Team:Groningen/Project/Promoters
From 2009.igem.org
m (→Results) |
Svenjurgens (Talk | contribs) (→Zinc Induced Promoters) |
||
| (5 intermediate revisions not shown) | |||
| Line 205: | Line 205: | ||
homeostasis (Rensing et al., 2000; Rensing and Grass, 2003). CueR, a MerR-family transcription factor, stimulates | homeostasis (Rensing et al., 2000; Rensing and Grass, 2003). CueR, a MerR-family transcription factor, stimulates | ||
copper-induced transcription of both copA encoding Cu(I)-translocating P-type ATPase pump (exporter), that is the central component for maintenance of the copper homeostasis, and cueO encoding a periplasmic multicopper | copper-induced transcription of both copA encoding Cu(I)-translocating P-type ATPase pump (exporter), that is the central component for maintenance of the copper homeostasis, and cueO encoding a periplasmic multicopper | ||
| - | oxidase for detoxification (Outten et al | + | oxidase for detoxification ([[Team:Groningen/Literature#Outten2000|Outten C.E. et al, 2000]]; [[Team:Groningen/Literature#Petersen2000| Petersen and Moller, 2000]])." (from [[Team:Groningen/Literature#Yamamoto|Yamamoto K., 2005) |
Promoter cusCp is associated with the two component system CusR and CusS for the copper induced transcription of genes involved in copper efflux (cusC, cusF, cusB and cusA, which is present on the genome of <i>E. coli </i> str. K-12 substrain MG1655). The sequence shows the typical -10 and -35 region of the promoter and can be found through the following [http://biocyc.org/ECOLI/NEW-IMAGE?type=OPERON&object=TU0-1821 link]. A second region, located at -53.5 from the transcription start site, is thought to bind CusR. Upon binding of CusR, the RNA polymerase is able to recognize the site and attach itself, and can also be found in the same [http://biocyc.org/ECOLI/NEW-IMAGE?type=OPERON&object=TU0-1821 link]. | Promoter cusCp is associated with the two component system CusR and CusS for the copper induced transcription of genes involved in copper efflux (cusC, cusF, cusB and cusA, which is present on the genome of <i>E. coli </i> str. K-12 substrain MG1655). The sequence shows the typical -10 and -35 region of the promoter and can be found through the following [http://biocyc.org/ECOLI/NEW-IMAGE?type=OPERON&object=TU0-1821 link]. A second region, located at -53.5 from the transcription start site, is thought to bind CusR. Upon binding of CusR, the RNA polymerase is able to recognize the site and attach itself, and can also be found in the same [http://biocyc.org/ECOLI/NEW-IMAGE?type=OPERON&object=TU0-1821 link]. | ||
| Line 219: | Line 219: | ||
====Cloning strategy==== | ====Cloning strategy==== | ||
| - | The CueR CueO sensitive promotor was designed by substracting its sequence from the genome database of ''E. coli'' str K12.It's binding region was established by Yamamoto and co worker. The promotor region was designed in silico with its own RBS and the pre and suffix were ''in silico'' cuted with ''Eco''RI and ''Spe''I creating sticky ends. See parts registry {{Part| | + | The CueR CueO sensitive promotor was designed by substracting its sequence from the genome database of ''E. coli'' str K12.It's binding region was established by Yamamoto and co worker. The promotor region was designed in silico with and without its own RBS and the pre and suffix were ''in silico'' cuted with ''Eco''RI and ''Spe''I creating sticky ends. See parts registry {{Part|BBa_K190017}} |
====Results==== | ====Results==== | ||
| Line 236: | Line 236: | ||
===Conclusion=== | ===Conclusion=== | ||
The fluorescence measurements of the CueR promotor show that there is no fluorescence without induction of CuSO<sub>4</sub>. Upon induction with CuSO<sub>4</sub> the cells show an increase in RFP fluorescence which keeps increasing over 22 hours after induction. | The fluorescence measurements of the CueR promotor show that there is no fluorescence without induction of CuSO<sub>4</sub>. Upon induction with CuSO<sub>4</sub> the cells show an increase in RFP fluorescence which keeps increasing over 22 hours after induction. | ||
| - | + | <!-- | |
===Parts Registry=== | ===Parts Registry=== | ||
| Line 260: | Line 260: | ||
''Lactococcus lactis'' <br> | ''Lactococcus lactis'' <br> | ||
'''Abs.''': Two regulatory genes (<i>lcoR</i> and <i>lcoS</i>) were identified from a plasmid-borne lactococcal copper resistance determinant and characterized by transcriptional fusion to the promoterless chloramphenicol acetyltransferase gene (<i>cat</i>). The transcription start site involved in copper induction was mapped by primer extension [[Team:Groningen/Literature#Khunajakr1999|Khunajakr 1999]]. | '''Abs.''': Two regulatory genes (<i>lcoR</i> and <i>lcoS</i>) were identified from a plasmid-borne lactococcal copper resistance determinant and characterized by transcriptional fusion to the promoterless chloramphenicol acetyltransferase gene (<i>cat</i>). The transcription start site involved in copper induction was mapped by primer extension [[Team:Groningen/Literature#Khunajakr1999|Khunajakr 1999]]. | ||
| + | --> | ||
==Zinc Induced Promoters== | ==Zinc Induced Promoters== | ||
| - | Zinc is essential for the functioning of cells, and must be maintained at certain levels within the cell. However, apart from its function, zinc is also harmful at elevated concentrations. Zinc starvation and zinc toxicity both lead to transcription of a number of recently characterized ''E. coli'' genes that encode Zn(II) uptake or export proteins. (from Outten C.E. et al, 1999) | + | Zinc is essential for the functioning of cells, and must be maintained at certain levels within the cell. However, apart from its function, zinc is also harmful at elevated concentrations. Zinc starvation and zinc toxicity both lead to transcription of a number of recently characterized ''E. coli'' genes that encode Zn(II) uptake or export proteins. (from [[Team:Groningen/Literature#Outten1999|Outten C.E. et al, 1999]]) |
| - | ZntR protein found in ''E. coli'', a homologue of MerR, has recently been shown to mediate Zn(II)-responsive regulation of zntA, a gene involved in Zn(II) detoxification. ZntR functions as a zinc receptor that is necessary to activate Zn-responsive transcription at the zntA promoter. ZntR binds in the atypical 20-base pair spacer region of the promoter and distorts the DNA in a manner that is similar to MerR. The addition of Zn(II) to ZntR converts it to a transcriptional activator protein that introduces changes in the DNA conformation. These changes apparently make the promoter a better substrate for RNA polymerase. The ZntR metalloregulatory protein is a direct Zn(II) sensor that catalyzes transcriptional activation of a zinc efflux gene, thus preventing intracellular Zn(II) from exceeding an optimal concentration. (from Outten C.E. et al, 1999 | + | ZntR protein found in ''E. coli'', a homologue of MerR, has recently been shown to mediate Zn(II)-responsive regulation of zntA, a gene involved in Zn(II) detoxification. ZntR functions as a zinc receptor that is necessary to activate Zn-responsive transcription at the zntA promoter. ZntR binds in the atypical 20-base pair spacer region of the promoter and distorts the DNA in a manner that is similar to MerR. The addition of Zn(II) to ZntR converts it to a transcriptional activator protein that introduces changes in the DNA conformation. These changes apparently make the promoter a better substrate for RNA polymerase. The ZntR metalloregulatory protein is a direct Zn(II) sensor that catalyzes transcriptional activation of a zinc efflux gene, thus preventing intracellular Zn(II) from exceeding an optimal concentration. (from [[Team:Groningen/Literature#Outten1999|Outten C.E. et al, 1999]]) |
| - | + | ||
| - | + | ||
| + | The sequence of zntRp has been used to design synthetic oligos ending in biobrick pre- and suffix with ''Eco''RI and ''Spe''I restriction overhangs. The promoter sequence contains the -35 and -10 sequence with the atypical 20-base pair spacer region for binding of ZntR ([http://partsregistry.org/wiki/index.php/Part:BBa_K190016 BBa_K190016]). In addition, the promoter was designed with a RBS found before the zntA gene ([http://partsregistry.org/wiki/index.php/Part:BBa_K190022 BBa_K190022]). The commonly used RBS part ([http://partsregistry.org/wiki/index.php/Part:BBa_B0034 BBa_B0034]) might be to strong and give unwanted leakage of the promoter. | ||
| + | <!-- | ||
====Other organisms==== | ====Other organisms==== | ||
''Bacillus subtilis'' | ''Bacillus subtilis'' | ||
| Line 282: | Line 283: | ||
''Staphylococcus aureus'' | ''Staphylococcus aureus'' | ||
| - | '''Abs.''': In ''Staphylococcus aureus'' CzrA, a member of the ArsR/SmtB family of DNA binding proteins, functions as a repressor of the czr operon, that consists of czrA and the gene encoding the CzcD homologue CzrB (Xiong and Jayaswal, 1998; Kuroda et al., 1999; | + | '''Abs.''': In ''Staphylococcus aureus'' CzrA, a member of the ArsR/SmtB family of DNA binding proteins, functions as a repressor of the czr operon, that consists of czrA and the gene encoding the CzcD homologue CzrB ([[Team:Groningen/Literature#Xiong1998|Xiong and Jayaswal, 1998]]; [[Team:Groningen/Literature#Kuroda1999|Kuroda et al., 1999]]; [[Team:Groningen/Literature#Kuroda1999|Kuroda et al., 1999]]). CzrA-mediated repression is alleviated in the presence of Zn<sup>2+</sup> and Co<sup>2+</sup> ([[Team:Groningen/Literature#Xiong1998|Xiong and Jayaswal, 1998]]; [[Team:Groningen/Literature#Kuroda1999|Kuroda et al., 1999]]; [[Team:Groningen/Literature#Kuroda1999|Kuroda et al., 1999]]). |
| - | + | ||
| + | --> | ||
{{Team:Groningen/Project/Footer}} | {{Team:Groningen/Project/Footer}} | ||
Latest revision as of 23:38, 21 October 2009
[http://2009.igem.org/Team:Groningen http://2009.igem.org/wiki/images/f/f1/Igemhomelogo.png]
|
|---|
- Transport
- Accumulation
- Metal-sensitive Promoters
- Gas Vesicles
PromotersA promoter is a part of DNA involved in the regulation of gene transcription by RNA polymerase. In general RNA polymerase tends to bind weakly to a strand of DNA until a suitable promoter is encountered and the binding becomes strong. Promoters are used to express genes of interest in cells in either a constitutive or induced manner. The constitutive promoters are used when a constant expression of enzymes is desired, and the amount of activity can be regulated by choosing from a range of promoters varying from low to high expression. If, however, expression is desired at certain points in time, or growth stage, inducible promoters are the best choice for regulating gene expression. In our system, we want to induce GVP production when the concentration of desired metal in the cells reaches a certain level. By choosing metal sensitive promoters already present in E. coli cells, the cells contain the necessary components for controlling the promoters, and the promoter sequence has only to be placed in front of the genes of interest.By cloning the ArsR and CueO promotor in front of RFP we have shown that by induction with respectively Arsenite and Copper repression of the promotor is reduced and expression of RFP enhanced. We took the following promoters into consideration:
Arsenic Induced PromotersBecause of the similarity to phosphate, sometimes arsenate is mistaken for phosphate, which is how it is introduced into living organisms, including E. coli, by the phosphate uptake system. Other molecules such as As(III) can also be introduced into the cells by various membrane transporters. E. coliPromoter arsRp is associated with the dimer of ArsR for the arsenic induced transcription of genes involved in arsenic efflux (arsR, arsB and arsC, which is present on the genome of E. coli str. K-12 substrain MG1655). The sequence shows the typical -10 and -35 region of the promoter and can be found through the following [http://biocyc.org/ECOLI/NEW-IMAGE?type=OPERON&object=TU00239 link]. A second region, located at -41.5 from the transcription start site, is thought to bind dimeric ArsR. Upon binding of arsenic, the dimer dissociates and allows the RNA polymerase space to attach itself, and can also be found in the same [http://biocyc.org/ECOLI/NEW-IMAGE?type=OPERON&object=TU00239 link].
(ArsR)2-DNA → ArsR-Ar + ArsR-Ar + DNA → Activation of transription The presence of all genes and promoters on the chromosome of E. coli makes the use of the arsRp for induction of the GVP cluster relatively straight forward. The promoter sequence of arsRp, with the upstream binding box for ArsR dimer, can either be synthesized completely with the required restriction sites, or acquired using PCR and carefully designed primers. It might even be an option to alter the -10/-35 promoter region for higher or lower transcription of the genes. Cloning strategyThe ArsR sensitive promotor was designed by substracting its sequence from the genome database of E. Coli str K12. Its binding region was established by Lee and co workers. The promotor region was designed in silico with its own RBS and the pre and suffix were in silico cuted with EcoRI and SpeI creating sticky ends. See parts registry [http://partsregistry.org/Part:BBa_K190015 BBa_K190015] ResultsThe functionality of pArsR () was tested by using a test construct, composed of pArsR and RFP on (Figure 1). 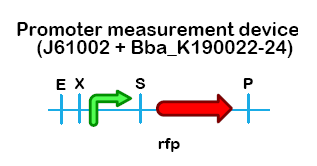
Fluorescence of resting cellsThe fluorescence of the red fluorescent protein was measured as described in protocols. Upon induction of the ArsR promoter the expression of RFP increased, as seen in figure 2. From the enhanced fluorescence a value for the relative promoter unit (RPU) was calculated according to Kelly 2009 (formula 9). Thereby an induction of 2.3 RPU was found, which was in consensus with the promoter activity found for arsenic metal sensitive promoter (used in expression of MTs) (personal communication, Dr. D. Wilcox). The arsenite uptake in E. coli with J61002- over time was measured using the arsenite uptake assay, this was done upon incubation with 10µM NaAsO2. This data was multiplied by the following ratio: As(III) uptake upon induction for 1hr with 100µM As(III) devided by As(III) uptake upon induction for with 10 µM As(III). The increasing intracellular concentration is shown in figure 3. 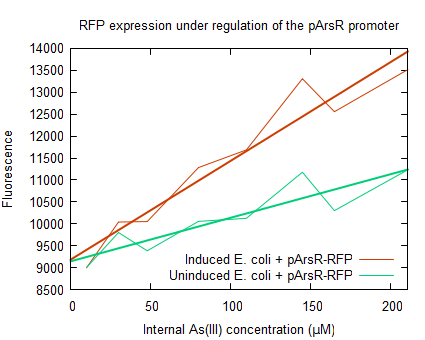
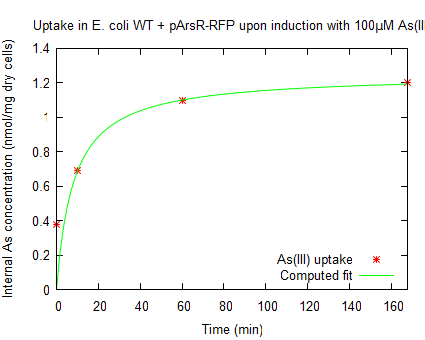
The raw data can be found at downloads. Fluorescence of growing cellsIn order to further characterize the ArsR promotor, measurements were done by inducing cells in the exponential phase. After induction the fluorescence was measured for 22 h. see protocols. The RFP was excited at 580 nm and emission was measured at 600 nm. In order to have a significant high enough signal cells were resuspended at OD600=0.5 in half the volume. The cells were induced to an end concentration of 5000, 500, 50, 5 and 0 µM. The fluorescence normalized to the OD600 is plotted in figure 4. In all measurements [http://partsregistry.org/Part:BBa_J23101 BBa_J23101] was taken along to serve as a reference. 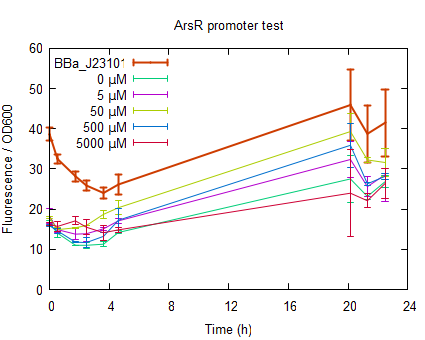
The fluorescence in figure 4 is normalized to the OD600 to correct for differences in cell concentration. As can be seen in figure 4 non induced ArsR RFP (0 µM)is already fluorescent at the time of induction, meaning that the promotor is leaking. What figure 4 also shows is that upon induction the fluorescence increases meaning that the promotor although leaking is less suppresed in the presence of Arsenite. The highest increase in fluorescence is upon induction to a concentration of 50 µM arsenite which is as high as 85% of the fluorescence from reference promotor . Almost all plots show a slight decrease of fluorescence in the beginning due to the recovery of resuspending the cells at 4 °C. Induction to a final concentration of 5000 µM of Arsenite gives after 1 hour already an increase but decreases after 2 hours and shows only a slow increase in fluorescence after 5 hours. Reason for the lower fluorescence intensity of induction to 5000 µM is the poisoning of the cells with Arsenite. The poisoning of the cells is best seen in the OD plotted against time as shown in figure 5. The cells induced to a concentration of 5000 µM Arsenite shows a big decrease in OD between 5 and 22 hours after induction due to Arsenite poisoning. 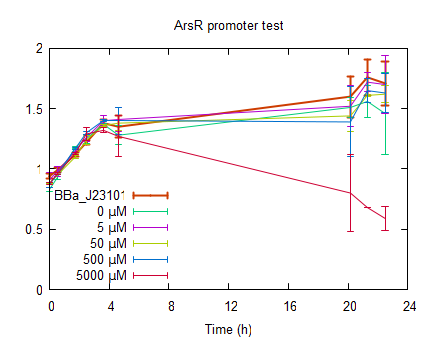
ConclusionBoth promoter tests, with resting cells and growing cells, show clearly that the pArsR promoter is functional. The negative transcriptional regulator ArsR releases the promoter region upon induction with arsenite. The promoter strength was calculated in relative promoter units, upon induction of resting cells with 100 µM As(III) an increase of 2.3 was found. A disadvantage of the usage of pArsR, also clearly shown by the two measurements, is that the negative regulation is leaky as there is already some RFP expressed without addition of arsenite. The OD600 measurements of the growing cell measurements showed that concentrations as high as 5000 µM Arsenite are poisonous for E. coli TOP 10 cells. Modelling
The three graphs below illustrate the promoter response after induction with arsenic (directly in the cell, with the equivalent of 1 µM in the solution) with and without constitutive expression of ArsR (the first two graphs) and with slower production and degradation of ArsR (the two left graphs). Also, each graph has a line showing the formation of a product behind the ars promoter that does not degrade (and has production rate 1), subtracting the production that would have occurred without induction to show the effect of adding arsenic. Some conclusions:
Other organismsBacillus subtilis In B. subtilis, an ArsR family repressor (ArsRBS) responds to As(III) and Sb(III) and regulates the ars operon encoding itself (ArsR), and arsenate reductase (ArsC), an arsenite efflux pump (ArsB) and a protein of unknown function (YqcK). The order in which ArsRBS recognises metals is as follows: As(III)>As(V)>Cd(II)~Ag(I). A second protein, AseR, negatively regulates itself and AseA, an As(III) efflux pump which contributes to arsenite resistance in cells lacking a functional ars operon. The order in which AseR recognises metals is as follows: As(III)>As(V). Copper Induced PromotersCopper is an essential element that becomes highly cytotoxic when concentrations exceed the capacity of cells to sequester the ion. The toxicity of copper is largely due to its tendency to alternate between its cuprous, Cu(I), and cupric, Cu(II), oxidation states, differentiating copper from other trace metals, such as zinc or nickel. Under aerobic conditions, this redox cycling leads to the generation of highly reactive hydroxyl radicals that readily and efficiently damage biomolecules, such as DNA, proteins, and lipids. Most organisms have specialized mechanisms to deal with dangerous levels of heavy metals, like the production of efflux pumps. These genes are regulated by promoters, which are inducible by the respective metals. E. coli"The intracellular level of copper in E. coli is controlled by the export of excess copper, but the entire systems of copper uptake and intracellular copper delivery are not fully understood. Two regulatory systems, the CueR and CusR systems, have been identified to be involved in transcription regulation of the genes for copper homeostasis (Rensing et al., 2000; Rensing and Grass, 2003). CueR, a MerR-family transcription factor, stimulates copper-induced transcription of both copA encoding Cu(I)-translocating P-type ATPase pump (exporter), that is the central component for maintenance of the copper homeostasis, and cueO encoding a periplasmic multicopper oxidase for detoxification (Outten C.E. et al, 2000; Petersen and Moller, 2000)." (from [[Team:Groningen/Literature#Yamamoto|Yamamoto K., 2005) Promoter cusCp is associated with the two component system CusR and CusS for the copper induced transcription of genes involved in copper efflux (cusC, cusF, cusB and cusA, which is present on the genome of E. coli str. K-12 substrain MG1655). The sequence shows the typical -10 and -35 region of the promoter and can be found through the following [http://biocyc.org/ECOLI/NEW-IMAGE?type=OPERON&object=TU0-1821 link]. A second region, located at -53.5 from the transcription start site, is thought to bind CusR. Upon binding of CusR, the RNA polymerase is able to recognize the site and attach itself, and can also be found in the same [http://biocyc.org/ECOLI/NEW-IMAGE?type=OPERON&object=TU0-1821 link].
Cu → CusS → +P → CusR → Activation of transription The problem so far is the site of detection of copper. The CusS protein senses the external copper concentrations and not the internal. For our project it would be nice to have an internal sensor for the induction of the buoyancy genes, so it will float after uptake. In addition to CusR, three other systems involved in copper resistence are present (CueR, CpxR and YedW). Both CpxR and YedW have the same problem of sensing external copper instead of internal copper, CueR is thought to respond to intracellular concentrations of copper. The choice for CusR over CueR would be based on the frequency of binding sites of both on the genome of E. coli (1 vs. 197 times), which gives CusR more chance of binding to our promoter. However, the idea behind our project is to induce gvp transcription at a high intracellular concentration, and results in the CueR related promoter. Cloning strategyThe CueR CueO sensitive promotor was designed by substracting its sequence from the genome database of E. coli str K12.It's binding region was established by Yamamoto and co worker. The promotor region was designed in silico with and without its own RBS and the pre and suffix were in silico cuted with EcoRI and SpeI creating sticky ends. See parts registry [http://partsregistry.org/Part:BBa_K190017 BBa_K190017] ResultsIn order characterize the CueO promotor, measurements were done by inducing cells in the exponential phase. After induction the fluorescence was measured for 22 h. see protocols. The RFP was excited at 580 nm and emission was measured at 600 nm. In order to have a significant high enough signal cells were resuspended at OD600=0.5 in half the volume. The cells were induced to an end concentration of 5000, 500, 50, 5 and 0 µM. The fluorescence normalized to the OD600 is plotted in figure 4. In all measurements [http://partsregistry.org/Part:BBa_J23101 BBa_J23101] was taken along to serve as a reference. 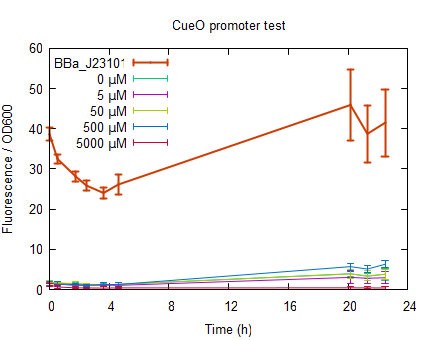
The fluorescence in figure 6 is normalized to the OD600 to correct for differences in cell concentration. As can be seen in figure 6 non induced CueO RFP (0 µM) shows no fluorescence meaning that the promotor is not leaking. The Fluorescence for CuSO4 induced cells shows only slight increase in the order of 0 < 5000 < 5 < 50 < 500 µM CuSO4. The cells induced to a concentration of 5000 µM CuSO4 show no increase in fluorescence which could be due to poisoning of the cells by the CuSO4. In figure 7 can be seen that the OD600 of the copper induced cells is increasing in first 5 hours and then stabilizes or even decreases in case of induction to 5000 µM CuSO4. 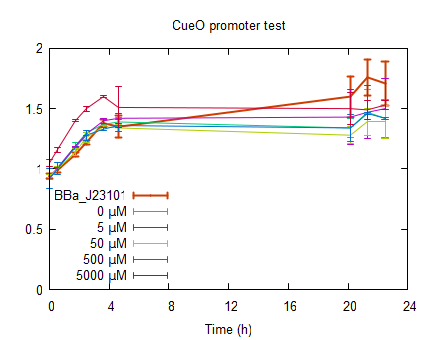
ConclusionThe fluorescence measurements of the CueR promotor show that there is no fluorescence without induction of CuSO4. Upon induction with CuSO4 the cells show an increase in RFP fluorescence which keeps increasing over 22 hours after induction. Zinc Induced PromotersZinc is essential for the functioning of cells, and must be maintained at certain levels within the cell. However, apart from its function, zinc is also harmful at elevated concentrations. Zinc starvation and zinc toxicity both lead to transcription of a number of recently characterized E. coli genes that encode Zn(II) uptake or export proteins. (from Outten C.E. et al, 1999) ZntR protein found in E. coli, a homologue of MerR, has recently been shown to mediate Zn(II)-responsive regulation of zntA, a gene involved in Zn(II) detoxification. ZntR functions as a zinc receptor that is necessary to activate Zn-responsive transcription at the zntA promoter. ZntR binds in the atypical 20-base pair spacer region of the promoter and distorts the DNA in a manner that is similar to MerR. The addition of Zn(II) to ZntR converts it to a transcriptional activator protein that introduces changes in the DNA conformation. These changes apparently make the promoter a better substrate for RNA polymerase. The ZntR metalloregulatory protein is a direct Zn(II) sensor that catalyzes transcriptional activation of a zinc efflux gene, thus preventing intracellular Zn(II) from exceeding an optimal concentration. (from Outten C.E. et al, 1999) The sequence of zntRp has been used to design synthetic oligos ending in biobrick pre- and suffix with EcoRI and SpeI restriction overhangs. The promoter sequence contains the -35 and -10 sequence with the atypical 20-base pair spacer region for binding of ZntR ([http://partsregistry.org/wiki/index.php/Part:BBa_K190016 BBa_K190016]). In addition, the promoter was designed with a RBS found before the zntA gene ([http://partsregistry.org/wiki/index.php/Part:BBa_K190022 BBa_K190022]). The commonly used RBS part ([http://partsregistry.org/wiki/index.php/Part:BBa_B0034 BBa_B0034]) might be to strong and give unwanted leakage of the promoter.
|
 "
"






