Team:Cambridge/Notebook/Week2
From 2009.igem.org
MikeDavies (Talk | contribs) (→Weekend Work) |
MikeDavies (Talk | contribs) (→Violacin) |
||
| (17 intermediate revisions not shown) | |||
| Line 4: | Line 4: | ||
{{Template:CambridgeNotepad}} | {{Template:CambridgeNotepad}} | ||
| + | |||
| + | ''Competent cell are made - the mel and vio plasmids are transformed - two biobricks (the orange pigment and the constituative promotor) are transformed - more research is done into our pigment biosynthesis'' | ||
==Monday== | ==Monday== | ||
| - | Finalized our project into a few '''objectives''' | + | ===Objectives=== |
| + | Finalized our project into a few '''objectives'''; | ||
*Pigment production | *Pigment production | ||
:*Transform E. coli to produce pigment | :*Transform E. coli to produce pigment | ||
:*Hook up the pigmentation genes to inducible promoters | :*Hook up the pigmentation genes to inducible promoters | ||
*Shutter mechanism | *Shutter mechanism | ||
| - | :* | + | :*Build, test a shutter mechanism |
:*Hook up pigmentation genes under inducible promoters to the shutter mechanism | :*Hook up pigmentation genes under inducible promoters to the shutter mechanism | ||
*Improve on previous iGEM projects | *Improve on previous iGEM projects | ||
| - | :* | + | :*Arsenic sensor? |
| - | :* | + | :*Bacterial photography? |
*Modelling | *Modelling | ||
| - | :* | + | :*Shutter mechanism |
:*Growth rate regulation | :*Growth rate regulation | ||
| - | + | ===Goals for Wet Work=== | |
*Inducible pigment production integrated with a shutter mechanism | *Inducible pigment production integrated with a shutter mechanism | ||
*Possibly attach the arsenic bio-sensor to our green/red pigment output | *Possibly attach the arsenic bio-sensor to our green/red pigment output | ||
| Line 27: | Line 30: | ||
==Tuesday== | ==Tuesday== | ||
| - | + | ===Procedure to make competent cells started=== | |
*One colony of TOP10 ''E. coli'' added to 50ml of LB broth. Incubated overnight in the shaking incubator | *One colony of TOP10 ''E. coli'' added to 50ml of LB broth. Incubated overnight in the shaking incubator | ||
| - | + | ===Planned and prepared for transformation=== | |
| - | *Media made up | + | *Media made up: |
:*Two litres of 0.1 mM HEPES | :*Two litres of 0.1 mM HEPES | ||
| - | :*One litre of LB broth (for making | + | :*One litre of LB broth (for making competent cells) |
:*100ml of 10% glycerol | :*100ml of 10% glycerol | ||
*Ampicillin made up with water to make 8 x 1ml aliquots of 100mg/ml | *Ampicillin made up with water to make 8 x 1ml aliquots of 100mg/ml | ||
*15 LB agar and 100ug/ml Ampicillin plates made up | *15 LB agar and 100ug/ml Ampicillin plates made up | ||
| - | + | ===Dry work=== | |
*Discussed how genetic amplifiers work, including transient and steady state response | *Discussed how genetic amplifiers work, including transient and steady state response | ||
*Looked at ways in which a latch might be made, by positive feedback, mutual inhibition, or other schemes | *Looked at ways in which a latch might be made, by positive feedback, mutual inhibition, or other schemes | ||
| Line 45: | Line 48: | ||
| - | + | ===Plan for tomorrow=== | |
| - | * | + | *Prepare trimethoprim stock solution |
| - | * | + | *Make LB agar and trimethoprim plates as well as LB agar and copper + tyrosine plates |
| - | *Complete | + | *Complete procedure for making competent cells (transfer bacteria into big flask, wash with HEPES, and aliquot into glycerol) |
*Attempt transformations by electroporation | *Attempt transformations by electroporation | ||
:*pPSX plasmid (violet) | :*pPSX plasmid (violet) | ||
| Line 56: | Line 59: | ||
==Wednesday== | ==Wednesday== | ||
| - | + | ===Competent Cell Procedure=== | |
| - | *Completed (see protocol page for details of the procedure). Competent cells stored in aliquots at -80 degrees. | + | *Completed [https://2009.igem.org/Team:Cambridge/Protocols (see protocol page for details of the procedure)]. Competent cells stored in aliquots at -80 degrees. |
| - | + | ===Planned and Prepared for Electroporation Tomorrow=== | |
| - | *Made up 4 LB agar and 100ug/ml Ampicillin, 0. | + | *Made up 4 LB agar and 100ug/ml Ampicillin, 0.2mg/mL tyrosine (NOTE: protocol actually calls for 0.3mg/mL), 15 ug/mL CuSO4 plates |
*Plan to transform TOP10 competent bacteria with | *Plan to transform TOP10 competent bacteria with | ||
:*Melanin plasmid, pTRCmelA plasmid -- Plate onto tyrosine/copper/ampicillin plate | :*Melanin plasmid, pTRCmelA plasmid -- Plate onto tyrosine/copper/ampicillin plate | ||
| Line 73: | Line 76: | ||
==Thursday== | ==Thursday== | ||
| - | + | ===Electroporation=== | |
| - | * | + | *Transformed Top10 competent cells, incubated the following plates: |
:*Top10 + pTRCmelA plasmid on tyrosine/copper/ampicillin plate | :*Top10 + pTRCmelA plasmid on tyrosine/copper/ampicillin plate | ||
:*Top10 + Plasmid pPSX-Vio+ on trimethoprim plate | :*Top10 + Plasmid pPSX-Vio+ on trimethoprim plate | ||
| Line 82: | Line 85: | ||
*All plates were stored overnight in the 37 degree incubator | *All plates were stored overnight in the 37 degree incubator | ||
| - | + | ===Making BioBricks=== | |
With the aim of making a brown-pigmented BioBrick, we looked at constructing primers to cut out the MelA gene from the plasmid, add the prefix and suffix, and remove the two forbidden restriction sites. This would then be pasted into one of the recommended vectors and could be submitted to the registry. | With the aim of making a brown-pigmented BioBrick, we looked at constructing primers to cut out the MelA gene from the plasmid, add the prefix and suffix, and remove the two forbidden restriction sites. This would then be pasted into one of the recommended vectors and could be submitted to the registry. | ||
| Line 88: | Line 91: | ||
[[Image:Mela_biobrick.JPG]] | [[Image:Mela_biobrick.JPG]] | ||
| - | + | ===Input and Processing systems=== | |
We split into groups to plan and discuss the following possible ideas for input systems: | We split into groups to plan and discuss the following possible ideas for input systems: | ||
| Line 94: | Line 97: | ||
*Siming and Alan = Bistable switch | *Siming and Alan = Bistable switch | ||
*Crispian and Megan = Population control and growth | *Crispian and Megan = Population control and growth | ||
| - | *Viv = Coloured spores ( | + | *Viv = Coloured spores (Idea discarded due to high levels of complexity involved in understanding, designing and working with the spore system. It also required the use of a chassis that was not E-Coli.) |
| - | + | ===Plans for Friday=== | |
Meeting at 10am to discuss ideas | Meeting at 10am to discuss ideas | ||
| Line 104: | Line 107: | ||
==Friday== | ==Friday== | ||
| - | + | ===Results of Wet Work=== | |
| - | + | ||
All transformations were completed sucessfully. Single cononies were observed on the 1/10 dilution plates and could easily be picked out for plasmid propagation procedure (to be started on Monday). However no colours were observed. | All transformations were completed sucessfully. Single cononies were observed on the 1/10 dilution plates and could easily be picked out for plasmid propagation procedure (to be started on Monday). However no colours were observed. | ||
| - | Reasons: | + | '''Reasons:''' |
*Melanin plasmid = plasmid required the addition of IPTG | *Melanin plasmid = plasmid required the addition of IPTG | ||
*pPSX plasmid (violet) = the pigment is on a low copy number plasmid | *pPSX plasmid (violet) = the pigment is on a low copy number plasmid | ||
| - | The melanin plates were left out on the bench to see if the brown pigment developed over time. The rest of the plates were stored in the fridge for making plasmid stocks ( | + | The melanin plates were left out on the bench to see if the brown pigment developed over time. The rest of the plates were stored in the fridge for making plasmid stocks (procedure starting next week). |
| - | + | ===BioBrick Construction=== | |
Used ApE to examine the melanin plasmid: | Used ApE to examine the melanin plasmid: | ||
| Line 129: | Line 131: | ||
The other possible pigments are the orange biobrick system and the pseudomonas red/green pigment. | The other possible pigments are the orange biobrick system and the pseudomonas red/green pigment. | ||
| - | |||
| - | + | ||
| - | + | ===Pigment Plans=== | |
| - | + | ||
| + | ====Melanin==== | ||
| + | *Chemistry of Pigment Production: | ||
| + | :*The MelA gene is a tyrosinase. Tyrosinases catalyze two reactions, as described in the figure below. Melanin is a macromolecular compound produced by the polymerization of the quinone product of the second reaction, and has a characteristic brown colour. | ||
[[Image:tyrosinase action.jpg]] | [[Image:tyrosinase action.jpg]] | ||
| Line 139: | Line 143: | ||
From H. Claus and H. Decker, Bacterial tyrosinases, Syst. Appl. Microbiol. 29 (2006), pp. 3–14. [http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B7GVX-4H21K91-1&_user=1495569&_coverDate=01%2F24%2F2006&_fmt=full&_orig=search&_cdi=20442&view=c&_acct=C000053194&_version=1&_urlVersion=0&_userid=1495569&md5=de451c5e1d1d18ad12b7e39b10b80408&ref=full] | From H. Claus and H. Decker, Bacterial tyrosinases, Syst. Appl. Microbiol. 29 (2006), pp. 3–14. [http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B7GVX-4H21K91-1&_user=1495569&_coverDate=01%2F24%2F2006&_fmt=full&_orig=search&_cdi=20442&view=c&_acct=C000053194&_version=1&_urlVersion=0&_userid=1495569&md5=de451c5e1d1d18ad12b7e39b10b80408&ref=full] | ||
| - | + | ====Forward plans:==== | |
| - | + | *Troubleshoot pigment production | |
| - | + | *Make our own plasmid stocks | |
| - | + | *As above, make the pigment into a biobrick | |
| - | + | *Characterize the pigment by making an absorbance spectrum (max absorption at 400 nm); incidentally, the melanin gene we have is actually a mutant, and maximum pigment production was achieved in a strain that over-produces tyrosine, as described in: | |
Santos, C. N., and G. Stephanopoulos. 2008. Melanin-based high-throughput screen for L-tyrosine production in Escherichia coli. Appl. Environ. Microbiol. 74:1190-1197. [http://aem.asm.org/cgi/content/abstract/74/4/1190] | Santos, C. N., and G. Stephanopoulos. 2008. Melanin-based high-throughput screen for L-tyrosine production in Escherichia coli. Appl. Environ. Microbiol. 74:1190-1197. [http://aem.asm.org/cgi/content/abstract/74/4/1190] | ||
| - | + | :*Goal: To use as a single BioBrick output (possibly as a 'fall-back' pigment to test processing systems) | |
| - | + | ||
| - | + | ||
| - | + | ====Violacein==== | |
| + | *Chemistry of pigment production: | ||
| + | :*There are 4 genes that convert tryptophan into violacein; the biosynthetic pathway is detailed below | ||
[[Image:violacein pigment production.jpg]] | [[Image:violacein pigment production.jpg]] | ||
| Line 156: | Line 162: | ||
From P.R. August, T.H. Grossman, C. Minor, M.P. Draper, I.A. MacNeil, J.M. Pemberton, K.M. Call, d. Holt, and M. S. Osbourne, Sequence Analysis and Functional Characterization of the Violacein Biosynthetic Pathway from ''Chromobacterium violaceum'', J. Mol. Microbiol. Biotechnol. (2000) 2(4): 513-519. [http://www.horizonpress.com/jmmb/v2/v2n4/26.pdf] | From P.R. August, T.H. Grossman, C. Minor, M.P. Draper, I.A. MacNeil, J.M. Pemberton, K.M. Call, d. Holt, and M. S. Osbourne, Sequence Analysis and Functional Characterization of the Violacein Biosynthetic Pathway from ''Chromobacterium violaceum'', J. Mol. Microbiol. Biotechnol. (2000) 2(4): 513-519. [http://www.horizonpress.com/jmmb/v2/v2n4/26.pdf] | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | + | *Forward plans: | |
| - | [[Image: | + | :*troubleshoot pigment production |
| + | :*make our own plasmid sticks | ||
| + | :*As above, synthesize the operon with the biobrick prefix and suffix | ||
| + | :*Use the synthesized operon as a template to PCR out the individual genes | ||
| + | :*Characterize both violacein and the cyan molecule by making an absorption spectrum for each. | ||
| + | :*Explore the possibilities of constructing logic gates | ||
| + | [[Image:diagram2.jpg]] | ||
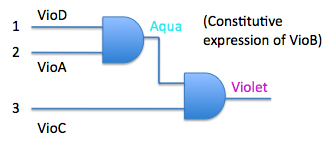
| + | For example, | ||
| + | [[Image:logic gate2.jpg]] | ||
| - | + | ::*Need to analyze the flux through the pathway | |
| - | + | ::*The products of the reactions catalyzed by VioD and VioA are combined in a 1:1 ratio by VioB; what happens to the production of cyan and violacein if flux through VioD is much greater than flux through VioA? Which might be the case if VioD and VioA are induced by different inputs that are not reliably equal. | |
| - | + | ::*What are the dynamics of the flux through VioC? Would analyze by producing cyan, and then switching on VioC and measuring absorbance. Considerations for biosensors - if presence of whatever the biosensor is sensing causes cyan production, and then low rate of VioC expression slowly turns cyan into violet, violet being an indication that the sensor is old and past its shelf life . | |
| - | + | ====Red and Green from Pseudomonas==== | |
| - | + | *Chemistry of the pigment | |
[[Image:pseudomonas.jpg]] | [[Image:pseudomonas.jpg]] | ||
From Mavrodi, D. V., R. F. Bonsall, S. M. Delaney, M. J. Soule, G. Phillips, and L. S. Thomashow. 2001. Functional analysis of genes for biosynthesis of pyocyanin and phenazine-1-carboxamide from Pseudomonas aeruginosa PAO1. J. Bacteriol. 183:6454-6465. [http://jb.asm.org/cgi/content/abstract/183/21/6454?ijkey=41531d004be13e87d3ba32abcbbd672bcc9d6cf3&keytype2=tf_ipsecsha] | From Mavrodi, D. V., R. F. Bonsall, S. M. Delaney, M. J. Soule, G. Phillips, and L. S. Thomashow. 2001. Functional analysis of genes for biosynthesis of pyocyanin and phenazine-1-carboxamide from Pseudomonas aeruginosa PAO1. J. Bacteriol. 183:6454-6465. [http://jb.asm.org/cgi/content/abstract/183/21/6454?ijkey=41531d004be13e87d3ba32abcbbd672bcc9d6cf3&keytype2=tf_ipsecsha] | ||
::*Pyocyanin is a green pigment that diffuses out of the cell, the red pigment also diffuses out of the cell. | ::*Pyocyanin is a green pigment that diffuses out of the cell, the red pigment also diffuses out of the cell. | ||
| - | :* | + | :*Currently waiting for the arrival of plasmids containing PhzM and PhzS; if they do arrive, hoping to explore logic gate construction. |
| - | |||
| - | + | ===Planning a Processing System=== | |
| - | + | ====Amplification:==== | |
| + | An amplifier consists of a ribosome binding site, a protein coding region, a terminator and a promoter. The protein produced in the device binds to the promoter as an activator, such that an incoming gene transcription will be amplified to various levels with appropriate choices of activator and promoter. The Cambridge team in 2007 looked at these devices. | ||
| - | + | We are taking this idea forward. | |
| + | ====Population Control:==== | ||
| + | If we had three different pigments in a single plate, we would like to make sure that the colonies all grow to the same size. | ||
| + | |||
| + | We therefore looked into an AHL-regulated arrangement, where the slowest-growing strain would produce signalling molecules, allowing the faster strains to grow. The medium speed strain could also have a signalling molecule for induction of the fast strain. | ||
| + | |||
| + | The idea has a good chance of working in a liquid medium, where the relevant signalling molecules can easily disperse. However, on a plate, spatial effects have to be considered. Due to this difficulty, we chose not to continue this idea further. | ||
| + | |||
| + | ====Bioswitch:==== | ||
A basic bistable switch requires two inverters, and some way of making it change state. | A basic bistable switch requires two inverters, and some way of making it change state. | ||
| Line 213: | Line 227: | ||
However, we decided that given the large amount of work which we need to do on pigments, and the fact that a non-functioning bistable would not be novel, we would not progress further with this idea. | However, we decided that given the large amount of work which we need to do on pigments, and the fact that a non-functioning bistable would not be novel, we would not progress further with this idea. | ||
| - | ==Weekend | + | ==Weekend== |
| - | + | ===RED=== | |
| - | + | ||
Looked at red pigment synthesis briefly as optional easy extra | Looked at red pigment synthesis briefly as optional easy extra | ||
link to relevant paper - https://static.igem.org/mediawiki/2009/f/ff/1128.pdf | link to relevant paper - https://static.igem.org/mediawiki/2009/f/ff/1128.pdf | ||
| - | + | ===Degredation tags=== | |
Interesting articles: | Interesting articles: | ||
| - | + | *Engineering controllable protein degredation [http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6WSR-4K47X21-G&_user=1495569&_rdoc=1&_fmt=&_orig=search&_sort=d&_docanchor=&view=c&_searchStrId=967086878&_rerunOrigin=google&_acct=C000053194&_version=1&_urlVersion=0&_userid=1495569&md5=8603cc30cea1e63c39b7a605a8f2a477] | |
| - | Engineering controllable protein degredation | + | |
Primers for melanin designed! | Primers for melanin designed! | ||
| - | <!--Do not remove the first and last lines in this page!--> | + | <!--Do not remove the first and last lines in this page!-->{{Template:CambridgeBottom}} |
Latest revision as of 16:28, 7 October 2009
Categories :
Project :
-
Overview
Sensitivity Tuner
--- Characterisation
--- Modelling
Colour Generators
--- Carotenoids (Orange/Red)
--- Melanin (Brown)
--- Violacein (Purple/Green)
The Future
Safety
Notebook :
Team Logistics :
Week 2 - The Project
Competent cell are made - the mel and vio plasmids are transformed - two biobricks (the orange pigment and the constituative promotor) are transformed - more research is done into our pigment biosynthesis
Monday
Objectives
Finalized our project into a few objectives;
- Pigment production
- Transform E. coli to produce pigment
- Hook up the pigmentation genes to inducible promoters
- Shutter mechanism
- Build, test a shutter mechanism
- Hook up pigmentation genes under inducible promoters to the shutter mechanism
- Improve on previous iGEM projects
- Arsenic sensor?
- Bacterial photography?
- Modelling
- Shutter mechanism
- Growth rate regulation
Goals for Wet Work
- Inducible pigment production integrated with a shutter mechanism
- Possibly attach the arsenic bio-sensor to our green/red pigment output
Tuesday
Procedure to make competent cells started
- One colony of TOP10 E. coli added to 50ml of LB broth. Incubated overnight in the shaking incubator
Planned and prepared for transformation
- Media made up:
- Two litres of 0.1 mM HEPES
- One litre of LB broth (for making competent cells)
- 100ml of 10% glycerol
- Ampicillin made up with water to make 8 x 1ml aliquots of 100mg/ml
- 15 LB agar and 100ug/ml Ampicillin plates made up
Dry work
- Discussed how genetic amplifiers work, including transient and steady state response
- Looked at ways in which a latch might be made, by positive feedback, mutual inhibition, or other schemes
- Positive feedback relies in part on an amplifier component: this would need to be fixed on toxicity grounds before the secondary application could be used.
Plan for tomorrow
- Prepare trimethoprim stock solution
- Make LB agar and trimethoprim plates as well as LB agar and copper + tyrosine plates
- Complete procedure for making competent cells (transfer bacteria into big flask, wash with HEPES, and aliquot into glycerol)
- Attempt transformations by electroporation
- pPSX plasmid (violet)
- melA plasmid (brown)
- biobricks
Wednesday
Competent Cell Procedure
- Completed (see protocol page for details of the procedure). Competent cells stored in aliquots at -80 degrees.
Planned and Prepared for Electroporation Tomorrow
- Made up 4 LB agar and 100ug/ml Ampicillin, 0.2mg/mL tyrosine (NOTE: protocol actually calls for 0.3mg/mL), 15 ug/mL CuSO4 plates
- Plan to transform TOP10 competent bacteria with
- Melanin plasmid, pTRCmelA plasmid -- Plate onto tyrosine/copper/ampicillin plate
- Violacein plasmid,Plasmid pPSX-Vio+ -- Plate onto trimethoprim plate
- Orange biobrick, BBa_K152005 -- Plate onto ampicillin plate
- Promoter, R0011 -- Plate onto ampicillin plate
- More?
- Need to make trimethoprim stock solution 10mg/mL acetone, and plate at 50ug/mL
- Tried to make the stock solution today, couldn't get the solid trimethoprim to dissolve in acetone
- Need to work out exact volumes of plasmid to be added to the cell (concentrations written on the side of the bottles)
Thursday
Electroporation
- Transformed Top10 competent cells, incubated the following plates:
- Top10 + pTRCmelA plasmid on tyrosine/copper/ampicillin plate
- Top10 + Plasmid pPSX-Vio+ on trimethoprim plate
- Top10 + BBa_K152005 on ampicillin plate
- Top 10 + R0011 on ampicillin plate
- 1:10 dilutions of each were also plated
- All plates were stored overnight in the 37 degree incubator
Making BioBricks
With the aim of making a brown-pigmented BioBrick, we looked at constructing primers to cut out the MelA gene from the plasmid, add the prefix and suffix, and remove the two forbidden restriction sites. This would then be pasted into one of the recommended vectors and could be submitted to the registry.
Input and Processing systems
We split into groups to plan and discuss the following possible ideas for input systems:
- Mike and Shuna = Amplifier
- Siming and Alan = Bistable switch
- Crispian and Megan = Population control and growth
- Viv = Coloured spores (Idea discarded due to high levels of complexity involved in understanding, designing and working with the spore system. It also required the use of a chassis that was not E-Coli.)
Plans for Friday
Meeting at 10am to discuss ideas Meeting with faculty at 2.30pm to further discuss ideas and present work done so far
Wet work = colony PCR (to be explained by James) and observation of (hopefully!) coloured plates
Friday
Results of Wet Work
All transformations were completed sucessfully. Single cononies were observed on the 1/10 dilution plates and could easily be picked out for plasmid propagation procedure (to be started on Monday). However no colours were observed.
Reasons:
- Melanin plasmid = plasmid required the addition of IPTG
- pPSX plasmid (violet) = the pigment is on a low copy number plasmid
The melanin plates were left out on the bench to see if the brown pigment developed over time. The rest of the plates were stored in the fridge for making plasmid stocks (procedure starting next week).
BioBrick Construction
Used ApE to examine the melanin plasmid:
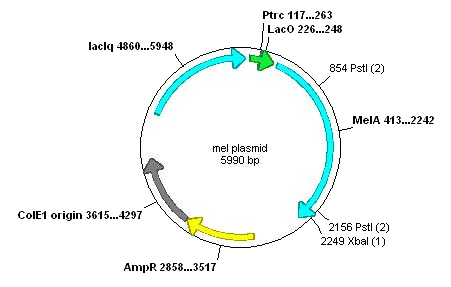
This contains only two forbidden restriction sites within the actual gene. These could be removed by PCR.
Also looked at the vio operon:
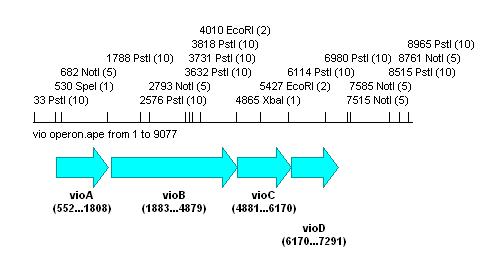
As this contains a large number of the forbidden biobrick restriction sites, it would be impossible to prepare the biobrick using PCR alone. The operon would have to be synthesised with these sites removed to be feasible as a biobrick.
The other possible pigments are the orange biobrick system and the pseudomonas red/green pigment.
Pigment Plans
Melanin
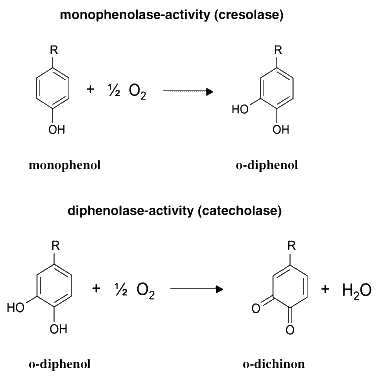
- Chemistry of Pigment Production:
- The MelA gene is a tyrosinase. Tyrosinases catalyze two reactions, as described in the figure below. Melanin is a macromolecular compound produced by the polymerization of the quinone product of the second reaction, and has a characteristic brown colour.
From H. Claus and H. Decker, Bacterial tyrosinases, Syst. Appl. Microbiol. 29 (2006), pp. 3–14. [http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B7GVX-4H21K91-1&_user=1495569&_coverDate=01%2F24%2F2006&_fmt=full&_orig=search&_cdi=20442&view=c&_acct=C000053194&_version=1&_urlVersion=0&_userid=1495569&md5=de451c5e1d1d18ad12b7e39b10b80408&ref=full]
Forward plans:
- Troubleshoot pigment production
- Make our own plasmid stocks
- As above, make the pigment into a biobrick
- Characterize the pigment by making an absorbance spectrum (max absorption at 400 nm); incidentally, the melanin gene we have is actually a mutant, and maximum pigment production was achieved in a strain that over-produces tyrosine, as described in:
Santos, C. N., and G. Stephanopoulos. 2008. Melanin-based high-throughput screen for L-tyrosine production in Escherichia coli. Appl. Environ. Microbiol. 74:1190-1197. [http://aem.asm.org/cgi/content/abstract/74/4/1190]
- Goal: To use as a single BioBrick output (possibly as a 'fall-back' pigment to test processing systems)
Violacein
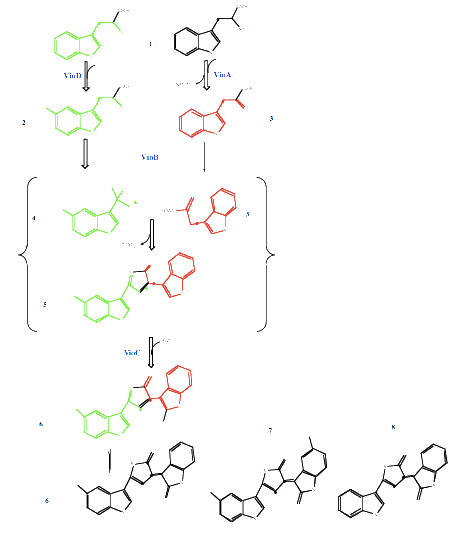
- Chemistry of pigment production:
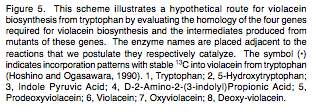
- There are 4 genes that convert tryptophan into violacein; the biosynthetic pathway is detailed below
Molecule 5 is Cyan coloured, allowing two different colour states to be produced by this system.
From P.R. August, T.H. Grossman, C. Minor, M.P. Draper, I.A. MacNeil, J.M. Pemberton, K.M. Call, d. Holt, and M. S. Osbourne, Sequence Analysis and Functional Characterization of the Violacein Biosynthetic Pathway from Chromobacterium violaceum, J. Mol. Microbiol. Biotechnol. (2000) 2(4): 513-519. [http://www.horizonpress.com/jmmb/v2/v2n4/26.pdf]
- Forward plans:
- troubleshoot pigment production
- make our own plasmid sticks
- As above, synthesize the operon with the biobrick prefix and suffix
- Use the synthesized operon as a template to PCR out the individual genes
- Characterize both violacein and the cyan molecule by making an absorption spectrum for each.
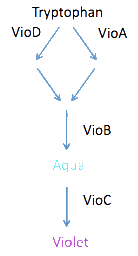
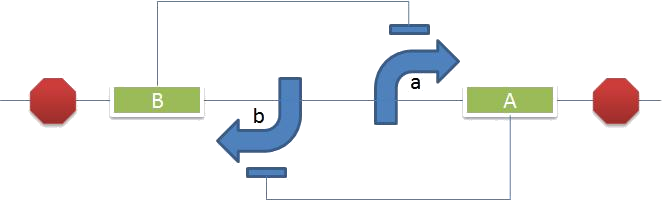
- Explore the possibilities of constructing logic gates
- Need to analyze the flux through the pathway
- The products of the reactions catalyzed by VioD and VioA are combined in a 1:1 ratio by VioB; what happens to the production of cyan and violacein if flux through VioD is much greater than flux through VioA? Which might be the case if VioD and VioA are induced by different inputs that are not reliably equal.
- What are the dynamics of the flux through VioC? Would analyze by producing cyan, and then switching on VioC and measuring absorbance. Considerations for biosensors - if presence of whatever the biosensor is sensing causes cyan production, and then low rate of VioC expression slowly turns cyan into violet, violet being an indication that the sensor is old and past its shelf life .
Red and Green from Pseudomonas
- Chemistry of the pigment
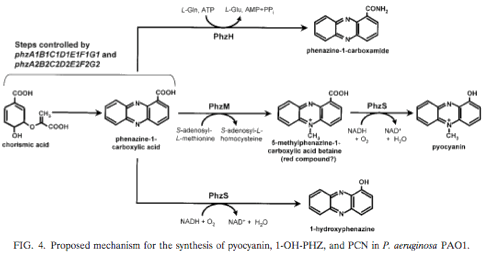
From Mavrodi, D. V., R. F. Bonsall, S. M. Delaney, M. J. Soule, G. Phillips, and L. S. Thomashow. 2001. Functional analysis of genes for biosynthesis of pyocyanin and phenazine-1-carboxamide from Pseudomonas aeruginosa PAO1. J. Bacteriol. 183:6454-6465. [http://jb.asm.org/cgi/content/abstract/183/21/6454?ijkey=41531d004be13e87d3ba32abcbbd672bcc9d6cf3&keytype2=tf_ipsecsha]
- Pyocyanin is a green pigment that diffuses out of the cell, the red pigment also diffuses out of the cell.
- Currently waiting for the arrival of plasmids containing PhzM and PhzS; if they do arrive, hoping to explore logic gate construction.
Planning a Processing System
Amplification:
An amplifier consists of a ribosome binding site, a protein coding region, a terminator and a promoter. The protein produced in the device binds to the promoter as an activator, such that an incoming gene transcription will be amplified to various levels with appropriate choices of activator and promoter. The Cambridge team in 2007 looked at these devices.
We are taking this idea forward.
Population Control:
If we had three different pigments in a single plate, we would like to make sure that the colonies all grow to the same size.
We therefore looked into an AHL-regulated arrangement, where the slowest-growing strain would produce signalling molecules, allowing the faster strains to grow. The medium speed strain could also have a signalling molecule for induction of the fast strain.
The idea has a good chance of working in a liquid medium, where the relevant signalling molecules can easily disperse. However, on a plate, spatial effects have to be considered. Due to this difficulty, we chose not to continue this idea further.
Bioswitch:
A basic bistable switch requires two inverters, and some way of making it change state.
Important papers on this concept in synthetic biology include:
"Construction of a genetic toggle switch in Escherichia coli" Gardner, Cantor & Collins 2000 http://www.nature.com/nature/journal/v403/n6767/pdf/403339a0.pdf
This paper was followed up with a second, "Programmable cells: Interfacing natural and engineered gene networks", 2004 http://www.pnas.org/content/101/22/8414.figures-only
The authors showed in 2000 that a genetic toggle switch was possible. They investigated using different promoters, and found only some combinations to give bistability.
Many iGEM teams have attempted to recreate a bistable switch; none have so far fully succeeded! The most recent were Peking 2007 and Bologna 2008.
Peking 2007 designed a bistable circuit as part of their "push on, push off" device. They connected together two mutual inhibitors, and showed that this gave an agar spread that had red in places and green in others. This suggests that they made a bistable device. However, they did not progress to activation by external sources.
Bologna's iGEM project in 2008 was a bistable circuit, called "E.coli PROM". They performed extensive modelling of the device, but did not complete fabrication.
This leaves an iGEM team with a good opportunity. Building on Peking and Bologna, amongst others, a working, and inducible bistable could be made, an iGEM first.
However, we decided that given the large amount of work which we need to do on pigments, and the fact that a non-functioning bistable would not be novel, we would not progress further with this idea.
Weekend
RED
Looked at red pigment synthesis briefly as optional easy extra link to relevant paper - https://static.igem.org/mediawiki/2009/f/ff/1128.pdf
Degredation tags
Interesting articles:
- Engineering controllable protein degredation [http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6WSR-4K47X21-G&_user=1495569&_rdoc=1&_fmt=&_orig=search&_sort=d&_docanchor=&view=c&_searchStrId=967086878&_rerunOrigin=google&_acct=C000053194&_version=1&_urlVersion=0&_userid=1495569&md5=8603cc30cea1e63c39b7a605a8f2a477]
Primers for melanin designed!
 "
"