
Sporulation Tuning/Chassis Team
Introduction
- It has been proved that we got 9 strains which contain right sleB clone. Yesterday, we cultured 50ml cells for Midi Prep. We'll perform a Midi Prep today to get large amount of plasmid with sleB biobrick.
- Also we had get the cwlJ fragment successful, to get the cwlJ biobrick, we'll do the same procedure as we did to sleB.
- To get a double clone of cwlJ:sleB, we need digest plasmid pSB1AT3:sleB and ligate with cwlJ fragment.
Experiment procedure
Midi Prep
- Before we carried out Midi Prep, we take out 1ml of cell mix and put them into 1.5ml eppendorf tube to keep in fridge.
- 50ml of cells were used for Midi Prep.
- Midi prep processed by following the standard protocal of Plasmid Midi Prep Kit(Sigma).
Ligate cwlJ with pSB1AT3 and transform into E.coli
- Ligation procedure was same as the procedure of sleB ligation in day 08/09/09. Ligase and buffer were came from Fast Ligation and Transformation Kit(Fermentas).
Both cwlJ and pSB1AT3 were cut by EcoRI and speI
- Transformation process was carried out by the same procedure in day 09/09/09.
Digest pSB1AT3:sleB plasmid for double clone
- Prepare the digest reaction.
dd H2O 7ul
10X fast digest buffer 7ul
Fast EcoRI 3ul
Fast speI 3ul
pSB1AT3:sleB plasmid 50ul
------------------------------
70ul
- Incubate 1 hour at 37 degree.
- Store at -20C.
Conclusion
- The concentration of our Midi Prep plasmid pSB1AT3:sleB is 109.3ng/ul.
Futher plan
- Ligate cwlJ with pSB1AT3:sleB and transform the ligated plasmid into E.coli.
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
- Newcastle iGEM Twitter
- [http://www.facebook.com/home.php#/group.php?gid=131709337641 Newcastle on Facebook]
- [http://www.youtube.com/user/newcastle2009igem Newcastle Youtube Channel]
 "
"