Team:Newcastle/Labwork/2 September 2009
From 2009.igem.org
(→Lab Work - 02/09/09) |
|||
| Line 2: | Line 2: | ||
{{:Team:Newcastle/Header}} | {{:Team:Newcastle/Header}} | ||
{{:Team:Newcastle/Left}} | {{:Team:Newcastle/Left}} | ||
| - | |||
__NOTOC__ | __NOTOC__ | ||
| - | =Lab | + | [[Image:Team Newcastle 2009 iGEM ProbationaryP-Sign.PNG|50px|right]] |
| + | =Formal Lab Session - 2nd September 2009= | ||
[[Image:Team Newcastle 2009 iGEM IMG 0681.JPG|400px|center]] | [[Image:Team Newcastle 2009 iGEM IMG 0681.JPG|400px|center]] | ||
<br> | <br> | ||
Revision as of 16:07, 6 October 2009
Formal Lab Session - 2nd September 2009
Metal Sensing Team
Introduction
Before anything else is carried out, the Metal Sensing team will analyse the PCR products from yesterday's PCR reactions by DNA gel electrophoresis - this includes the czrA, sac, ara and sspb genes. The results from this will then decide where we go from here.
If our PCR reactions do not go to plan then today, we will prepare genomic DNA from B.subtilis 168 using the Genomic DNA kit by Sigma-Aldrich as we will need some more to be able to PCR up required genes for all 3 teams potentially.
Practical Outline
These tasks should be carried out by the Metal Sensing team for today:
- Analyse the PCR products from yesterday's PCR reaction by DNA gel electrophoresis
- If PCR reactions have failed:
- Prepare genomic DNA (for Bacillus subtilis) using Sigma-Aldrich's kit
- Reattempt PCR reactions
Observations
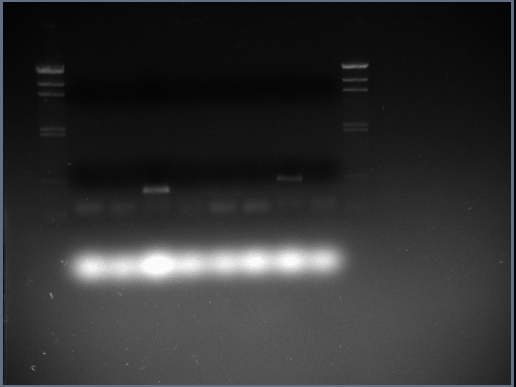
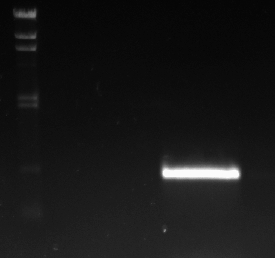
Here is the gel photograph in which the PCR products were analsyed. The lanes are as follows:
- Lane 1 = blank
- Lane 2 = HindIII DNA ladder
- Lane 3 = czrA PCR product
- Lane 4 = czrA PCR control
- Lane 5 = sac PCR product
- Lane 6 = sac PCR control
- Lane 7 = ara PCR product
- Lane 8 = ara PCR control
- Lane 9 = sspb PCR product
- Lane 10 = sspb PCR control
- Lane 11 = HindIII DNA ladder
It can be seen that the only PCR products produced (i.e. PCR reactions that worked) are the sac and sspb coding sequences - the other PCR reactions didn't work.
Procedure
In light of the unsuccessful PCR reaction (attempting to amplify the czrA coding sequence) the Metal Sensing Team will carry out a second PCR attempt. This time it will be done on the prepared genome of B. subtilis
B.subtilis 168 Genomic DNA preparation
We followed the instructions in the kit and got our genomic DNA.
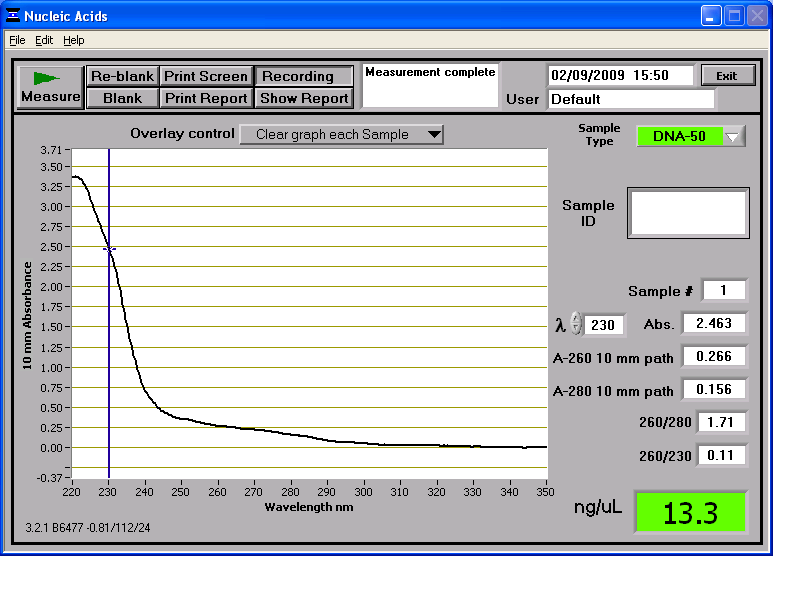
Results from the NanoDrop reading for our 6 preparations:
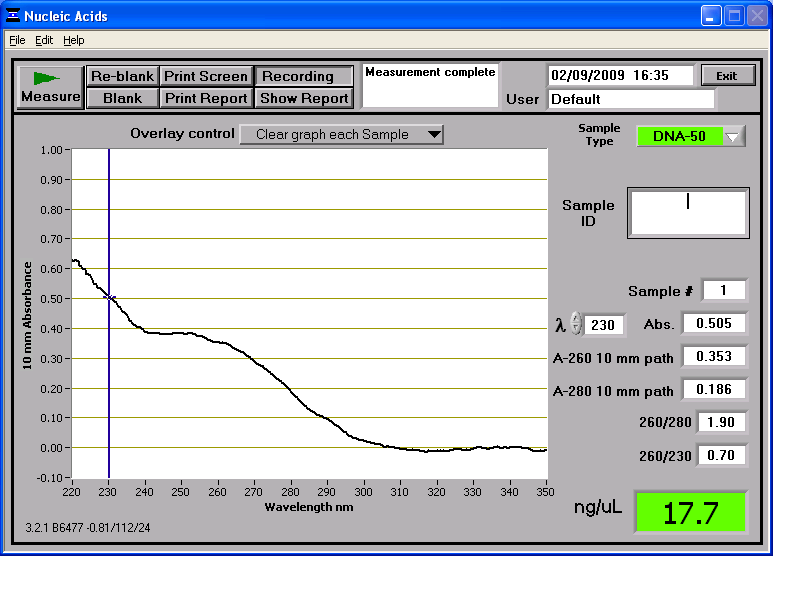
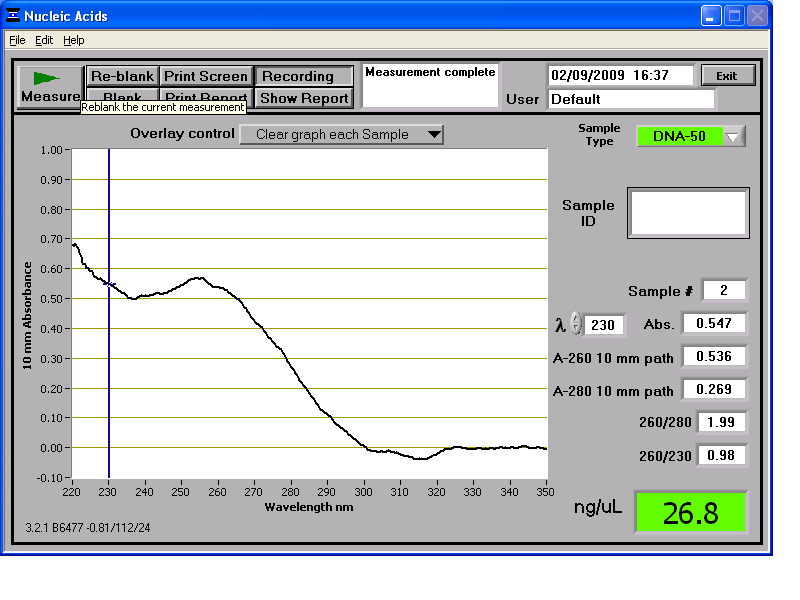

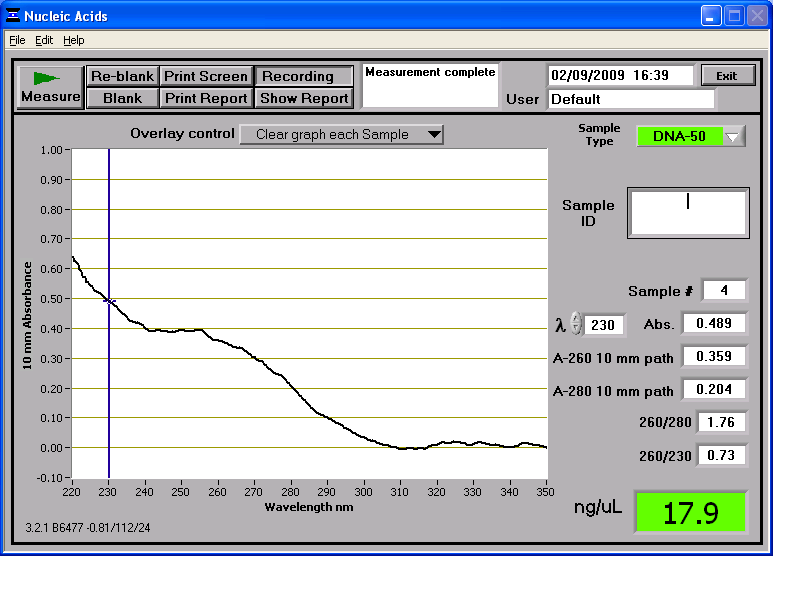
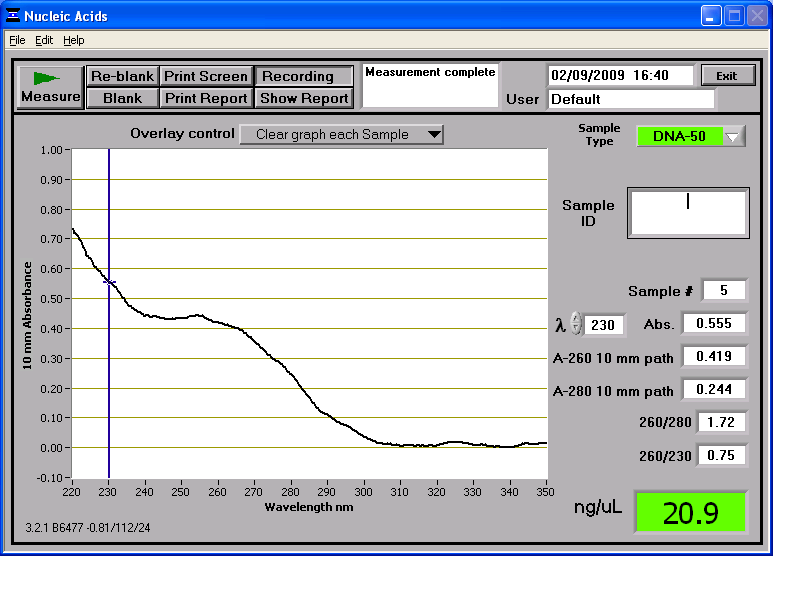
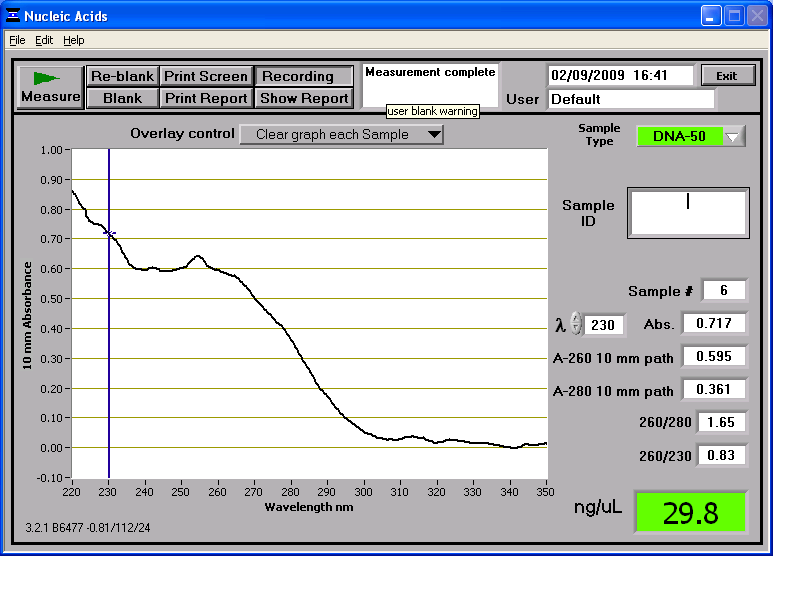
We then stored the genomic DNA in the -20C freezer in the midiprep box.
PCR
We carried out PCR again to amplify czrA and ara genes in the same way as on the 21/08/2009 PCR reaction.
| No. | Water (ul) | PCR Mix (ul) | Forward Primer (ul) | Reverse Primer (ul) | Template (ul) | Polymerase used (ul) | Total Volume (ul) |
|---|---|---|---|---|---|---|---|
| 1 | 24 | 15 | pMA1 2.5ul | pMA2 2.5ul | 168 DNA 5ul | MolTaq (1ul) | 50 |
| 2 | 29 | 15 | pMA1 2.5ul | pMA2 2.5ul | 0 | MolTaq (1ul) | 50 |
| 3 | 24 | 15 | araR_forward 2.5ul | araR_reverse 2.5ul | 168 DNA 5ul | MolTaq (1ul) | 50 |
| 4 | 29 | 15 | araR_forward 2.5ul | araR_reverse 2.5ul | 0 | MolTaq (1ul) | 50 |
PCR conditions:
1- 95C for 2min
2- 95C for 30sec
3- 50C for 1min
4- 75C for 2min
Step 2-3-4 x 30
5- 75C for 5min
6- 4C Pause
Promoter Library Sub-Project
Introduction
In yesterday's session, PCR amplifications for the three types of sigA promoter variants was carried out (i.e. V1, V2 and V3). The amplifications of these three types of promoter variants will give rise to a library of different sigA promoters, hopefully of differing affinities.
In today's session the PCR products will be run on the gel and analysed. If there are signs that the PCR reactions have worked successfully then the PCR products will be purified. If it is apparent that the PCR reactions haven't worked at all then another attempt at PCR shall be made.
Preparing the gel and the DNA gel electrophoresis
The fragments we are expecting to be seen in the gel are roughly 100bp long therefore a fine gel is needed to capture these PCR products. Instead of making up 0.8% agarose gel, 1.5% agarose gel was made up and melted in the way the protocol dictates.
With the gel now set in the tray 6ul (5ul of PCR product and 1ul of loading buffer) of each PCR product was placed into the wells of the gel. The electrophoresis machine was switched on to 90mv and the process allowed to run for 30 minutes.
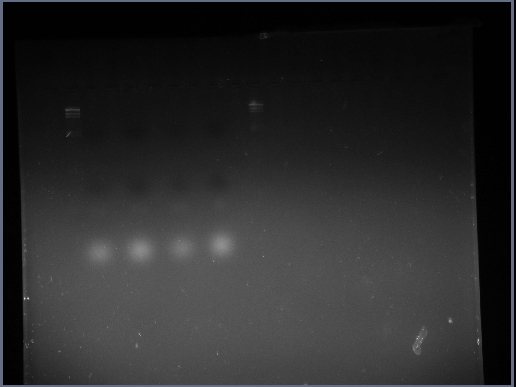
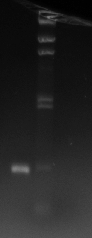
The resulting gel photograph can be seen here:
The samples loaded into the gel were as follows:
- Lane 1 = blank
- Lane 2 = HindIII DNA ladder
- Lane 3 = C0
- Lane 4 = V1
- Lane 5 = V2
- Lane 6 = V3
- Lane 7 = 1ul of neat V1 (control)
- Lane 8 = HindIII DNA ladder
At first glance it seems that the PCRs have completely failed but on closer inspection there are little clouds of flourescing material just below the second row of dark bands (caused by the colour bands)in the middle of the lanes. These could well be PCR products; another attempt at gel electrophoresis would clarify this. It is possible that the DNA gel electrophoresis has been left to run too long and additionally the voltage has been set too high. Of course, the white cloudy substance at the bottom of these wells are the primer-dimer complexes.
Second attempt at DNA gel electrophoresis
The result of the gel electrophoresis called for another attempt at running the PCR products on the 1.5% agarose gel. The samples were prepared in the following way:
- 5ul of the 4 PCR samples (C0, V1, V2, V3) were extracted and to each 1ul loading buffer was added
- 1ul of neat V1 (the control) was diluted in 8ul water - 1ul of loading buffer was added also
- 2 x 15ul of HindIII ladder
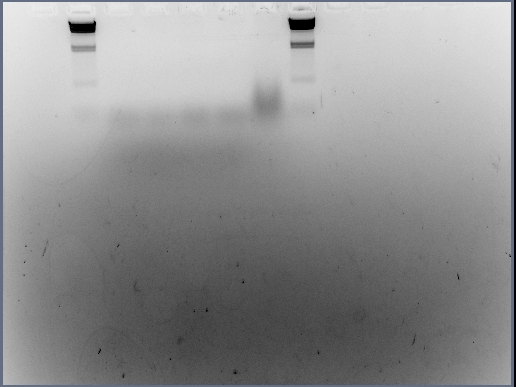
The voltage was cut down to 60mv and the gel was only allowed to run for 15-20 minutes - this was to ensure that the DNA was not run too fast through the gel. The result of this procedure can be seen below:
The samples displayed are as follows:
- Lane 1 = blank
- Lane 2 = 15ul of HindIII DNA ladder
- Lane 3 = 6ul of C0
- Lane 4 = 6ul of V1
- Lane 5 = 6ul of V2
- Lane 6 = 6ul of V3
- Lane 7 = 6ul of neat V1
- Lane 8 = 15ul of HindIII DNA ladder
This is certainly a better gel attempt than the previous one as 'bands' can be seen. However it is not clear whether the approx 100bp is excess template or the real product.
Conclusion: Repeat the PCR dropping the annealing temperature down to 45C and proceed with trying to clean up the products from lanes 3-6
Chassis team
Today we began the lab work on our PCR primers. We attempted to use two pairs of our primers; the ones to amplify the sleB and cwlJ genes from the B. subtilis genome. We prepared the solutions needed, and followed the PCR protocol. We rehydrated our primers, and diluted them accordingly. The TMs were calculated by using 5 degrees lower than the given temperature. We prepared two lots of solutions, one with genomic DNA and one without (our controls). They were loaded into the PCR machine, which ran for around 1 hour 53 minutes.
PCR
We carried out PCR of gene "sleB" and "cwlJ" using primer:
- sleBForward & sleBReverse
- cwlJForward & cwlJReverse
PCR conditions:
- No.1 & 2 : 95C 2min -->[95C 30S --> 53C 30S --> 75C 1min] 30 runs --> 75C 5min --> store at 4C
- No.3 & 4 : 95C 2min -->[95C 30S --> 55C 30S --> 75C 1min] 30 runs --> 75C 5min --> store at 4C
| No. | Water (ul) | PCR Mix (ul) | Forward Primer (ul) | Reverse Primer (ul) | Template (ul) | Polymerase used (ul) | Total Volume (ul) |
|---|---|---|---|---|---|---|---|
| 1 | 24 | 15 | sleBForward 2.5ul | sleBReverse 2.5ul | B.subtilis DNA 5ul | MolTaq (1ul) | 50 |
| 2 | 29 | 15 | sleBForward 2.5ul | sleBReverse 2.5ul | 0 | MolTaq (1ul) | 50 |
| 3 | 24 | 15 | cwlJForward 2.5ul | cwlJReverse 2.5ul | B.subtilis DNA 5ul | MolTaq (1ul) | 50 |
| 4 | 29 | 15 | cwlJForward 2.5ul | cwlJReverse 2.5ul | 0 | MolTaq (1ul) | 50 |
PCR product test
Run PCR product on 0.8% agarose gel
When we ran the products on a gel, we noticed a light blue band, a dark blue band and a yellow band - however these were for all of the solutions, control included. Putting the gel under UV light showed this too. We concluded that something had gone wrong. The stochastic switch team also used a well on the same gel, which worked, so we concluded that the gel was not the likely cause of the error. Therefore something went wrong at either the preparation for the PCR stage, or during the PCR. The gel image can be seen below:
We are going to try this again when we are next in the lab.
Stochastic Switch Team
Summary
Today we froze our strains into the TPA collection, by freezing them down in DMSO in the -80 freezers both in the lab as well as duplicates in the stock freezer. Goksel did a digest of the sspb PCR product that was cleaned up yesterday.
Yesterday a PCR was done of CzrA, sacA, ara, and sspb. The products of these were run on a gel, the results of which are shown below. It seemed that there was no PCR product for both the CzrA, and ara, however there is a shadow from all of the wells at around 200bp that could be contamination of PCR product- we are unsure.
Despite inconclusive gel results a PCR product clean up was done on all four reactions. Restriction digests were done on the sac and sspb purified products, which had reasonably clear bands on the gel. These were cut with EcoRI and pstI ready for ligation into the plasmid backbone.
sspB digest and DNA extract
We digested sspB and run the final 70ul of the mix + 10ul of loading buffer on the gel with large wells. We used the ingredients below to prepare the 70ul of the final solution
Water, 16ul Buffer, 7ul sspB PCR product, 41ul EcoRI, 3ul SpeI, 3ul
We mixed the solution well and incubated at 37C for an hour.
We then cut the gel fragment containing our sspB cut with EcoRI and SpeI sites. The DNA of the sspB insert was then extracted from this gel fragment.
Gel fragment was measured as 378 mg and we used 3x378=1134ul of gel soluzable gel and 378ul of isopropanol to extract the DNA.
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
- Newcastle iGEM Twitter
- [http://www.facebook.com/home.php#/group.php?gid=131709337641 Newcastle on Facebook]
- [http://www.youtube.com/user/newcastle2009igem Newcastle Youtube Channel]
 "
"