Team:Newcastle/Labwork/4 September 2009
From 2009.igem.org
Babyneurone (Talk | contribs) (→Overview) |
Babyneurone (Talk | contribs) (→Sequencing BBa_J33206 taken from the Spring Distribution) |
||
| (23 intermediate revisions not shown) | |||
| Line 2: | Line 2: | ||
{{:Team:Newcastle/Header}} | {{:Team:Newcastle/Header}} | ||
{{:Team:Newcastle/Left}} | {{:Team:Newcastle/Left}} | ||
| - | |||
__NOTOC__ | __NOTOC__ | ||
| - | =Lab | + | [[Image:Team Newcastle 2009 iGEM ProbationaryP-Sign.PNG|50px|right]] |
| + | =Formal Lab Session - 4th September 2009= | ||
| + | [[Image:Team Newcastle 2009 iGEM 04-09-09 IMG 0776.JPG|400px|center]] | ||
| + | <br> | ||
=<font color="Orange"><u>Overview</u></font>= | =<font color="Orange"><u>Overview</u></font>= | ||
<font color="Orange"> | <font color="Orange"> | ||
| - | *[[#Promoter Library Sub-Project|Promoter Library Team]] '''- PCR products analysis''' | + | *[[#Promoter Library Sub-Project|Promoter Library Team]] '''- C0, V1, V2 and V3 PCR products analysis''' |
<br> | <br> | ||
*[[#Stochastic Switch Team|Stochastic Switch Team]] '''- E.Coli transformation with pSB1AT3 + sspb''' | *[[#Stochastic Switch Team|Stochastic Switch Team]] '''- E.Coli transformation with pSB1AT3 + sspb''' | ||
<br> | <br> | ||
| - | *[[#Metal Sensing Team|Metal Sensing Team]] '''- Sequencing ''BBa_J33206'' taken from the Spring Distribution and ''czrA'', ''ArsR'' and ''cadA'' promoter PCR''' | + | *[[#Metal Sensing Team|Metal Sensing Team]] '''- Sequencing ''BBa_J33206'' taken from the Spring Distribution and ''czrA'', ''ArsR'' and ''cadA'' promoter PCR amplification''' |
</font> | </font> | ||
| Line 19: | Line 21: | ||
==<u>Promoter Library Sub-Project</u>== | ==<u>Promoter Library Sub-Project</u>== | ||
===Introduction=== | ===Introduction=== | ||
| + | [[Image:Team Newcastle 2009 iGEM 04-09-09 IMG 0822.JPG|200px|right]] | ||
In yesterday's lab session the products from the PCR reactions on the C0, V1, V2 and V3 samples were purified. The gel electrophoresis carried out on the unpurified samples the previous day showed hazy bands however it was uncertain that these bands were the promoters or just primer-dimers. Today, we aim to carry out DNA gel electrophoresis on these purified samples (in 1.5% agarose gel) to determine whether any PCR amplification reactions worked. | In yesterday's lab session the products from the PCR reactions on the C0, V1, V2 and V3 samples were purified. The gel electrophoresis carried out on the unpurified samples the previous day showed hazy bands however it was uncertain that these bands were the promoters or just primer-dimers. Today, we aim to carry out DNA gel electrophoresis on these purified samples (in 1.5% agarose gel) to determine whether any PCR amplification reactions worked. | ||
<br> | <br> | ||
| + | |||
===Preparing samples and DNA Gel Electrophoresis=== | ===Preparing samples and DNA Gel Electrophoresis=== | ||
The purified PCR products were treated in a similar way to which DNA samples are treated in the gel electrophoresis protocol except 5ul of each sample was extracted and to each of these 1ul of loading buffer (dye) added. 15ul of ''HindIII''' ladder will be added in the first and last wells of the 1.5% agarose gel. This is how the wells will be organised: | The purified PCR products were treated in a similar way to which DNA samples are treated in the gel electrophoresis protocol except 5ul of each sample was extracted and to each of these 1ul of loading buffer (dye) added. 15ul of ''HindIII''' ladder will be added in the first and last wells of the 1.5% agarose gel. This is how the wells will be organised: | ||
| Line 37: | Line 41: | ||
[[Image:Team Newcastle 2009 iGEM Geldoc 2009-09-04 16hr 52min.jpg|450px|center]] | [[Image:Team Newcastle 2009 iGEM Geldoc 2009-09-04 16hr 52min.jpg|450px|center]] | ||
==<u>Stochastic Switch Team</u>== | ==<u>Stochastic Switch Team</u>== | ||
| + | [[Image:Team Newcastle 2009 iGEM 04-09-09 IMG 0782.JPG|200px|right]] | ||
Today we ran the sac and ara PCR products from yesterday on an 0.8% agarose gel in order to determine whether they were worth using the clean up kit on. We ran two PCRs for each set of primers so that resulting fragments could be combined for a more successful ligation. As it happens there was a good result for the sac product but no fully clear bands for the ara product. However both PCR products for sac and ara were combined and the PCR clean up procedure was carried out according to the protocol. Following clean up a digest was done on both products using EcoRI and SpeI. This was done using the fast digest enzymes and buffer, however the reactions were left for an hour. | Today we ran the sac and ara PCR products from yesterday on an 0.8% agarose gel in order to determine whether they were worth using the clean up kit on. We ran two PCRs for each set of primers so that resulting fragments could be combined for a more successful ligation. As it happens there was a good result for the sac product but no fully clear bands for the ara product. However both PCR products for sac and ara were combined and the PCR clean up procedure was carried out according to the protocol. Following clean up a digest was done on both products using EcoRI and SpeI. This was done using the fast digest enzymes and buffer, however the reactions were left for an hour. | ||
Rather than running these products on a wide well agarose gel the PCR clean up procedure was carried out to save time and avoid losing as much product. The resulting elutions were labaeeled and put into the -20 freezer in the midi-preps box. | Rather than running these products on a wide well agarose gel the PCR clean up procedure was carried out to save time and avoid losing as much product. The resulting elutions were labaeeled and put into the -20 freezer in the midi-preps box. | ||
| - | Goksel carried out an ''E.coli'' transformation using the ligation product from yesterday (pSB1AT3 + sspb) however we are unsure whether the protocol used was appropriate for the ''E.coli'' strains used. For this reason on Monday we will try ligating all three fragments (sac, ara as well as sspb) with the pSB1AT3 backbone again. For this we will try the new Fast Ligation/Transformation kit to try to save time. | + | Goksel carried out an ''E.coli'' transformation using the ligation product from yesterday (pSB1AT3 + sspb) however we are unsure whether the protocol used was appropriate for the ''E.coli'' strains used. For this reason on Monday we will try ligating all three fragments (sac, ara as well as sspb) with the pSB1AT3 backbone again. For this we will try the new Fast Ligation/Transformation kit to try to save time. |
| + | |||
| + | ===Transformation=== | ||
| + | We carried out the ''E. coli'' transformation using commercial cells with ligated pSB1AT3 + sspB. | ||
| + | We prepared four tubes with the following mix | ||
| + | |||
| + | Tube 1: | ||
| + | 40ul competent cells | ||
| + | 30ul backbone + insert from the ligation step(psB1AT3 + sspB) | ||
| + | |||
| + | Tube 2: | ||
| + | 40ul competent cells | ||
| + | 20ul backbone from the ligation step (cut backbone) | ||
| + | |||
| + | Tube 3: | ||
| + | 40ul competent cells | ||
| + | 3ul uncut pSB1AT3(As the positive transformation control) | ||
| + | |||
| + | Tube 4: | ||
| + | 40ul competen t cells | ||
| + | 3ul water (As the negative transformation control) | ||
| + | |||
| + | After carrying out the transformation steps and incubating at shaking incubator at 37C, we plated them out | ||
| + | Tube 1 plates: | ||
| + | 400ul plated out to LB + Amp | ||
| + | 200ul plated out to LB + Amp | ||
| + | (20ul + 180ul water) plated out to LB + Amp | ||
| + | 200ul lated out to LB | ||
| + | |||
| + | Tube 2 plates | ||
| + | 200ul to LB + Amp (Should not grow) | ||
| + | 200ul to LB | ||
| + | |||
| + | Tube 3 plates | ||
| + | 200ul to LB | ||
| + | 2ooul to LB+Amp | ||
| + | |||
| + | Tube 4: | ||
| + | 200ul to LB + Amp (Should not grow) | ||
| + | 200ul to LB | ||
==<u>Metal Sensing Team</u>== | ==<u>Metal Sensing Team</u>== | ||
===Introduction=== | ===Introduction=== | ||
In light of the ''BBa_J33206'' BioBrick (the arsR gene + arsR binding site) from the Spring Distribution possibly lacking the ''lacZ'' gene (as described in the Parts Registry), Chris French from Edinburgh University has kindly sent us the original BioBrick DNA as well as forward and reverse sequencing primers and forward and reverse PCR primers. | In light of the ''BBa_J33206'' BioBrick (the arsR gene + arsR binding site) from the Spring Distribution possibly lacking the ''lacZ'' gene (as described in the Parts Registry), Chris French from Edinburgh University has kindly sent us the original BioBrick DNA as well as forward and reverse sequencing primers and forward and reverse PCR primers. | ||
| + | [[Image:Team Newcastle 2009 iGEM 04-09-09 IMG 0785.JPG|200px|right]] | ||
The sequencing primers should enable us to determine whether the ''BBa_J33206'' BioBrick DNA (which we have midi-prepped)from the Spring Distribution contains the ''lacZ'' gene; if it does not then this means the unusually light bands seen in the gel (created from digests of the ''pSB1A2'' plasmid which released the ''BBa_J33206'' BioBrick) are not erroneous at all. No matter what the result, we should transform some ''E.coli'' with the 5ul of DNA sent to us. | The sequencing primers should enable us to determine whether the ''BBa_J33206'' BioBrick DNA (which we have midi-prepped)from the Spring Distribution contains the ''lacZ'' gene; if it does not then this means the unusually light bands seen in the gel (created from digests of the ''pSB1A2'' plasmid which released the ''BBa_J33206'' BioBrick) are not erroneous at all. No matter what the result, we should transform some ''E.coli'' with the 5ul of DNA sent to us. | ||
| Line 63: | Line 108: | ||
'''50ng x 30ul = 1,500ng''' ...... this is approximately the desired amount of DNA present in 30ul | '''50ng x 30ul = 1,500ng''' ...... this is approximately the desired amount of DNA present in 30ul | ||
<br> | <br> | ||
| + | |||
'''1,500ng / 1171.4ng/ul = 1.28ul''' ...... this is approx. the amount of DNA at a concentration of 1171.4ng/ul we will need in the 30ul of solution. | '''1,500ng / 1171.4ng/ul = 1.28ul''' ...... this is approx. the amount of DNA at a concentration of 1171.4ng/ul we will need in the 30ul of solution. | ||
| Line 78: | Line 124: | ||
These solutions were then labelled with the data recorded on a database. The DNA sample plus sequencing primers were then packaged and placed in the collection box. | These solutions were then labelled with the data recorded on a database. The DNA sample plus sequencing primers were then packaged and placed in the collection box. | ||
<br> | <br> | ||
| + | |||
===Transforming ''E. coli'' cells with ''BBa_J33206''=== | ===Transforming ''E. coli'' cells with ''BBa_J33206''=== | ||
| - | The decision was taken to transform some pre-made competent ''DH5-α E. coli'' cells with the ''BBa_J33206'' BioBrick in ''pSB1A2'' plasmid DNA sent to us by Chris French. The protocol used was [https://2009.igem.org/Team:Newcastle/Project/Labwork/PhilsProtocols#Transforming_DNA_.22Phil_Style.22 Dr. Aldridge's Transforming by Heat Shock protocol]. There were only a couple of changes to the protocol and these changes were seen in steps 2 and 5; the time allowed for these steps was 15 minutes instead of 30 minutes. | + | The decision was taken to transform some pre-made competent ''DH5-α E. coli'' cells with the ''BBa_J33206'' BioBrick in ''pSB1A2'' plasmid DNA sent to us by Chris French. |
| + | [[Image:Team Newcastle 2009 iGEM 04-09-09 IMG 0802.JPG|200px|right]] | ||
| + | The protocol used was [https://2009.igem.org/Team:Newcastle/Project/Labwork/PhilsProtocols#Transforming_DNA_.22Phil_Style.22 Dr. Aldridge's Transforming by Heat Shock protocol]. There were only a couple of changes to the protocol and these changes were seen in steps 2 and 5; the time allowed for these steps was 15 minutes instead of 30 minutes. | ||
Once the process was completed, 50ul of the 'transformed' cells was placed on an LB + ampicillin plate (ampicillin being the antibiotic the ''pSB1A2'' offers resistance too) and 100ul of the 'transformed' cells was placed on another LB + ampicillin plate. | Once the process was completed, 50ul of the 'transformed' cells was placed on an LB + ampicillin plate (ampicillin being the antibiotic the ''pSB1A2'' offers resistance too) and 100ul of the 'transformed' cells was placed on another LB + ampicillin plate. | ||
| + | <br> | ||
| + | |||
| + | <br> | ||
===PCR=== | ===PCR=== | ||
| Line 136: | Line 188: | ||
We then will run the PCR products on a gel on the 07/09/2009. | We then will run the PCR products on a gel on the 07/09/2009. | ||
| + | [[Image:Team Newcastle 2009 iGEM 04-09-09 IMG 0818.JPG|350px|center]] | ||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | <br> | ||
| + | |||
| + | ==<u> Chassis team </u>== | ||
| + | [[Image:Team Newcastle 2009 iGEM 04-09-09 IMG 0841.JPG|200px|right]] | ||
| + | Today we design the new primers for gene ''sleB'' and ''cwlJ'', the new primers will arrive at 3 days later. At the same time, we try to change the PCR experiment with new polymerase, reduce the B.subtilis 168's concentration to 2ul. | ||
| + | The total reaction volumn still be 50ul. | ||
| + | |||
| + | PCR reaction contains: | ||
| + | dd H2O xul | ||
| + | GoTag Buffer 10ul | ||
| + | dNTP 2ul | ||
| + | Primer Forward 2.5ul | ||
| + | Primer Reverse 2.5ul | ||
| + | GoTag(5u/ul) 0.25ul | ||
| + | Template 2ul | ||
| + | |||
| + | {{:Team:Newcastle/Project/Labwork/CalTemplate}} | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
{{:Team:Newcastle/Footer}} | {{:Team:Newcastle/Footer}} | ||
{{:Team:Newcastle/Right}} | {{:Team:Newcastle/Right}} | ||
Latest revision as of 12:11, 20 August 2010
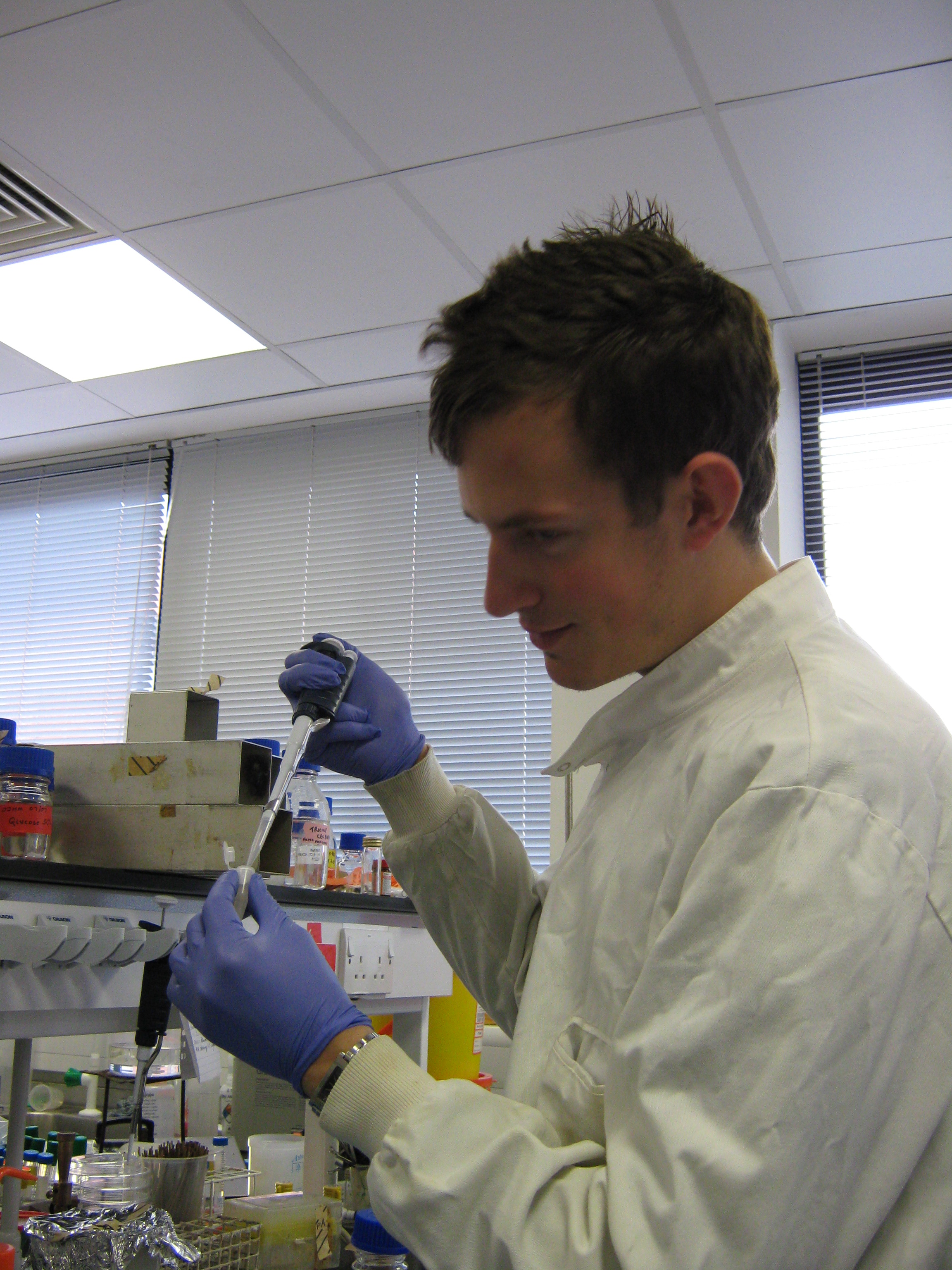
Formal Lab Session - 4th September 2009
Overview
- Promoter Library Team - C0, V1, V2 and V3 PCR products analysis
- Stochastic Switch Team - E.Coli transformation with pSB1AT3 + sspb
- Metal Sensing Team - Sequencing BBa_J33206 taken from the Spring Distribution and czrA, ArsR and cadA promoter PCR amplification
Promoter Library Sub-Project
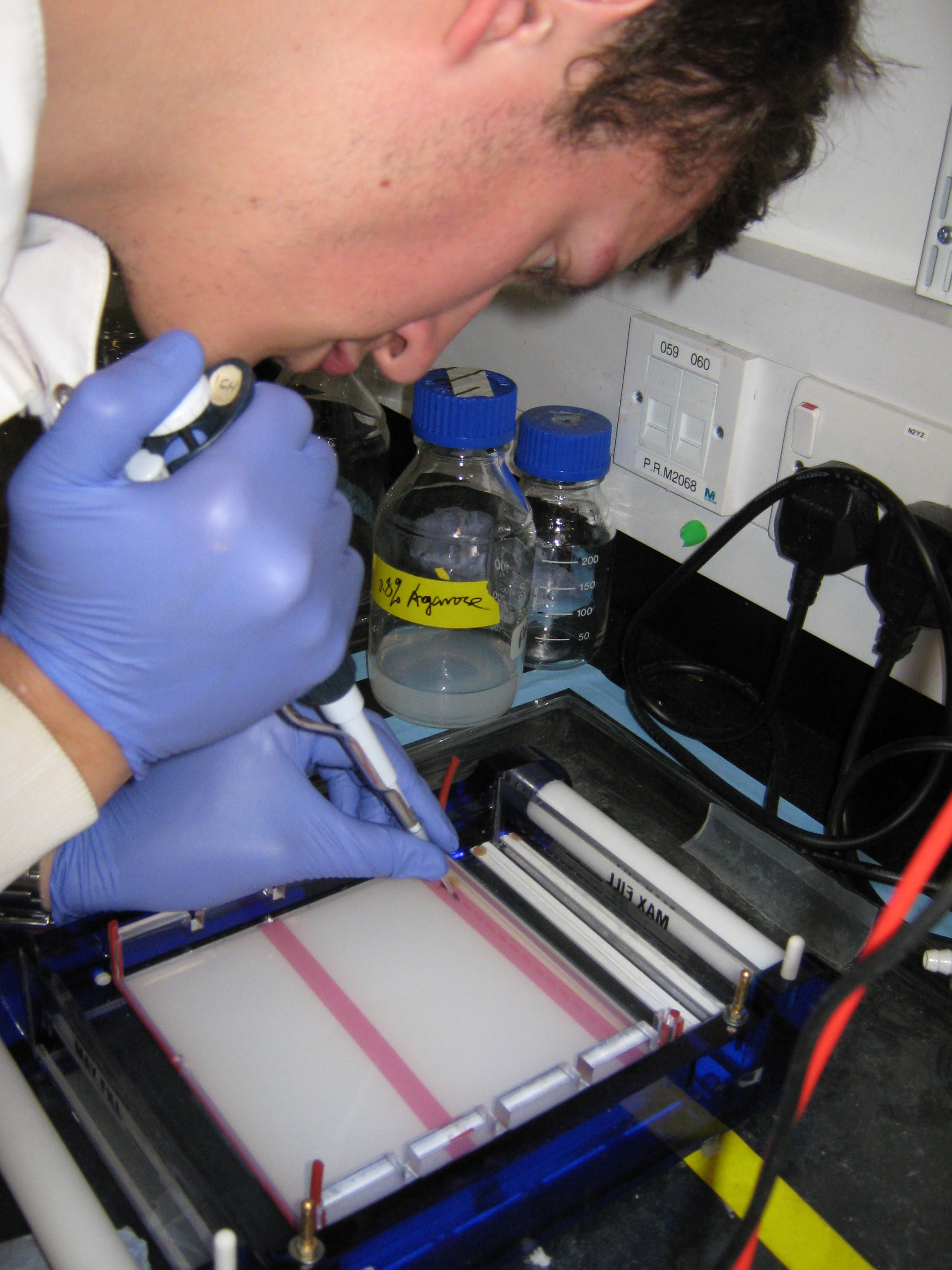
Introduction
In yesterday's lab session the products from the PCR reactions on the C0, V1, V2 and V3 samples were purified. The gel electrophoresis carried out on the unpurified samples the previous day showed hazy bands however it was uncertain that these bands were the promoters or just primer-dimers. Today, we aim to carry out DNA gel electrophoresis on these purified samples (in 1.5% agarose gel) to determine whether any PCR amplification reactions worked.
Preparing samples and DNA Gel Electrophoresis
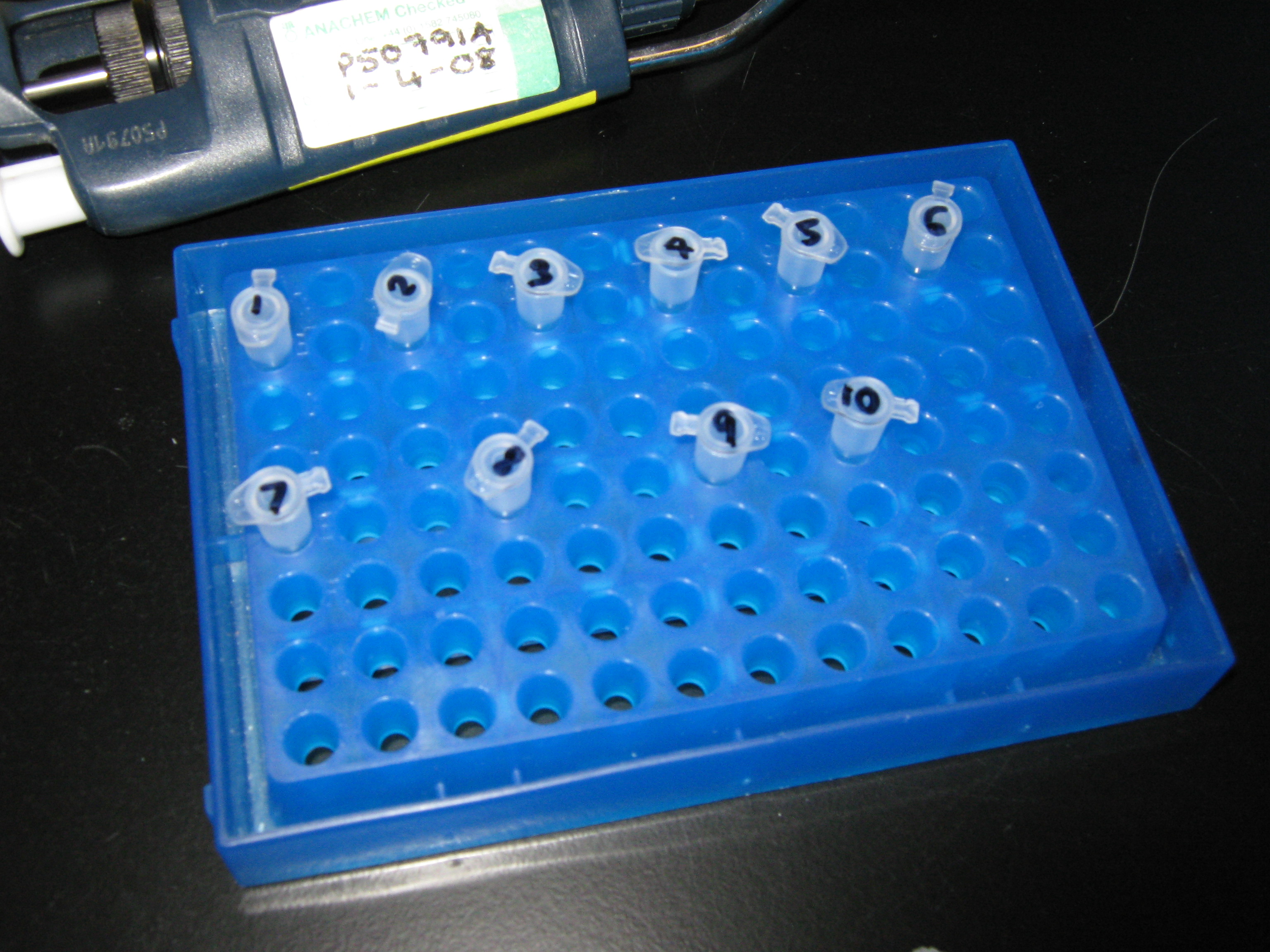
The purified PCR products were treated in a similar way to which DNA samples are treated in the gel electrophoresis protocol except 5ul of each sample was extracted and to each of these 1ul of loading buffer (dye) added. 15ul of HindIII' ladder will be added in the first and last wells of the 1.5% agarose gel. This is how the wells will be organised:
- Lane 1 = blank
- Lane 2 = 15ul of HindIII DNA ladder
- Lane 3 = 6ul of C0 (Sample 1)
- Lane 4 = 6ul of V1 (Sample 2)
- Lane 5 = 6ul of V2 (Sample 3)
- Lane 6 = 6ul of V3 (Sample 4)
- Lane 7 = 15ul of HindIII DNA ladder
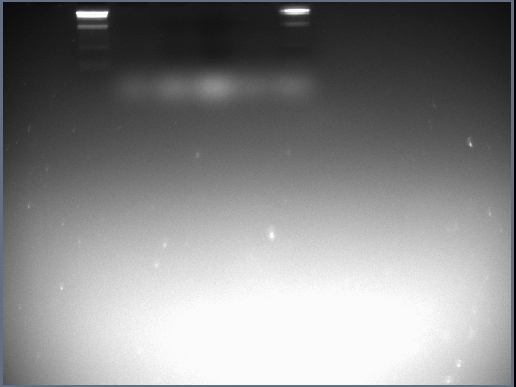
The 1.5% agarose was prepared and poured in the same way as the DNA gel electrophoresis protocol suggests. The samples were then added to the wells in the way described above; the voltage was set to 60mv and left for 15-20 minutes. The photograph obtained from the resulting gel can be seen in the 'Results' section.
Results
Stochastic Switch Team
Today we ran the sac and ara PCR products from yesterday on an 0.8% agarose gel in order to determine whether they were worth using the clean up kit on. We ran two PCRs for each set of primers so that resulting fragments could be combined for a more successful ligation. As it happens there was a good result for the sac product but no fully clear bands for the ara product. However both PCR products for sac and ara were combined and the PCR clean up procedure was carried out according to the protocol. Following clean up a digest was done on both products using EcoRI and SpeI. This was done using the fast digest enzymes and buffer, however the reactions were left for an hour. Rather than running these products on a wide well agarose gel the PCR clean up procedure was carried out to save time and avoid losing as much product. The resulting elutions were labaeeled and put into the -20 freezer in the midi-preps box.
Goksel carried out an E.coli transformation using the ligation product from yesterday (pSB1AT3 + sspb) however we are unsure whether the protocol used was appropriate for the E.coli strains used. For this reason on Monday we will try ligating all three fragments (sac, ara as well as sspb) with the pSB1AT3 backbone again. For this we will try the new Fast Ligation/Transformation kit to try to save time.
Transformation
We carried out the E. coli transformation using commercial cells with ligated pSB1AT3 + sspB. We prepared four tubes with the following mix
Tube 1:
40ul competent cells 30ul backbone + insert from the ligation step(psB1AT3 + sspB)
Tube 2:
40ul competent cells 20ul backbone from the ligation step (cut backbone)
Tube 3:
40ul competent cells 3ul uncut pSB1AT3(As the positive transformation control)
Tube 4:
40ul competen t cells 3ul water (As the negative transformation control)
After carrying out the transformation steps and incubating at shaking incubator at 37C, we plated them out Tube 1 plates:
400ul plated out to LB + Amp 200ul plated out to LB + Amp (20ul + 180ul water) plated out to LB + Amp 200ul lated out to LB
Tube 2 plates
200ul to LB + Amp (Should not grow) 200ul to LB
Tube 3 plates
200ul to LB 2ooul to LB+Amp
Tube 4:
200ul to LB + Amp (Should not grow) 200ul to LB
Metal Sensing Team
Introduction
In light of the BBa_J33206 BioBrick (the arsR gene + arsR binding site) from the Spring Distribution possibly lacking the lacZ gene (as described in the Parts Registry), Chris French from Edinburgh University has kindly sent us the original BioBrick DNA as well as forward and reverse sequencing primers and forward and reverse PCR primers.
The sequencing primers should enable us to determine whether the BBa_J33206 BioBrick DNA (which we have midi-prepped)from the Spring Distribution contains the lacZ gene; if it does not then this means the unusually light bands seen in the gel (created from digests of the pSB1A2 plasmid which released the BBa_J33206 BioBrick) are not erroneous at all. No matter what the result, we should transform some E.coli with the 5ul of DNA sent to us.
Today's experiments involve sending our midi-prepped BBa_J33206 + pSB1A2 DNA (derived from the Spring Distributions) off to be sequenced using the sequencing primers and also to transform E. coli with the BBa_J33206 BioBrick DNA sent to us.
Also, we will carry out PCR reactions along with the sporulation team in order to amplify czrA gene, ArsR BioBrick without the promoter region, and the cadA promoter region.
Sequencing BBa_J33206 taken from the Spring Distribution
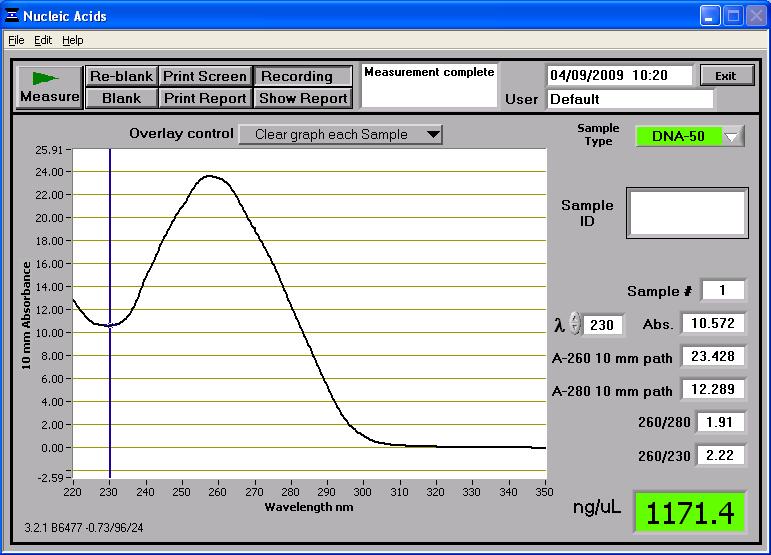
The first step taken was to quantify the midi-prepped BBa_J33206 + pSB1A2 DNA taken from the Spring Distribution (this is the arsR gene plus arsR binding site). When applied to the Nanodrop spectrophotometer, 1171.4ng/ul of DNA was detected. The analysis of the BBa_J33206 DNA concentration can be seen below, including DNA-to-Protein ratios:
The company who sequence the DNA require us to submit 30ul of the DNA and with a recommended concentration of ~50ng/ul. To calculate how much DNA to add to water to make up 30ul of solution with a concentration of 50ng/ul, we did the following:
50ng x 30ul = 1,500ng ...... this is approximately the desired amount of DNA present in 30ul
1,500ng / 1171.4ng/ul = 1.28ul ...... this is approx. the amount of DNA at a concentration of 1171.4ng/ul we will need in the 30ul of solution.
We rounded the 1.28ul up to 2ul (doesn't have to be exactly 50ng/ul) and to this 2ul of DNA we added 28ul of sterile distilled water
A similar step was taken with the sequencing primers. The company which carries out the sequencing requires us to submit 30ul of primers set at a concentration of ~2pmol/ul. The primers (both forward and reverse) have arrived in a concentration of 5pmol/ul.
18ul of water + 12ul primer = 30 ul of primer with a concentration of 2pmol/ul.
These solutions were then labelled with the data recorded on a database. The DNA sample plus sequencing primers were then packaged and placed in the collection box.
Transforming E. coli cells with BBa_J33206
The decision was taken to transform some pre-made competent DH5-α E. coli cells with the BBa_J33206 BioBrick in pSB1A2 plasmid DNA sent to us by Chris French.
The protocol used was Dr. Aldridge's Transforming by Heat Shock protocol. There were only a couple of changes to the protocol and these changes were seen in steps 2 and 5; the time allowed for these steps was 15 minutes instead of 30 minutes.
Once the process was completed, 50ul of the 'transformed' cells was placed on an LB + ampicillin plate (ampicillin being the antibiotic the pSB1A2 offers resistance too) and 100ul of the 'transformed' cells was placed on another LB + ampicillin plate.
PCR
Today we received our new polymerase GoTaq, therefore we are carrying out PCR reactions with thisd polymerase on previous reations which didn't work; czrA gene for the metal sensing team as well as the ArsR BioBrick without it's promoter region and the cadA promoter region, and the sleB and cwlJ gene for the sporulation team.
| No. | Water (ul) | PCR Mix (ul) | Forward Primer (ul) | Reverse Primer (ul) | Template (ul) | Polymerase used (ul) | Total Volume (ul) |
|---|---|---|---|---|---|---|---|
| 1 | 30.75 | 12 | pMA1 2.5ul | pMA2 2.5ul | 168 DNA 2ul | GoTaq (0.25ul) | 50 |
| 2 | 32.75 | 12 | pMA1 2.5ul | pMA2 2.5ul | 0 | GoTaq (0.25ul) | 50 |
| 3 | 30.75 | 12 | pMA7 2.5ul | pMA8 2.5ul | ArsR BioBrick | GoTaq (0.25ul) | 50 |
| 4 | 32.75 | 12 | pMA7 2.5ul | pMA8 2.5ul | 0 | GoTaq (0.25ul) | 50 |
| 5 | 30.75 | 12 | pMA9 2.5ul | pMA10 2.5ul | 168 DNA 2ul | GoTaq (0.25ul) | 50 |
| 6 | 32.75 | 12 | pMA9 2.5ul | pMA10 2.5ul | 0 | GoTaq (0.25ul) | 50 |
| 7 | 30.75 | 12 | sleB 2.5ul | sleB 2.5ul | 168 DNA 2ul | GoTaq (0.25ul) | 50 |
| 8 | 32.75 | 12 | sleB 2.5ul | sleB 2.5ul | 0 | GoTaq (0.25ul) | 50 |
| 9 | 30.75 | 12 | cwlJ 2.5ul | cwlJ 2.5ul | 168 DNA 2ul | GoTaq (0.25ul) | 50 |
| 10 | 32.75 | 12 | cwlJ 2.5ul | cwlJ 2.5ul | 0 | GoTaq (0.25ul) | 50 |
PCR conditions:
For all samples except 3 and 4
1- 95C for 2min
2- 95C for 30sec
3- 50C for 1min
4- 75C for 2min
Step 2-3-4 x 30
5- 75C for 5min
6- 4C Pause
For samples 3 and 4
1- 95C for 2min
2- 95C for 30sec
3- 50C for 3min
4- 75C for 2min
Step 2-3-4 x 30
5- 75C for 5min
6- 4C Pause
We then will run the PCR products on a gel on the 07/09/2009.
Chassis team
Today we design the new primers for gene sleB and cwlJ, the new primers will arrive at 3 days later. At the same time, we try to change the PCR experiment with new polymerase, reduce the B.subtilis 168's concentration to 2ul. The total reaction volumn still be 50ul.
PCR reaction contains:
dd H2O xul GoTag Buffer 10ul dNTP 2ul Primer Forward 2.5ul Primer Reverse 2.5ul GoTag(5u/ul) 0.25ul Template 2ul
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
- Newcastle iGEM Twitter
- [http://www.facebook.com/home.php#/group.php?gid=131709337641 Newcastle on Facebook]
- [http://www.youtube.com/user/newcastle2009igem Newcastle Youtube Channel]
 "
"