Team:Groningen/Notebook/4 September 2009
From 2009.igem.org
(→GVP Cluster) |
(→Wet) |
||
| (18 intermediate revisions not shown) | |||
| Line 7: | Line 7: | ||
[http://partsregistry.org/Part:BBa_F2620 BBa_F2620] | [http://partsregistry.org/Part:BBa_F2620 BBa_F2620] | ||
| + | :→ {{done}} Isolate plasmid from o.n. precultures of pLacI-GVP and pBad-araC-GVP in pSB1A2 | ||
| + | :→ {{done}} Restriction control of plasmids and purification of wanted fragments for ligation in pSB2K3 vector | ||
| + | |||
| + | :→ {{done}} Ligate pArsR-GVP, pZntR-GVP, pCueO-GVP, and pLacI-GVP into vector pSB2A3 | ||
| + | :→ {{done}} Transform E.coli Top10 cells with ligation products and pSB2K3 (7C and 7K) | ||
| + | |||
| + | :→ {{done}} Order synthetic DNA for GVP | ||
| + | :→ {{todo}} Order primer for PstI site removal | ||
| + | |||
| + | :→ {{done}} Test promoter strenght compared to BBa_J23101 promoter (Sven) | ||
| + | :→ {{todo}} Enter sequences of constructs to Sandbox | ||
'''O.n. precultures''' | '''O.n. precultures''' | ||
| Line 61: | Line 72: | ||
|Yes (EcoRI/PstI) | |Yes (EcoRI/PstI) | ||
|} | |} | ||
| + | |||
| + | |||
| + | '''Restriction for Assembly''' | ||
| + | |||
| + | The vector [http://partsregistry.org/wiki/index.php?title=Part:pSB1A2 pSB1A2] containing the pLacI and pBad-araC with [http://partsregistry.org/Part:BBa_I750016 GVP] composite parts were cut with PstI and EcoRI to create correct ends for insert into [http://partsregistry.org/Part:pSB2K3 pSB2K3], which was also cut with EcoRI and PstI (4x) on 3-9. | ||
| + | |||
| + | {|cellpadding="2" cellspacing="1" border="4" | ||
| + | |'''Plasmid''' | ||
| + | |'''Amount μL''' | ||
| + | |'''MQ μL''' | ||
| + | |'''Fast digest buffer''' | ||
| + | |'''EcoRI fast digest enzyme''' | ||
| + | |'''XbaI fast digest enzyme''' | ||
| + | |'''SpeI fast digest enzyme''' | ||
| + | |'''PstI fast digest enzyme''' | ||
| + | |- | ||
| + | |pSB1A2 pLacI-GVP no.1 | ||
| + | |3.0 | ||
| + | |13.0 | ||
| + | |3.0 | ||
| + | |1.0 | ||
| + | |x | ||
| + | |x | ||
| + | |1.0 | ||
| + | |- | ||
| + | |pSB1A2 pLacI-GVP no.2 | ||
| + | |16.0 | ||
| + | |x | ||
| + | |3.0 | ||
| + | |1.0 | ||
| + | |x | ||
| + | |x | ||
| + | |1.0 | ||
| + | |- | ||
| + | |pSB1A2 pBad/araC-GVP no.1 | ||
| + | |16.0 | ||
| + | |x | ||
| + | |3.0 | ||
| + | |1.0 | ||
| + | |x | ||
| + | |x | ||
| + | |1.0 | ||
| + | |- | ||
| + | |pSB1A2 pBad/araC-GVP no.2 | ||
| + | |16.0 | ||
| + | |x | ||
| + | |3.0 | ||
| + | |1.0 | ||
| + | |x | ||
| + | |x | ||
| + | |1.0 | ||
| + | |} | ||
| + | |||
| + | Restriction was kept at 37C for 40 min. and put on ice until used for gel purification. | ||
| + | |||
| + | |||
| + | [[Image:Zymoclean Gel DNA Recovery Kit (D4001) 2.jpg|thumb|300px| www.zymoresearch.com]] | ||
| + | |||
| + | |||
| + | '''Gel Purification''' | ||
| + | |||
| + | * In step 7 an amount of 10μL MQ was added to elute the DNA fragments. | ||
| + | |||
| + | |||
| + | [[Image:4-9 no.1.jpg|270px]] [[Image:Generulers_1kb_marker_Fermentas.jpg]] | ||
| + | |||
| + | :→ Only the first restriction of pLacI-GVP seems to have worked judging from the fragments. | ||
| + | |||
| + | '''Ligation''' | ||
| + | |||
| + | A total amount of vector of 100ng was used (GVP) in a 1:3 ratio with insert. | ||
| + | |||
| + | {|cellpadding="2" cellspacing="1" border="4" | ||
| + | |'''Ligase buffer (uL)''' | ||
| + | |'''T4 Ligase (uL)''' | ||
| + | |'''Plasmid''' | ||
| + | |'''Insert''' | ||
| + | |'''MQ''' | ||
| + | |- | ||
| + | |3.0 | ||
| + | |1.0 | ||
| + | |5.0 pSB2K3 (no.2)(EcoRI,PstI) | ||
| + | |3.0 pLacI-GVP (EcoRI,PstI) | ||
| + | |8.0 | ||
| + | |- | ||
| + | |3.0 | ||
| + | |1.0 | ||
| + | |5.0 pSB2K3 (no.2)(EcoRI,PstI) | ||
| + | |8.0 pArsR-GVP (EcoRI,PstI) | ||
| + | |3.0 | ||
| + | |- | ||
| + | |3.0 | ||
| + | |1.0 | ||
| + | |6.0 pSB2K3 (no.3)(EcoRI,PstI) | ||
| + | |8.0 pZntR-GVP (EcoRI,PstI) | ||
| + | |2.0 | ||
| + | |- | ||
| + | |3.0 | ||
| + | |1.0 | ||
| + | |6.0 pSB2K3 (no.3)(EcoRI,PstI) | ||
| + | |8.0 pCueO-GVP (EcoRI,PstI) | ||
| + | |2.0 | ||
| + | |} | ||
| + | |||
| + | |||
| + | ''Incubate:'' | ||
| + | :* 25°C 50min. | ||
| + | :* kept on ice for 10min. | ||
| + | |||
| + | '''Tranformation''' | ||
| + | |||
| + | Together with these four ligation products, the plasmids from plate 1, 7C and 7K were transformed and grown on LB-kan<sub>50</sub>-IPTG plates to see if a higher colony number could be reached. | ||
| + | :* add 10uL of the ligation product to 50uL competent E.coli TOP10 cells. | ||
| + | :* add 2uL of the 7C and 7K plasmids to 50uL competent E.coli TOP10 cells. | ||
| + | ''Incubate:'' | ||
| + | :* 30 min @ ice | ||
| + | :* 90 sec 42°C | ||
| + | :* 2 min @ ice | ||
| + | :* add 800uL LB-medium-IPTG | ||
| + | :* incubate for 1 h at 37°C | ||
| + | :* plate on LB-kan<sub>50</sub>-IPTG plates | ||
| + | |||
| + | === phase contrast microscope === | ||
| + | |||
| + | The three pictures below show 1000X magnification of pNL29induced for one day (1) for a week(2) and non induced (3) | ||
| + | |||
| + | <gallery> | ||
| + | Image:PNL291day.tif.png|1 | ||
| + | Image:PNL297days.tif.png|2 | ||
| + | Image:PNL290days.tif.png|3 | ||
| + | </gallery> | ||
| + | |||
| + | In the first picture we see some light cell contend compared tot non induced cells. The second picture of an old culture we believe we see the gas vesicles clustered in the bright spot. | ||
===Transporters=== | ===Transporters=== | ||
===Metal Accumulation=== | ===Metal Accumulation=== | ||
| + | Made a [[Team:Groningen/Notebook/24_August_2009#Metal_Accumulation| SmtA-1]] ON culture, the assumption that the fragment should have been ~1200 bp was false and the sequencing results showed that neither [[Team:Groningen/Notebook/24_August_2009#Metal_Accumulation| SmtA-2]] or [[Team:Groningen/Notebook/24_August_2009#Metal_Accumulation| SmtA-3]] contained the proper construct. <partinfo>BBa_K190021</partinfo> should be around ~700 bp including prefix and suffix, instead of 900 bp. As [[Team:Groningen/Notebook/24_August_2009#Metal_Accumulation| SmtA-1]] was positioned lower than the others it might contain the proper construct. | ||
| + | |||
| + | ===Glycerol stocks (-80 °C)=== | ||
| + | {|cellpadding="5" cellspacing="1" border="5" style="font-size: small;" | ||
| + | |'''Component''' | ||
| + | |'''Description''' | ||
| + | |'''Part or Accession #''' | ||
| + | |'''Base Pairs (bp)''' | ||
| + | |'''Plasmid (backbone)''' | ||
| + | |'''Resistance''' | ||
| + | |'''Well''' | ||
| + | |'''Quality control''' | ||
| + | |'''Storage Medium+Antibiotics''' | ||
| + | |'''Organism''' | ||
| + | |'''Date of storage''' | ||
| + | |- | ||
| + | |fMT #4 | ||
| + | |Metallothionein for As<sup>3+</sup> | ||
| + | |<partinfo>BBa_K190019</partinfo> | ||
| + | |222 | ||
| + | |<partinfo>pSB1AC3</partinfo> | ||
| + | |Ampicillin/chloramphenicol | ||
| + | |plate 3, pos. 60 | ||
| + | |{{done|Sequence Okay!}} | ||
| + | |LB-Ampicillin | ||
| + | |[http://openwetware.org/wiki/E._coli_genotypes#TOP10_.28Invitrogen.29 <i>E. coli</i> TOP10] | ||
| + | |04september2009 | ||
| + | |} | ||
===Vectors=== | ===Vectors=== | ||
Latest revision as of 11:10, 7 September 2009
[http://2009.igem.org/Team:Groningen http://2009.igem.org/wiki/images/f/f1/Igemhomelogo.png]
|
|---|
Wet
GVP Cluster
[http://partsregistry.org/Part:BBa_F2620 BBa_F2620]
- → DONE Isolate plasmid from o.n. precultures of pLacI-GVP and pBad-araC-GVP in pSB1A2
- → DONE Restriction control of plasmids and purification of wanted fragments for ligation in pSB2K3 vector
- → DONE Ligate pArsR-GVP, pZntR-GVP, pCueO-GVP, and pLacI-GVP into vector pSB2A3
- → DONE Transform E.coli Top10 cells with ligation products and pSB2K3 (7C and 7K)
- → DONE Order synthetic DNA for GVP
- → TODO Order primer for PstI site removal
- → DONE Test promoter strenght compared to BBa_J23101 promoter (Sven)
- → TODO Enter sequences of constructs to Sandbox
O.n. precultures
- → All four o.n. precultures showed bacterial growth, and could be used to isolate plasmid. The four are registry vector pSB1A2 with pLacI/pBad-araC and GVP. The isolated plasmid is used for transformation into pSB2K3 and sent for sequencing.
Plasmid Purification
Plasmid isolation was performed on the cultures of E.coli TOP10 containing the above mentioned plasmids with the "Sygma-Aldrich™ [http://www.sigmaaldrich.com/life-science/molecular-biology/dna-and-rna-purification/plasmid-miniprep-kit.html GenElute™] Plasmid Miniprep Kit".
- From each tube 4mL of culture was collected in a 2.0mL cup, and the cells were pelleted by centrifugation for 1 min. at max. speed and the supernatant discarded.
- Plasmids were eluted with 30μL MQ and stored in the fridge
Concentrations
| Plasmid | Conc. ng/μL | 260/280 | 260/230 | -20 box (michael | Restriction Control |
| pSB1A2 pLacI-GVP no.1 | 405.3 | 1.85 | 2.36 | D-6 | Yes (EcoRI/PstI) |
| pSB1A2 pLacI-GVP no.2 | 34.0 | 2.01 | 2.20 | D-7 | Yes (EcoRI/PstI) |
| pSB1A2 pBad/araC-GVP no.1 | 27.5 | 2.09 | 2.16 | D-8 | Yes (EcoRI/PstI) |
| pSB1A2 pBad/araC-GVP no.2 | 33.3 | 1.96 | 2.06 | D-9 | Yes (EcoRI/PstI) |
Restriction for Assembly
The vector [http://partsregistry.org/wiki/index.php?title=Part:pSB1A2 pSB1A2] containing the pLacI and pBad-araC with [http://partsregistry.org/Part:BBa_I750016 GVP] composite parts were cut with PstI and EcoRI to create correct ends for insert into [http://partsregistry.org/Part:pSB2K3 pSB2K3], which was also cut with EcoRI and PstI (4x) on 3-9.
| Plasmid | Amount μL | MQ μL | Fast digest buffer | EcoRI fast digest enzyme | XbaI fast digest enzyme | SpeI fast digest enzyme | PstI fast digest enzyme |
| pSB1A2 pLacI-GVP no.1 | 3.0 | 13.0 | 3.0 | 1.0 | x | x | 1.0 |
| pSB1A2 pLacI-GVP no.2 | 16.0 | x | 3.0 | 1.0 | x | x | 1.0 |
| pSB1A2 pBad/araC-GVP no.1 | 16.0 | x | 3.0 | 1.0 | x | x | 1.0 |
| pSB1A2 pBad/araC-GVP no.2 | 16.0 | x | 3.0 | 1.0 | x | x | 1.0 |
Restriction was kept at 37C for 40 min. and put on ice until used for gel purification.
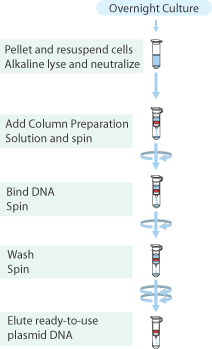
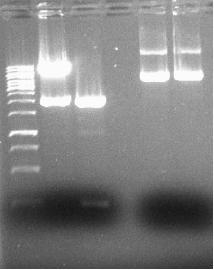
Gel Purification
- In step 7 an amount of 10μL MQ was added to elute the DNA fragments.
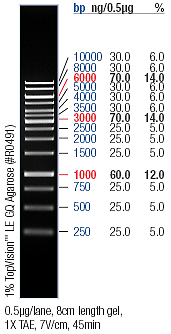
- → Only the first restriction of pLacI-GVP seems to have worked judging from the fragments.
Ligation
A total amount of vector of 100ng was used (GVP) in a 1:3 ratio with insert.
| Ligase buffer (uL) | T4 Ligase (uL) | Plasmid | Insert | MQ |
| 3.0 | 1.0 | 5.0 pSB2K3 (no.2)(EcoRI,PstI) | 3.0 pLacI-GVP (EcoRI,PstI) | 8.0 |
| 3.0 | 1.0 | 5.0 pSB2K3 (no.2)(EcoRI,PstI) | 8.0 pArsR-GVP (EcoRI,PstI) | 3.0 |
| 3.0 | 1.0 | 6.0 pSB2K3 (no.3)(EcoRI,PstI) | 8.0 pZntR-GVP (EcoRI,PstI) | 2.0 |
| 3.0 | 1.0 | 6.0 pSB2K3 (no.3)(EcoRI,PstI) | 8.0 pCueO-GVP (EcoRI,PstI) | 2.0 |
Incubate:
- 25°C 50min.
- kept on ice for 10min.
Tranformation
Together with these four ligation products, the plasmids from plate 1, 7C and 7K were transformed and grown on LB-kan50-IPTG plates to see if a higher colony number could be reached.
- add 10uL of the ligation product to 50uL competent E.coli TOP10 cells.
- add 2uL of the 7C and 7K plasmids to 50uL competent E.coli TOP10 cells.
Incubate:
- 30 min @ ice
- 90 sec 42°C
- 2 min @ ice
- add 800uL LB-medium-IPTG
- incubate for 1 h at 37°C
- plate on LB-kan50-IPTG plates
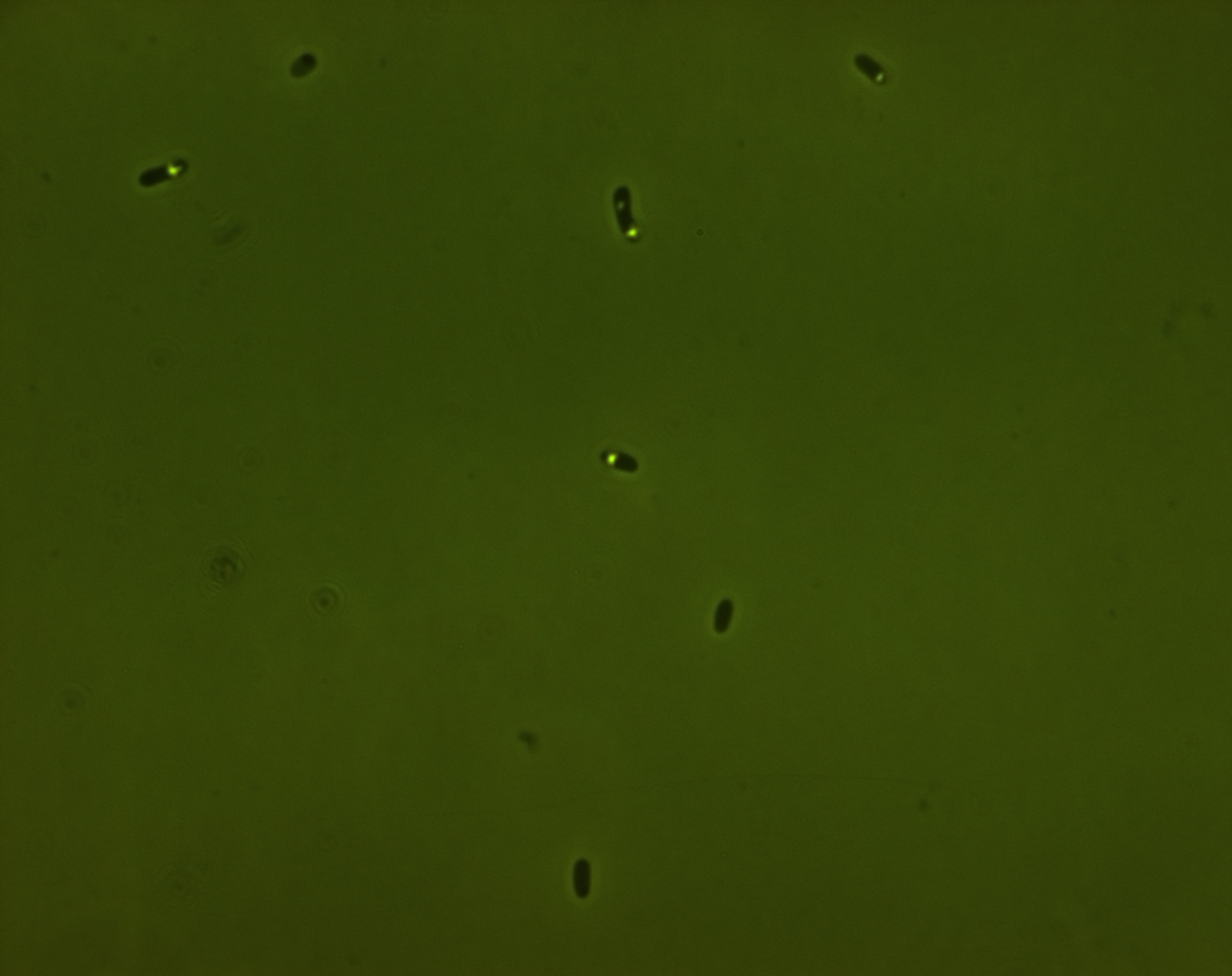
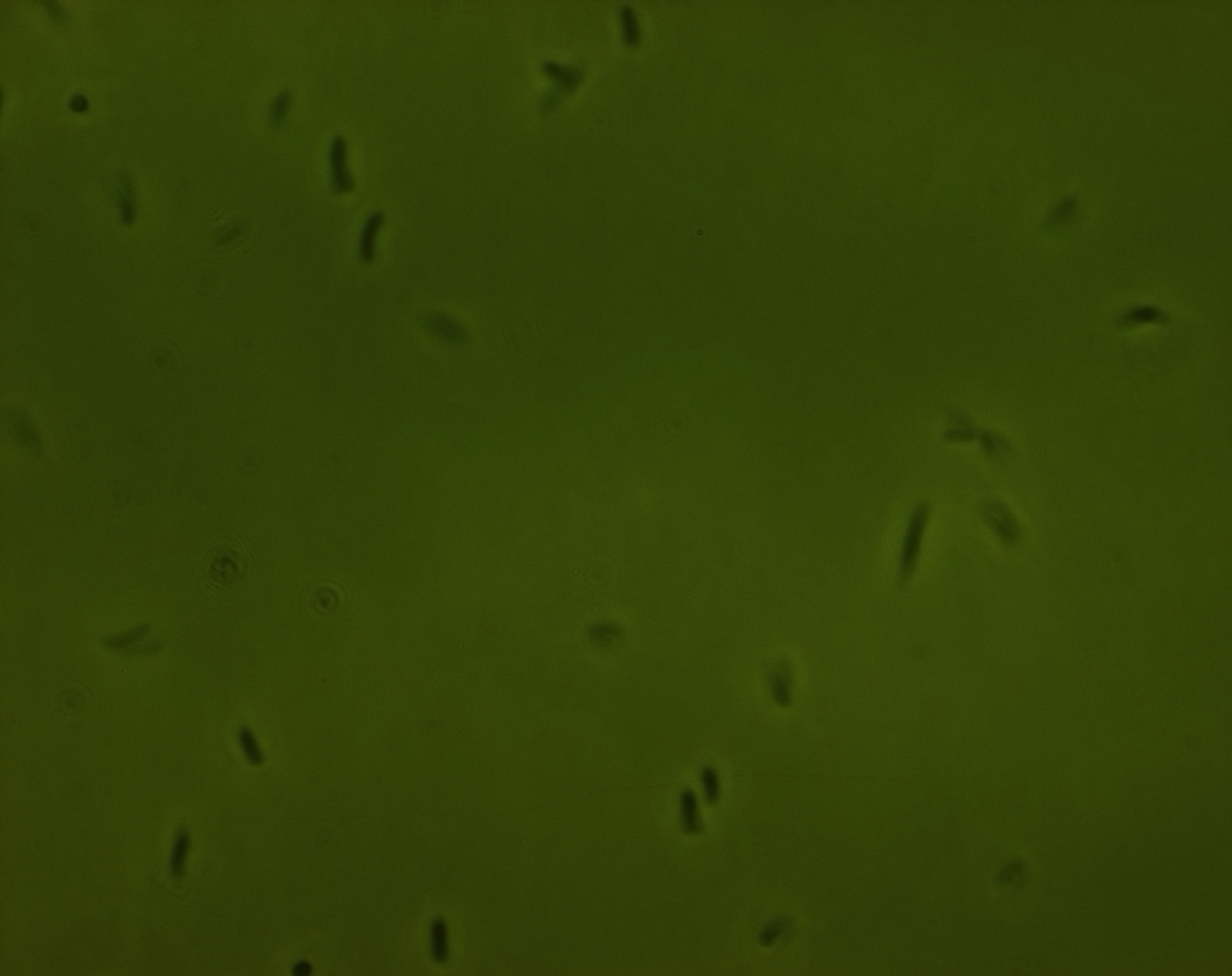
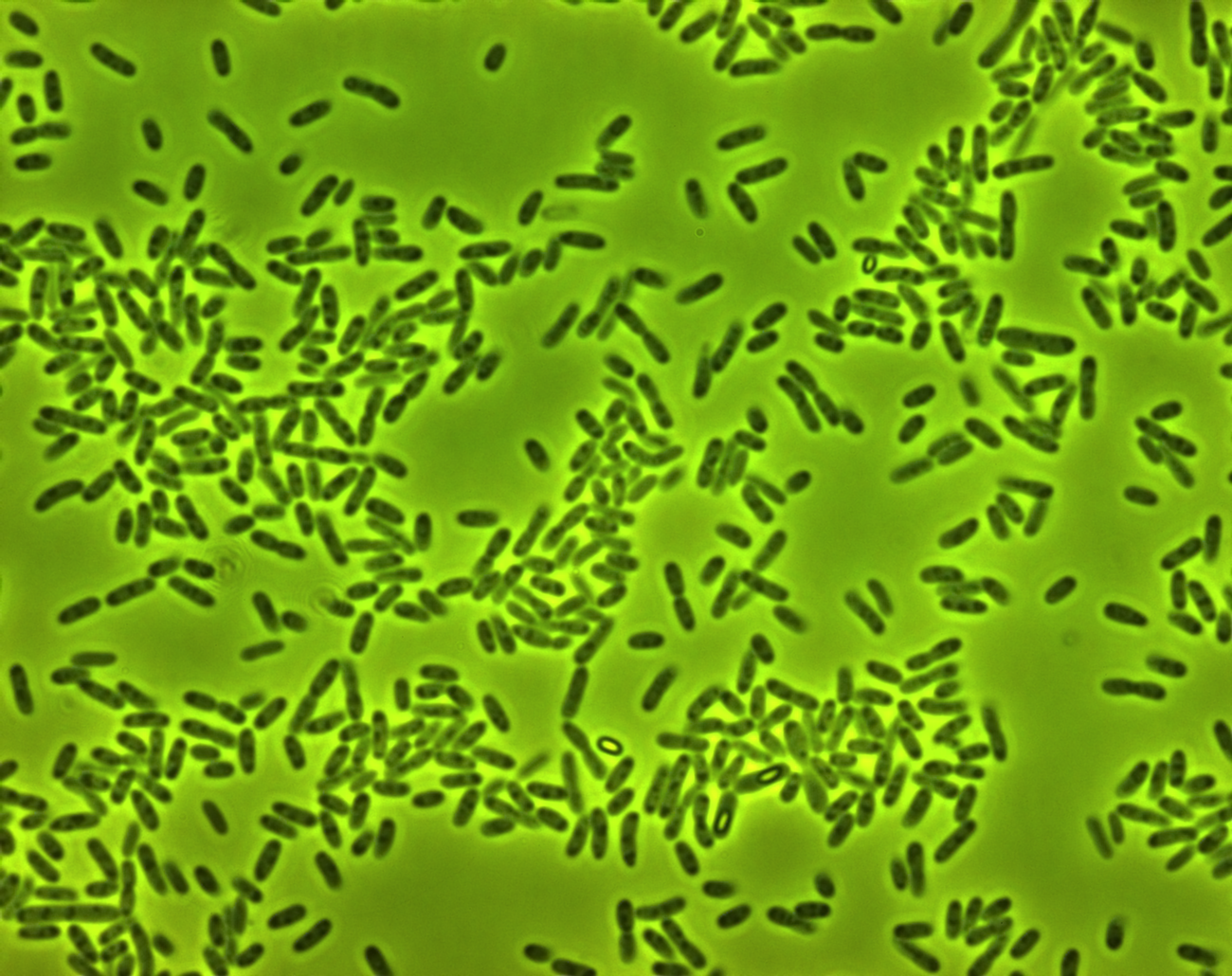
phase contrast microscope
The three pictures below show 1000X magnification of pNL29induced for one day (1) for a week(2) and non induced (3)
In the first picture we see some light cell contend compared tot non induced cells. The second picture of an old culture we believe we see the gas vesicles clustered in the bright spot.
Transporters
Metal Accumulation
Made a SmtA-1 ON culture, the assumption that the fragment should have been ~1200 bp was false and the sequencing results showed that neither SmtA-2 or SmtA-3 contained the proper construct. should be around ~700 bp including prefix and suffix, instead of 900 bp. As SmtA-1 was positioned lower than the others it might contain the proper construct.
Glycerol stocks (-80 °C)
| Component | Description | Part or Accession # | Base Pairs (bp) | Plasmid (backbone) | Resistance | Well | Quality control | Storage Medium+Antibiotics | Organism | Date of storage |
| fMT #4 | Metallothionein for As3+ | 222 | Ampicillin/chloramphenicol | plate 3, pos. 60 | Sequence Okay! | LB-Ampicillin | [http://openwetware.org/wiki/E._coli_genotypes#TOP10_.28Invitrogen.29 E. coli TOP10] | 04september2009 |
Vectors
Dry
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"