Team:Brown/Project Histamine Sensor
From 2009.igem.org
(→Histamine Sensor) |
|||
| (45 intermediate revisions not shown) | |||
| Line 6: | Line 6: | ||
| - | = | + | <html><img src="http://img10.imageshack.us/img10/8299/thehistaminesensor.png"></html> |
| - | + | ||
| - | + | ||
During the allergic response, the concentration of histamine in the extracellular fluid of the nasal cavity increases. To initiate a response; therefore, a histamine sensor is necessary. | During the allergic response, the concentration of histamine in the extracellular fluid of the nasal cavity increases. To initiate a response; therefore, a histamine sensor is necessary. | ||
| Line 14: | Line 12: | ||
Because natural histamine receptors exist only in eukaryotic cells as G-coupled protein receptors, they are unusable for our prokaryotic system. Therefore we have set out to engineer our own receptor. This novel receptor will sense extracellular concentrations of histamine and initiate an intracellular cascade, signaling cells to respond appropriately to the increase in histamine concentration. | Because natural histamine receptors exist only in eukaryotic cells as G-coupled protein receptors, they are unusable for our prokaryotic system. Therefore we have set out to engineer our own receptor. This novel receptor will sense extracellular concentrations of histamine and initiate an intracellular cascade, signaling cells to respond appropriately to the increase in histamine concentration. | ||
| - | To engineer a novel histamine receptor, we are mutating two existing prokaryotic chemoreceptors so that they bind histamine rather than their wild type ligands. | + | To engineer a novel histamine receptor, we are mutating two existing prokaryotic chemoreceptors so that they bind histamine rather than their wild type ligands. |
| + | = Re-engineering Chemoreceptor #1: Ribose Binding Protein = | ||
| - | |||
| - | We modified ribose binding protein (RBP), which normally binds ribose in the periplasmic space of Escherichia coli, to bind histamine. Our computational approach to accomplish this task was modeled after that taken in Looger et al., "Computational design of receptor and sensor proteins with novel functions" (2003). Using Rosetta macromolecular modeling software, we modified the program's existing Enzyme Design Function (such as that used in Röthlisberger et al "Kemp elimination catalysts by computational enzyme design" (2008)) to enable the re-design of RBP, a non-enzymatic protein. The successful modification of RBP would result in its ability to bind histamine. | + | We modified ribose binding protein (RBP), which normally binds ribose in the periplasmic space of Escherichia coli, to bind histamine. Our computational approach to accomplish this task was modeled after that taken in Looger, et al., "Computational design of receptor and sensor proteins with novel functions", Nature (2003). Using Rosetta macromolecular modeling software, we modified the program's existing Enzyme Design Function (such as that used in Röthlisberger, et al "Kemp elimination catalysts by computational enzyme design", Nature (2008)) to enable the re-design of RBP, a non-enzymatic protein. The successful modification of RBP would result in its ability to bind histamine. [[Image:Rbp img.png|400px|thumb|The native E. coli ribose-binding protein]] |
| - | + | '''<big>Protein design:</big>''' | |
| - | + | ||
| - | ''' | + | |
1) Took the PDB file for the crystal structure of RBP cocrystallized with ribose (2DRI). Removed water molecules and added missing hydrogens. | 1) Took the PDB file for the crystal structure of RBP cocrystallized with ribose (2DRI). Removed water molecules and added missing hydrogens. | ||
| Line 29: | Line 25: | ||
2) Used UCSF Chimera to geometrically search for all van der Waals interactions between ribose and RBP in the crystal structure. Identified the amino acids responsible for these interactions as those most likely present in the ligand binding pocket of RBP. | 2) Used UCSF Chimera to geometrically search for all van der Waals interactions between ribose and RBP in the crystal structure. Identified the amino acids responsible for these interactions as those most likely present in the ligand binding pocket of RBP. | ||
| - | 3) Used UCSF Chimera to mutate all the identified residues in RBP to alanine (which has neutral chemical properties, and almost no side | + | 3) Used UCSF Chimera to mutate all the identified residues in RBP to alanine (which has neutral chemical properties, and almost no side chain), thereby effectively creating a "blank" version of RBP that has no specific binding pocket for any ligand (removing its binding affinity for ribose)--called the polyala RBP. [[Image:polyala_rbp_img.png|300px|thumb|The polyala RBP.]] |
| - | 4) | + | 4) Replace ribose in the alanine-mutated RBP (polyala RBP) PDB file with a 3D structure of histamine in a low energy conformation. |
| - | + | ||
| - | 5) | + | 5) Used Rosetta's Ligand Docking mode on 100 of Brown's Center for Computational Molecular Biology (http://www.brown.edu/Research/CCMB/) clustered servers to simulate the docking of histamine into the polyala RBP. To dock histamine into polyala RBP, Rosetta Ligand uses Monte Carlo minimization to find the relative orientations of the ligand and receptor that minimize steric contacts between the two while still keeping histamine roughly within the original ligand binding pocket. This produces 10000 PDB files of histamine docked to the polyala RBP. |
| - | 6) | + | 6) Next, we sorted the 10000 docked PDB files by their interface energies between ligand and protein. We selected the top 2500. |
| - | + | ||
| + | 7) Next, we took the top 2500 and input them into the Rosetta Enzyme Design mode, run on the same cluster. We specified the residues mutated to alanine as those to re-design by mutation. The redesign process attempts to stabilize the protein-ligand pair by mutating residues at certain specified locations, those we originally mutated to alanine. This redesign process used Rosetta Design’s probabilistic simulated annealing algorithm to find the particular residues that minimize total energy between protein and ligand (minimized energy indicates a stable state, favoring binding). Because this process is probabilistic, final designs are not guaranteed to yield the lowest possible energy conformations. However, by doing thousands of designs in parallel, we increased the likelihood of isolating mutations that result in histamine binding. | ||
| - | + | 8) Finally, we sorted through the output designs using several criteria: predicted interface energies between the protein and histamine, amount of hydrogen bonding between the protein and histamine (H-bonds are very good for ligand binding), and predicted folding ability of protein. We filtered the final designs to only those with a total score better than or equal to that of the scaffold RBP. We then set cutoffs for several of the relevant criteria, found the sets of the best designs in each relevant category, and found the designs that were in the intersection of those sets. The final design (in bold) selected had the best overall scores in each category. We had the DNA for the top design synthesized by GeneArt AG. We are in the process of testing the protein’s ability to bind histamine and its stability/folding ability. | |
| - | + | [[Image:Final_sel.png|800px|center|thumb|The final selection list of mRBP.]] | |
| + | [[Image:native_score.png|800px|center|thumb|The score of the native RBP]] | ||
| - | + | {| | |
| - | + | !total_score | |
| - | + | !tot_pstat_pm | |
| + | !tot_nlpstat_pm | ||
| + | !tot_burunsat_pm | ||
| + | !tot_hbond | ||
| + | !SR_1_interf_E_1_2 | ||
| + | !SR_1_dasa_1_2 | ||
| + | |- | ||
| + | |Weighted sum of the other terms. (Lower better) | ||
| + | |How well packed the protein is. (0-1) | ||
| + | |How well packed the protein is without the ligand. (0-1) | ||
| + | |Number buried unsatisfied polars. (Lower better) | ||
| + | |Number of H-bonds. (Higher better) | ||
| + | |How well the ligand binds to protein. (Lower better) | ||
| + | |How exposed the ligand is. (0-1) | ||
| + | |} | ||
| + | |||
| + | [[Image:MRBP.png|500px|center|thumb|The mutated Ribose Binding Protein (mRBP)]][[Image:ligandbindingprotein.png|500px|center|thumb|Histamine in the mRBP ligand binding pocket]] | ||
| + | <br/> | ||
| + | <br/> | ||
| + | |||
| + | |||
| + | [[Image:ompc_envz_pathway.png|400px|thumb|The TrgEnvZ-OmpR pathway. From Fig. 3 of Looger, et al., "Computational design of receptor and sensor proteins with novel functions", Nature (2003).]] | ||
In normal E. coli cells, ribose binding to RBP forms a ribose-RBP complex that interacts with the periplasmic domain of Trg, which in turn activates an intracellular cascade that induces chemotaxis. | In normal E. coli cells, ribose binding to RBP forms a ribose-RBP complex that interacts with the periplasmic domain of Trg, which in turn activates an intracellular cascade that induces chemotaxis. | ||
| - | To link activation of Trg to induction of gene expression, we created a fusion between Trg periplasmic receptor domain and the EnvZ intracellular kinase domain by stitching the two intracellular domains at a shared NdeI restriction site. (as done in Baumgartner JW, et al. “Transmembrane signalling by a hybrid protein: communication from the domain of chemoreceptor Trg that recognizes sugar-binding proteins to the kinase/phosphatase domain of osmosensor EnvZ” (1994)). EnvZ phosphorylates the transcription factor OmpR, which causes transcription of DNA regulated by OmpC promoter. | + | To link activation of Trg to induction of gene expression, we created a fusion between Trg periplasmic receptor domain and the EnvZ intracellular kinase domain by stitching the two intracellular domains at a shared NdeI restriction site. (as done in Baumgartner JW, et al. “Transmembrane signalling by a hybrid protein: communication from the domain of chemoreceptor Trg that recognizes sugar-binding proteins to the kinase/phosphatase domain of osmosensor EnvZ” (1994)). EnvZ phosphorylates the transcription factor OmpR, which causes transcription of DNA regulated by OmpC promoter. |
| - | + | = Re-engineering Chemoreceptor #2: Tar Receptor = | |
In parallel, we modified the receptor Tar, which normally binds aspartate in the periplasmic space of E. coli, to bind histamine. | In parallel, we modified the receptor Tar, which normally binds aspartate in the periplasmic space of E. coli, to bind histamine. | ||
| - | Loren Looger at the Howard Hughes Medical Institute's Janelia Farm campus used his protein design software Chameleon to calculate mutations that would transform Tar’s aspartate binding pocket to a histamine binding pocket. His algorithm gave us the top 16 receptor designs and we are currently in the process of creating this library of mutants. We have designed primers for each of these designs and are introducing these mutations by both the “Round-the-Horn Site-Directed Mutagenesis” protocol on OpenWetWare and Strategene’s Quikchange Mutagenesis II Kit. | + | Dr. Loren Looger at the Howard Hughes Medical Institute's Janelia Farm campus used his protein design software Chameleon to calculate mutations that would transform Tar’s aspartate binding pocket to a histamine binding pocket. His algorithm gave us the top 16 receptor designs and we are currently in the process of creating this library of mutants. We have designed primers for each of these designs and are introducing these mutations by both the “Round-the-Horn Site-Directed Mutagenesis” protocol on OpenWetWare and Strategene’s Quikchange Mutagenesis II Kit. |
| - | + | In normal E. coli cells, aspartate binds to the Tar receptor. The Tar-EnvZ chimera protein (created by Utsumi et. al in "Activation of bacterial porin gene expression by a chimeric signal transducer in response to aspartate" 1989) allowed the Taz protein to be linked to the EnvZ cascade in the same manner as Trg-EnvZ. Just as in that system, ligand binding to its receptor leads to an intracellular signaling cascade promoting gene transcription. | |
| - | + | = Initiating an Appropriate Response: Linking Histamine Sensation to Intracellular Transcription = | |
| - | '''Assay to Test | + | The creation of a novel histamine receptor to detect elevated histamine is only the first step to providing an appropriate cell response to allergens. Once a functional histamine sensor is inserted into the cell membrane, its activation must be linked to an intracellular cascade responsible for triggering [https://2009.igem.org/Team:Brown/Project_HBP gene transcription]. The successful mutation of the binding pockets on both RBP and Tar was therefore followed by the complete construction of each receptor as it would function in the cell membrane. For both RBP and Tar, this meant replicating the work done by Masayori Inouye et. al to create the chimeric proteins Trg-EnvZ (Trz) and Tar-EnvZ (Taz) (respectively). Chimeric proteins were created by fusing the intracellular domains of Tar and Trg (Trg is the membrane receptor associated with ligand-bound RBP) to the kinase domain of EnvZ. Activation of the functional receptor domains by histamine binding is thus able to initiate an intracellular cascade that phosphorylates the transcription factor ompR thereby activating gene transcription under the ompC promoter. By replacing the gene normally present under this promoter with our gene of interest, we successfully manipulated the cascade to produce an appropriate cellular response when allergens are present. |
| + | |||
| + | '''<big>Assay to Test Signaling Cascade Functionality</big>''' | ||
Our assay to test these receptors’ affinities for histamine is based on fluorescence. Because the intracellular cascades of both Trg-EnvZ and Tar-EnvZ induce transcription under the OmpC promoter, we have constructed a cassette that places the OmpC promoter over the RFP gene for red fluorescence. Testing for functionality of the cascade then simply involved observing whether RFP expression occurred under ompC, a simple fluorescence assay conducted on an epifluorescence microscope. Qualitative visualization of red-fluorescing colonies transformed with both the receptor and cascade components indicated a functional intracellular signaling system. | Our assay to test these receptors’ affinities for histamine is based on fluorescence. Because the intracellular cascades of both Trg-EnvZ and Tar-EnvZ induce transcription under the OmpC promoter, we have constructed a cassette that places the OmpC promoter over the RFP gene for red fluorescence. Testing for functionality of the cascade then simply involved observing whether RFP expression occurred under ompC, a simple fluorescence assay conducted on an epifluorescence microscope. Qualitative visualization of red-fluorescing colonies transformed with both the receptor and cascade components indicated a functional intracellular signaling system. | ||
| Line 89: | Line 108: | ||
[[Image:RU1012 BL.jpg|300px]] [[Image:RU1012 RFP.jpg|300px]] | [[Image:RU1012 BL.jpg|300px]] [[Image:RU1012 RFP.jpg|300px]] | ||
| + | |||
2. RU1012 with OmpC-RFP | 2. RU1012 with OmpC-RFP | ||
[[Image:RU1012 OmpC BL.jpg|300px]] [[Image:RU1012 OmpC RFP.jpg|300px]] | [[Image:RU1012 OmpC BL.jpg|300px]] [[Image:RU1012 OmpC RFP.jpg|300px]] | ||
| + | |||
3. RU1012 with Tar-EnvZ | 3. RU1012 with Tar-EnvZ | ||
[[Image:RU1012 Taz BL.jpg|300px]] [[Image:RU1012 Taz RFP.jpg|300px]] | [[Image:RU1012 Taz BL.jpg|300px]] [[Image:RU1012 Taz RFP.jpg|300px]] | ||
| + | |||
4. RU1012 with Trg-EnvZ | 4. RU1012 with Trg-EnvZ | ||
| - | [[Image:RU1012 Trg RFP.jpg|300px]] | + | [[Image:RU1012 Trg BL.jpg|300px]][[Image:RU1012 Trg RFP.jpg|300px]] |
| + | |||
5. RU1012 with OmpC-RFP + Tar-EnvZ | 5. RU1012 with OmpC-RFP + Tar-EnvZ | ||
[[Image:RU1012 Taz OmpC BL.jpg|300px]] [[Image:RU1012 Taz OmpC RFP.jpg|300px]] | [[Image:RU1012 Taz OmpC BL.jpg|300px]] [[Image:RU1012 Taz OmpC RFP.jpg|300px]] | ||
| + | |||
6. RU1012 with OmpC-RFP + Trg-EnvZ | 6. RU1012 with OmpC-RFP + Trg-EnvZ | ||
[[Image:RU1012 Trg OmpC BL.jpg|300px]] [[Image:RU1012 Trg OmpC RFP.jpg|300px]] | [[Image:RU1012 Trg OmpC BL.jpg|300px]] [[Image:RU1012 Trg OmpC RFP.jpg|300px]] | ||
| + | |||
| + | ---- | ||
| + | |||
| + | |||
| + | ''<big>References</big>'' | ||
| + | |||
| + | * Baumgartner, James W., Changhoon Kim, Renée E. Brisette, Masoyuri Inouye, Changkyu Park, and Gerald L. Hazelbauer. "Transmembrane signalling by a hybrid protein: communication from the domain of chemoreceptor Trg that recognizes sugar-binding proteins to the kinase/phosphatase domain of osmosensor EnvZ." Journal of Bacteriology 176: 1157-163. Journal of Bacteriology. Web. 25 July 2009. <http://www.ncbi.nlm.nih.gov:80/pmc/articles/PMC205168/>. | ||
| + | |||
| + | * Looger, Loren L., Mary A. Dwyer, James J. Smith, and Homme H. Hellinga. "Computational design of receptor and sensor proteins with novel functions." Nature 423 (2003): 185-90. PubMed. Web. 15 July 2009. <http://www.ncbi.nlm.nih.gov/pubmed/12736688>. | ||
| + | |||
| + | * Meiler, Jens, and David Baker. "ROSETTALIGAND: protein-small molecule docking with full side-chain flexibility." Protein 65.3 (2006): 538-48. PubMed. Web. 12 Aug. 2009. <http://www.ncbi.nlm.nih.gov/pubmed/16972285>. | ||
| + | |||
| + | * Rhiju, Das, and David Baker. "Macromolecular modeling with rosetta." Annual Review of Biochemistry 77: 368-82. PubMed. Web. 12 July 2009. <http://www.ncbi.nlm.nih.gov/pubmed/18410248>. | ||
| + | |||
| + | * Rothlisberger, Daniela, Olga Kersonsky, Andrew M. Wollacott, Lin Jiang, Jason Dechancie, Jamie Betger, Jasmine L. Gallaher, Eric A. Althoff, Alexandre Zanghellini, Orly Dym, Shira Albeck, Kendall N. Houk, Dan S. Tawfik, and David Baker. "Kemp elimination catalysts by computational enzyme design." Nature 453 (2008): 190-95. Nature. 19 Mar. 2008. Web. 5 July 2009. <http://www.nature.com/nature/journal/v453/n7192/abs/nature06879.html>. | ||
| + | |||
| + | * Utsumi, R., R. E. Brisette, A. Rampersaud, S. A. Forst, K. Oosawa, and M. Inouye. "Activation of bacterial porin gene expression by a chimeric signal transducer in response to aspartate." Science 245.4923 (1989): 1246-249. Science. Web. 27 June 2009. <http://www.sciencemag.org/cgi/content/abstract/245/4923/1246>. | ||
| + | |||
| + | <html> | ||
| + | <a href="https://2009.igem.org/Team:Brown/Project_HBP"> | ||
| + | <img src="https://static.igem.org/mediawiki/2009/d/d8/Brown_hbp_bottom_3.png"> | ||
| + | </html> | ||
Latest revision as of 03:50, 22 October 2009

During the allergic response, the concentration of histamine in the extracellular fluid of the nasal cavity increases. To initiate a response; therefore, a histamine sensor is necessary.
Because natural histamine receptors exist only in eukaryotic cells as G-coupled protein receptors, they are unusable for our prokaryotic system. Therefore we have set out to engineer our own receptor. This novel receptor will sense extracellular concentrations of histamine and initiate an intracellular cascade, signaling cells to respond appropriately to the increase in histamine concentration.
To engineer a novel histamine receptor, we are mutating two existing prokaryotic chemoreceptors so that they bind histamine rather than their wild type ligands.
Re-engineering Chemoreceptor #1: Ribose Binding Protein
We modified ribose binding protein (RBP), which normally binds ribose in the periplasmic space of Escherichia coli, to bind histamine. Our computational approach to accomplish this task was modeled after that taken in Looger, et al., "Computational design of receptor and sensor proteins with novel functions", Nature (2003). Using Rosetta macromolecular modeling software, we modified the program's existing Enzyme Design Function (such as that used in Röthlisberger, et al "Kemp elimination catalysts by computational enzyme design", Nature (2008)) to enable the re-design of RBP, a non-enzymatic protein. The successful modification of RBP would result in its ability to bind histamine.Protein design:
1) Took the PDB file for the crystal structure of RBP cocrystallized with ribose (2DRI). Removed water molecules and added missing hydrogens.
2) Used UCSF Chimera to geometrically search for all van der Waals interactions between ribose and RBP in the crystal structure. Identified the amino acids responsible for these interactions as those most likely present in the ligand binding pocket of RBP.
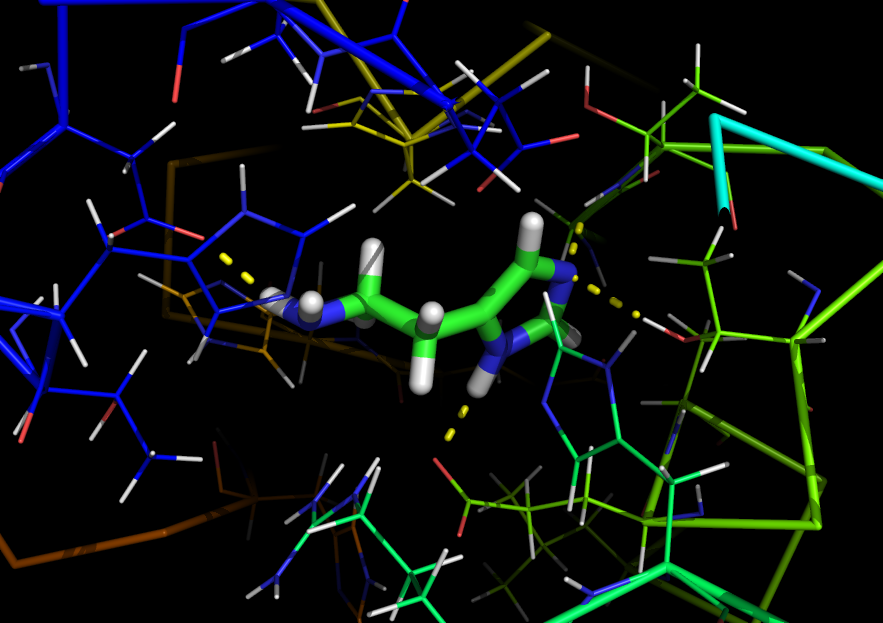
3) Used UCSF Chimera to mutate all the identified residues in RBP to alanine (which has neutral chemical properties, and almost no side chain), thereby effectively creating a "blank" version of RBP that has no specific binding pocket for any ligand (removing its binding affinity for ribose)--called the polyala RBP.4) Replace ribose in the alanine-mutated RBP (polyala RBP) PDB file with a 3D structure of histamine in a low energy conformation.
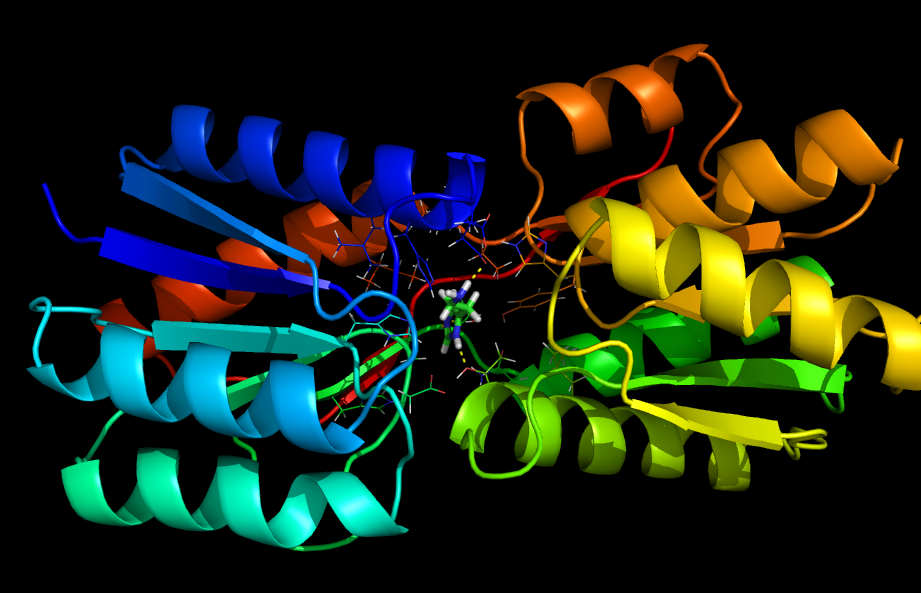
5) Used Rosetta's Ligand Docking mode on 100 of Brown's Center for Computational Molecular Biology (http://www.brown.edu/Research/CCMB/) clustered servers to simulate the docking of histamine into the polyala RBP. To dock histamine into polyala RBP, Rosetta Ligand uses Monte Carlo minimization to find the relative orientations of the ligand and receptor that minimize steric contacts between the two while still keeping histamine roughly within the original ligand binding pocket. This produces 10000 PDB files of histamine docked to the polyala RBP.
6) Next, we sorted the 10000 docked PDB files by their interface energies between ligand and protein. We selected the top 2500.
7) Next, we took the top 2500 and input them into the Rosetta Enzyme Design mode, run on the same cluster. We specified the residues mutated to alanine as those to re-design by mutation. The redesign process attempts to stabilize the protein-ligand pair by mutating residues at certain specified locations, those we originally mutated to alanine. This redesign process used Rosetta Design’s probabilistic simulated annealing algorithm to find the particular residues that minimize total energy between protein and ligand (minimized energy indicates a stable state, favoring binding). Because this process is probabilistic, final designs are not guaranteed to yield the lowest possible energy conformations. However, by doing thousands of designs in parallel, we increased the likelihood of isolating mutations that result in histamine binding.
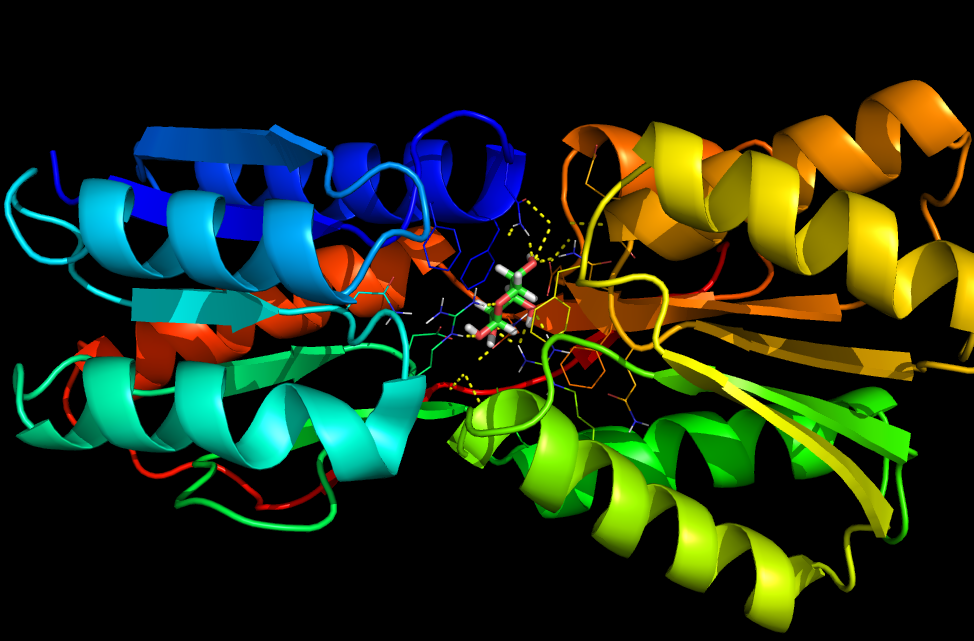
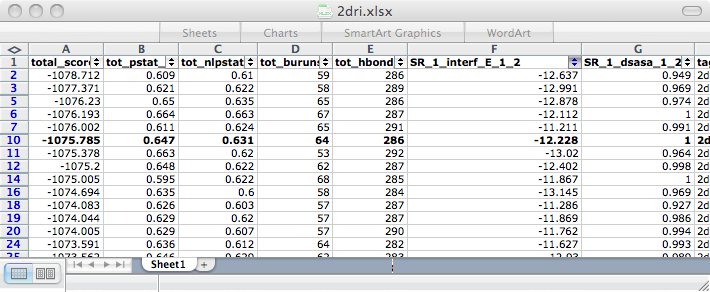
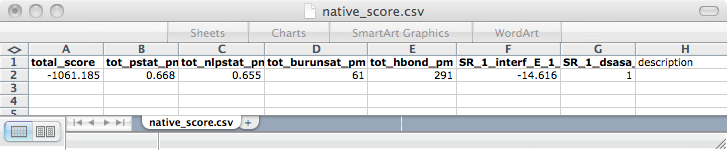
8) Finally, we sorted through the output designs using several criteria: predicted interface energies between the protein and histamine, amount of hydrogen bonding between the protein and histamine (H-bonds are very good for ligand binding), and predicted folding ability of protein. We filtered the final designs to only those with a total score better than or equal to that of the scaffold RBP. We then set cutoffs for several of the relevant criteria, found the sets of the best designs in each relevant category, and found the designs that were in the intersection of those sets. The final design (in bold) selected had the best overall scores in each category. We had the DNA for the top design synthesized by GeneArt AG. We are in the process of testing the protein’s ability to bind histamine and its stability/folding ability.
| total_score | tot_pstat_pm | tot_nlpstat_pm | tot_burunsat_pm | tot_hbond | SR_1_interf_E_1_2 | SR_1_dasa_1_2 |
|---|---|---|---|---|---|---|
| Weighted sum of the other terms. (Lower better) | How well packed the protein is. (0-1) | How well packed the protein is without the ligand. (0-1) | Number buried unsatisfied polars. (Lower better) | Number of H-bonds. (Higher better) | How well the ligand binds to protein. (Lower better) | How exposed the ligand is. (0-1) |
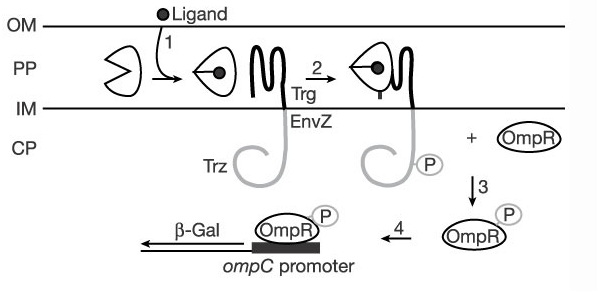
In normal E. coli cells, ribose binding to RBP forms a ribose-RBP complex that interacts with the periplasmic domain of Trg, which in turn activates an intracellular cascade that induces chemotaxis. To link activation of Trg to induction of gene expression, we created a fusion between Trg periplasmic receptor domain and the EnvZ intracellular kinase domain by stitching the two intracellular domains at a shared NdeI restriction site. (as done in Baumgartner JW, et al. “Transmembrane signalling by a hybrid protein: communication from the domain of chemoreceptor Trg that recognizes sugar-binding proteins to the kinase/phosphatase domain of osmosensor EnvZ” (1994)). EnvZ phosphorylates the transcription factor OmpR, which causes transcription of DNA regulated by OmpC promoter.
Re-engineering Chemoreceptor #2: Tar Receptor
In parallel, we modified the receptor Tar, which normally binds aspartate in the periplasmic space of E. coli, to bind histamine.
Dr. Loren Looger at the Howard Hughes Medical Institute's Janelia Farm campus used his protein design software Chameleon to calculate mutations that would transform Tar’s aspartate binding pocket to a histamine binding pocket. His algorithm gave us the top 16 receptor designs and we are currently in the process of creating this library of mutants. We have designed primers for each of these designs and are introducing these mutations by both the “Round-the-Horn Site-Directed Mutagenesis” protocol on OpenWetWare and Strategene’s Quikchange Mutagenesis II Kit.
In normal E. coli cells, aspartate binds to the Tar receptor. The Tar-EnvZ chimera protein (created by Utsumi et. al in "Activation of bacterial porin gene expression by a chimeric signal transducer in response to aspartate" 1989) allowed the Taz protein to be linked to the EnvZ cascade in the same manner as Trg-EnvZ. Just as in that system, ligand binding to its receptor leads to an intracellular signaling cascade promoting gene transcription.
Initiating an Appropriate Response: Linking Histamine Sensation to Intracellular Transcription
The creation of a novel histamine receptor to detect elevated histamine is only the first step to providing an appropriate cell response to allergens. Once a functional histamine sensor is inserted into the cell membrane, its activation must be linked to an intracellular cascade responsible for triggering gene transcription. The successful mutation of the binding pockets on both RBP and Tar was therefore followed by the complete construction of each receptor as it would function in the cell membrane. For both RBP and Tar, this meant replicating the work done by Masayori Inouye et. al to create the chimeric proteins Trg-EnvZ (Trz) and Tar-EnvZ (Taz) (respectively). Chimeric proteins were created by fusing the intracellular domains of Tar and Trg (Trg is the membrane receptor associated with ligand-bound RBP) to the kinase domain of EnvZ. Activation of the functional receptor domains by histamine binding is thus able to initiate an intracellular cascade that phosphorylates the transcription factor ompR thereby activating gene transcription under the ompC promoter. By replacing the gene normally present under this promoter with our gene of interest, we successfully manipulated the cascade to produce an appropriate cellular response when allergens are present.
Assay to Test Signaling Cascade Functionality
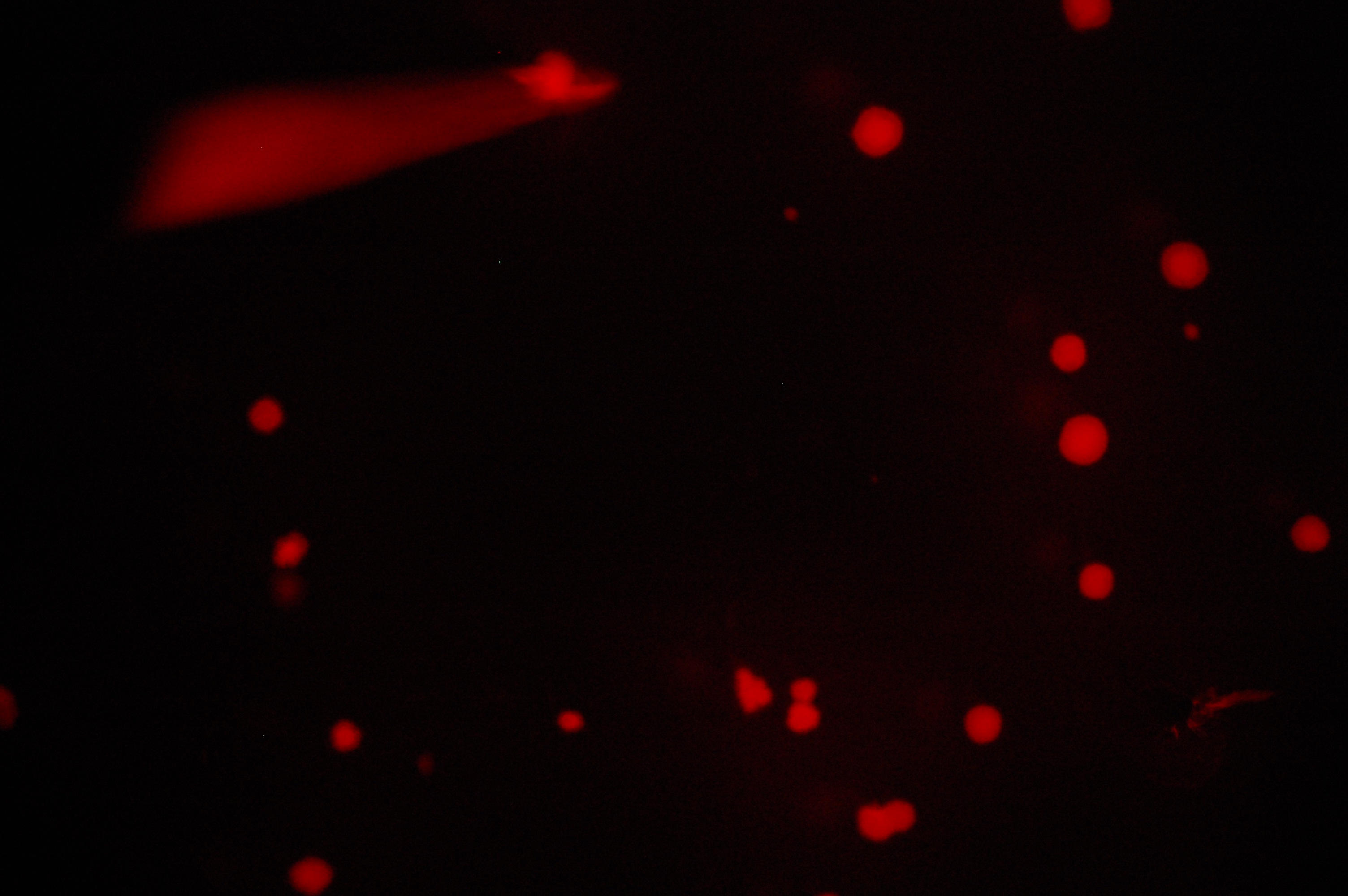
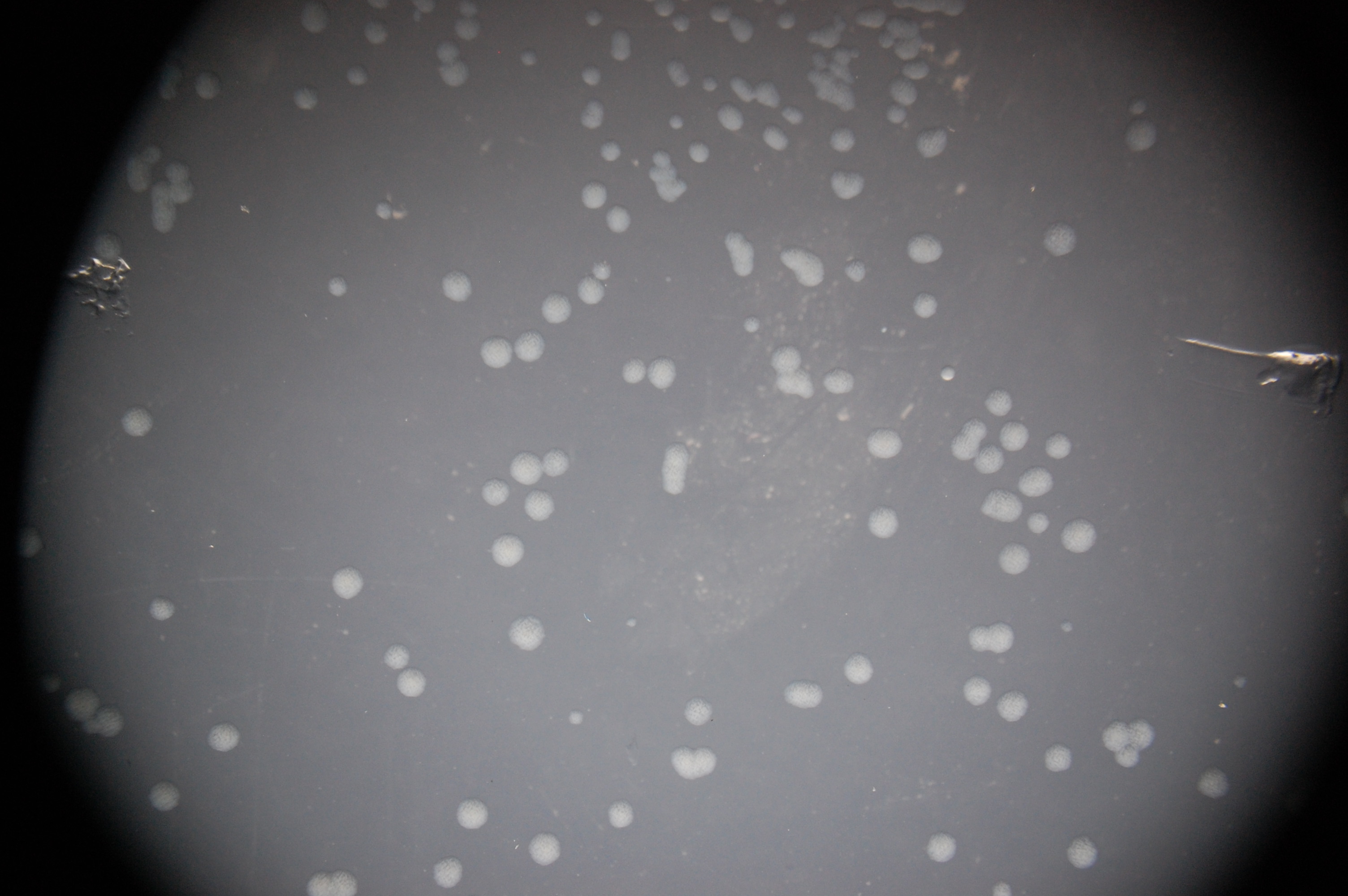
Our assay to test these receptors’ affinities for histamine is based on fluorescence. Because the intracellular cascades of both Trg-EnvZ and Tar-EnvZ induce transcription under the OmpC promoter, we have constructed a cassette that places the OmpC promoter over the RFP gene for red fluorescence. Testing for functionality of the cascade then simply involved observing whether RFP expression occurred under ompC, a simple fluorescence assay conducted on an epifluorescence microscope. Qualitative visualization of red-fluorescing colonies transformed with both the receptor and cascade components indicated a functional intracellular signaling system.
We have tested this signaling cascade by performing the following series of transformations. The E. coli strain RU1012 was used, because it is an EnvZ knockout strain. As efforts to construct a novel histamine receptor were being conducted in parallel to these assays, testing of the cascade was conducted with the original chimeric chemoreceptor Tar-EnvZ. The binding of the wild-type ligand aspartate to Tar and the intracellular transcription of ompC-RFP it initiated was thus used as a proof-of-concept of our future histamine-initiated system.
1. RU1012 with no plasmid
2. RU1012 with OmpC-RFP
3. RU1012 with Tar-EnvZ
4. RU1012 with Trg-EnvZ
5. RU1012 with OmpC-RFP + Tar-EnvZ
6. RU1012 with OmpC-RFP + Trg-EnvZ
Results: Testing the Cascade
Our fluorescence assays of these transformed colonies indicate that signal transduction is indeed effective.
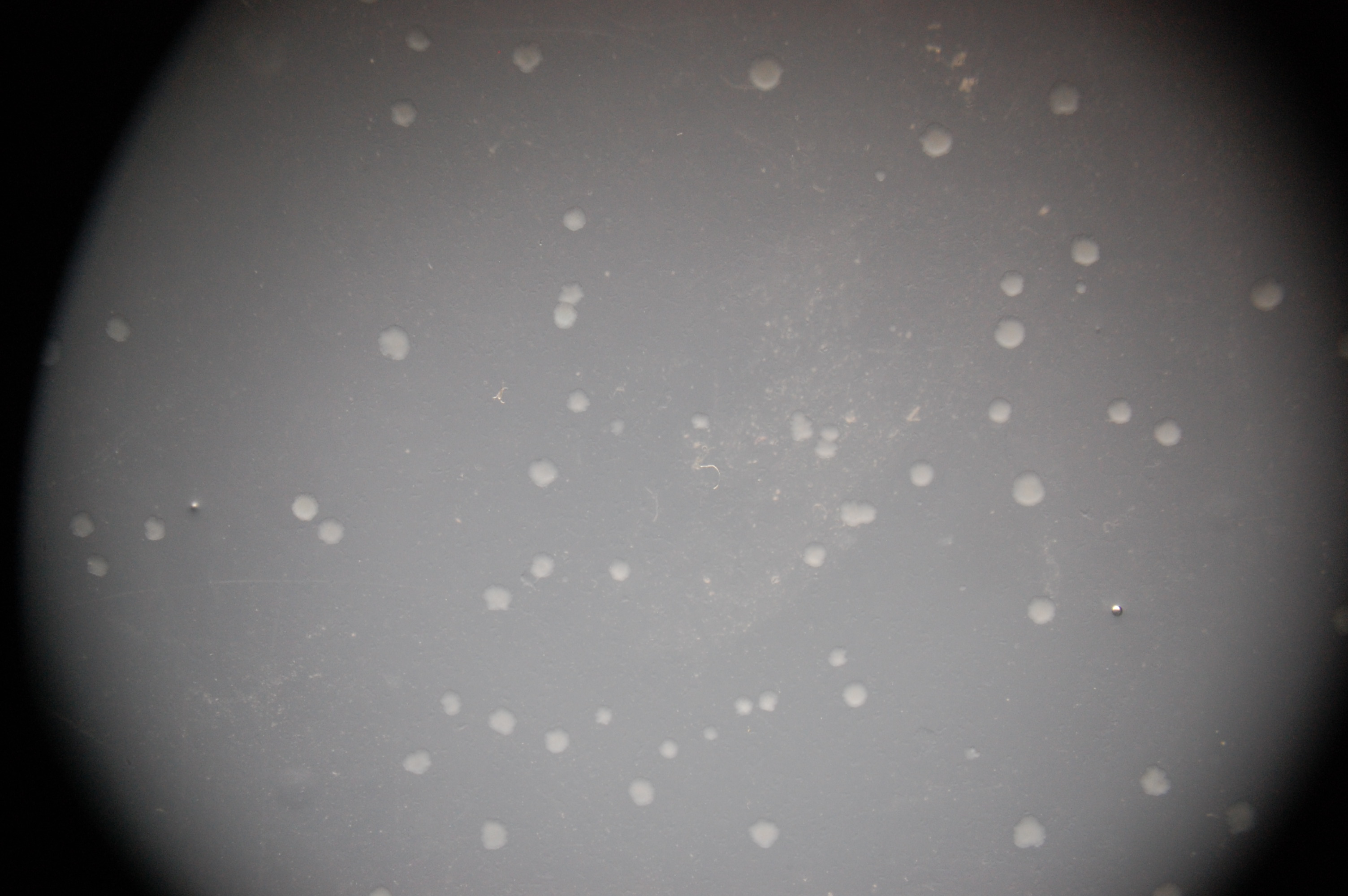
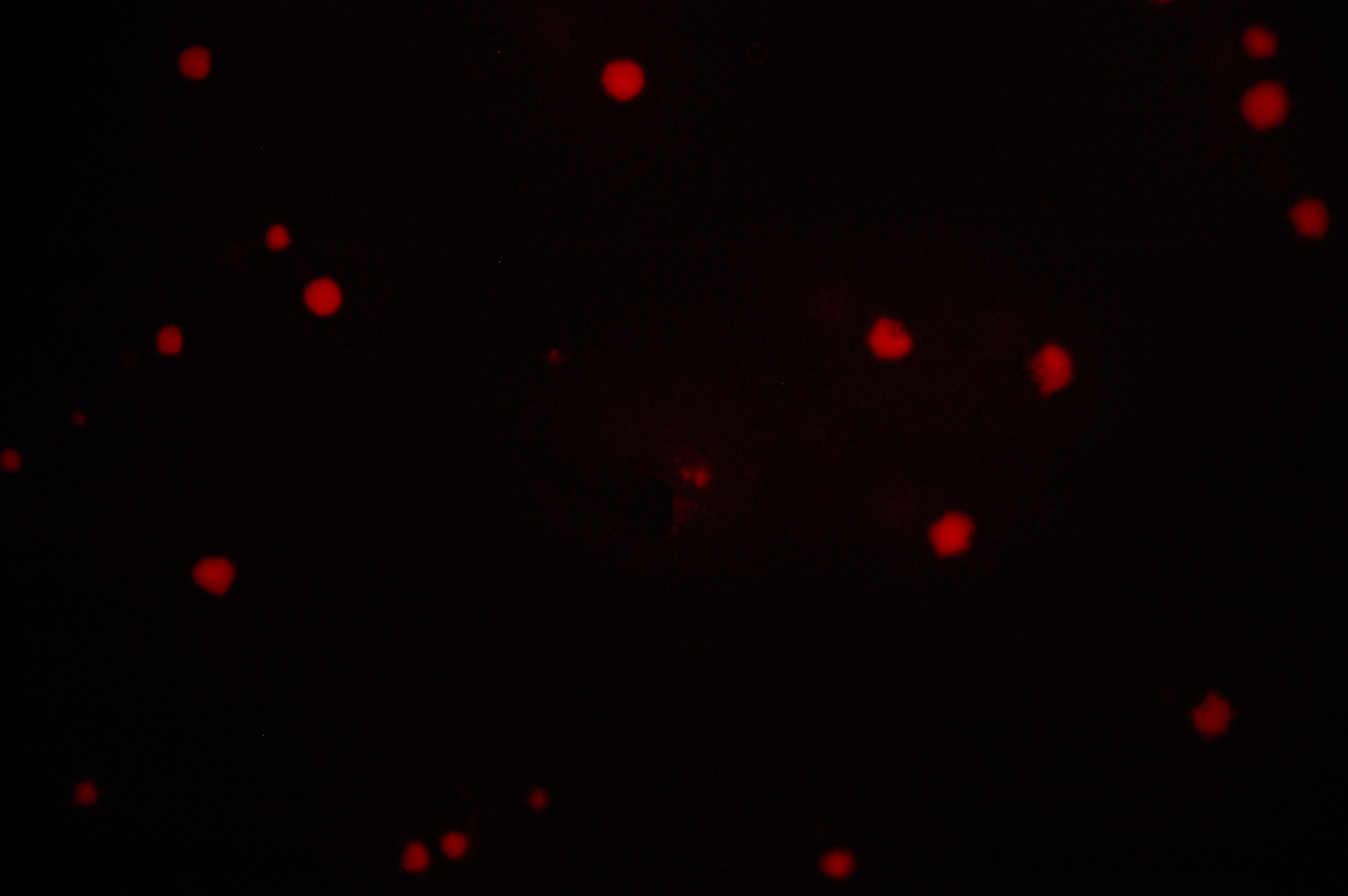
All photographs were taken on an epi-fluorescent microscope: Olympus SZX16, excitation source X-cite Series 120. The first image in each series is under bright light, the second under a fluorescent filter for RFP.
1. RU1012 with no plasmid
2. RU1012 with OmpC-RFP
3. RU1012 with Tar-EnvZ
4. RU1012 with Trg-EnvZ
5. RU1012 with OmpC-RFP + Tar-EnvZ
6. RU1012 with OmpC-RFP + Trg-EnvZ
References
- Baumgartner, James W., Changhoon Kim, Renée E. Brisette, Masoyuri Inouye, Changkyu Park, and Gerald L. Hazelbauer. "Transmembrane signalling by a hybrid protein: communication from the domain of chemoreceptor Trg that recognizes sugar-binding proteins to the kinase/phosphatase domain of osmosensor EnvZ." Journal of Bacteriology 176: 1157-163. Journal of Bacteriology. Web. 25 July 2009. <http://www.ncbi.nlm.nih.gov:80/pmc/articles/PMC205168/>.
- Looger, Loren L., Mary A. Dwyer, James J. Smith, and Homme H. Hellinga. "Computational design of receptor and sensor proteins with novel functions." Nature 423 (2003): 185-90. PubMed. Web. 15 July 2009. <http://www.ncbi.nlm.nih.gov/pubmed/12736688>.
- Meiler, Jens, and David Baker. "ROSETTALIGAND: protein-small molecule docking with full side-chain flexibility." Protein 65.3 (2006): 538-48. PubMed. Web. 12 Aug. 2009. <http://www.ncbi.nlm.nih.gov/pubmed/16972285>.
- Rhiju, Das, and David Baker. "Macromolecular modeling with rosetta." Annual Review of Biochemistry 77: 368-82. PubMed. Web. 12 July 2009. <http://www.ncbi.nlm.nih.gov/pubmed/18410248>.
- Rothlisberger, Daniela, Olga Kersonsky, Andrew M. Wollacott, Lin Jiang, Jason Dechancie, Jamie Betger, Jasmine L. Gallaher, Eric A. Althoff, Alexandre Zanghellini, Orly Dym, Shira Albeck, Kendall N. Houk, Dan S. Tawfik, and David Baker. "Kemp elimination catalysts by computational enzyme design." Nature 453 (2008): 190-95. Nature. 19 Mar. 2008. Web. 5 July 2009. <http://www.nature.com/nature/journal/v453/n7192/abs/nature06879.html>.
- Utsumi, R., R. E. Brisette, A. Rampersaud, S. A. Forst, K. Oosawa, and M. Inouye. "Activation of bacterial porin gene expression by a chimeric signal transducer in response to aspartate." Science 245.4923 (1989): 1246-249. Science. Web. 27 June 2009. <http://www.sciencemag.org/cgi/content/abstract/245/4923/1246>.
 "
"