Team:SJTU-BioX-Shanghai/The project
From 2009.igem.org
Pnstontale (Talk | contribs) |
Pnstontale (Talk | contribs) |
||
| Line 1: | Line 1: | ||
{{Template:SJTU09_header}} | {{Template:SJTU09_header}} | ||
{{Template:SJTU09_menubar}} | {{Template:SJTU09_menubar}} | ||
| - | + | <html><h0>Our project</h0></html> | |
| + | ==Background== | ||
It is universally acknowledged that bioclock works as a circadian regulator in most eukaryotic multicellular species. This mechanism controls higher plants’ blossom time, brings insects into metamorphosis, and also wakes us up every day. | It is universally acknowledged that bioclock works as a circadian regulator in most eukaryotic multicellular species. This mechanism controls higher plants’ blossom time, brings insects into metamorphosis, and also wakes us up every day. | ||
| Line 9: | Line 10: | ||
[[Image:SJTU09_Hypnos_happy.gif|left|thumb|250px|Lovely Hypnos]] | [[Image:SJTU09_Hypnos_happy.gif|left|thumb|250px|Lovely Hypnos]] | ||
[[Image:SJTU09_Hypnos_angry.gif|left|thumb|273px|Hypnos's curse]] | [[Image:SJTU09_Hypnos_angry.gif|left|thumb|273px|Hypnos's curse]] | ||
| - | |||
| - | |||
Hence, we constructed our bacteria bioclock by utilizing the toxin-antitoxin system (TA system), which forms an oscillator between two physiological states--dormancy and activity. | Hence, we constructed our bacteria bioclock by utilizing the toxin-antitoxin system (TA system), which forms an oscillator between two physiological states--dormancy and activity. | ||
| Line 18: | Line 17: | ||
Based on these mechanisms we have designed an ingenious genetic network which functions as a bacterial bioclock oscillating between the two states of dormancy and activity. It is exciting to imagine that we manage to manipulate the lifespan of E.coli by switching the oscillator on, since the metabolic process of microbes is vastly decelerated during the dormancy state, just like bears and hedgehogs in their hibernation. | Based on these mechanisms we have designed an ingenious genetic network which functions as a bacterial bioclock oscillating between the two states of dormancy and activity. It is exciting to imagine that we manage to manipulate the lifespan of E.coli by switching the oscillator on, since the metabolic process of microbes is vastly decelerated during the dormancy state, just like bears and hedgehogs in their hibernation. | ||
| - | [[Image:SJTU09_ecoli_mascot. | + | [[Image:SJTU09_ecoli_mascot.jpg|left|thumb|125px|E.Coli Mascot]] |
| - | [[Image:SJTU09_ecoli_asleep. | + | [[Image:SJTU09_ecoli_asleep.jpg|left|thumb|212px|E.Coli asleep]] |
| - | [[Image:SJTU09_ecoli_awake. | + | [[Image:SJTU09_ecoli_awake.jpg|left|thumb|161px|E.Coli awake]] |
| - | + | ||
| - | + | ||
An example of how this artificial bioclock could be applied might be the preservation of scientifically valuable bacteria which mutate frequently. During the dormancy state bacteria hardly undergo mutation; therefore their genetic characteristics are retained. | An example of how this artificial bioclock could be applied might be the preservation of scientifically valuable bacteria which mutate frequently. During the dormancy state bacteria hardly undergo mutation; therefore their genetic characteristics are retained. | ||
Other potential applications such as biologic timing and antibiotic resistance remain intriguing to explore. | Other potential applications such as biologic timing and antibiotic resistance remain intriguing to explore. | ||
| + | |||
| + | ==Principle & Network== | ||
| + | |||
| + | ;<center><font size=4>'''''E.coli''''' the napper</font></center> | ||
| + | |||
| + | Bacteria and at least some archaea harbor ‘suicide systems’, the existence of which is a mystery to those who hear about them for the first time. These systems consist of a lethal toxin that can be neutralized by an antitoxin. The production of antitoxin, which is labile, must at least parallel that of the toxin, which is usually stable, to circumvent the induction of cell poison. To achieve this, both proteins are encoded within a single operon. The toxins seem to target specific, essential cellular processes. Perhaps the best characterized of these system is relBE system, the genes of which are present on the '''''Escherichia coli''''' chromosome. The relE toxin is an RNase that preferentially cleaves mRNAs bound to the ribosome at the second position of stop codons. Stop codons not only signal the end of the protein coding sequence but also serve as the binding site for release factors, which promote release of the nascent polypeptide and facilitate recycling of ribosomes for further rounds of translation. Thus truncated mRNA by cleavage of relE lacks appropriate termination signals, which causes the accumulation of stalled ribosomes and these mRNAs are unable to promote release factor binding, nascent polypeptide release, and ribosome recycling. | ||
| + | |||
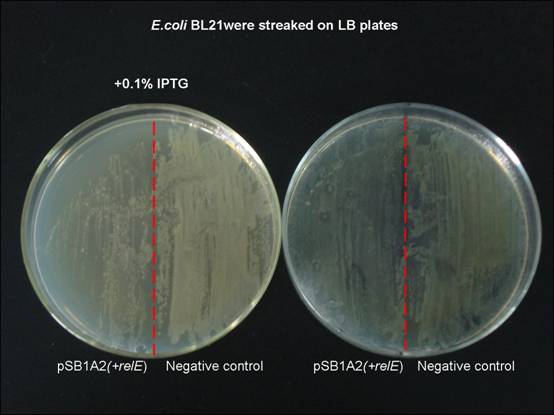
| + | As a result, expression of the '''''relE''''' gene has been shown to severely inhibit translation and prevent colony formation. | ||
| + | |||
| + | [[Image:SJTU09_Project_Overview1.jpg|center|thumb|554px|Fig. 1 expression of relE on growth of '''''E.coli''''' cells]] | ||
| + | |||
| + | However, the growth of '''''E.coli''''' can be restored by coexpression of the cognate relB antitoxin, which can form a heterotetrameric (relB-relE)<sub>2</sub> structure when binding with relE. Thus,this structure is too large to fit into the A site, so the toxic relE can be neutralized. Overexpression of relB restored protein synthesis and colony formation. From this followed that inhibition of protein synthesis by relE is bacteriostatic, a state of low metabolic rate and cell growth like the hibernation behavior of bears and hedgehogs in chilly winter, rather than bacteriocidal. But the unbound relB is an easy prey for the ATP-dependent Lon protease. | ||
| + | |||
| + | What is most intricating is that relE display codon-specific cleavage of mRNAs in the ribosomal A site, that is to say, among stop codons UAG is cleaved with fast, UAA intermediate and UGA slow rate(UAG>UAA>UGA). The rates of the inhibition reactions in all cases were proportional to the relE concentration and thus determined by k<sub>cat</sub>/K<sub>m</sub>. the k<sub>cat</sub>/K<sub>m</sub> Values (s<sup>-1</sup>μM<sup>-1</sup>) for RelE Cleavage of 3 stop codons in the Ribosomal A Site have been measured previously.(UGA:0.078, UAA:2.2, UAG:26). The physical rationale of the measurement is that k<sub>cat</sub>/K<sub>m </sub> is determined by the standard free energy difference between relE in activated complex with a ribosome and a ground state with free relE and ribosome. | ||
| + | |||
| + | In our project, this unique '''''relBE''''' operon is separated into two independent genes, that is , '''''relE''''' and '''''relB''''' are located on two plasmid vectors pSB1A2 and pSB1AK3 respectively. Thus, expression of '''''relE''''' can be induced by substrate IPTG and that of '''''relB''''' by Trp. And the ATP-dependent protease '''''Lon''''' gene follows '''''relE''''' to form a polycistron in the same operon, which means Lon can coexpress with relE once induced by IPTG. When two operons are induced simultaneously, in the protein level, relE, relB, and Lon will fight with each other. On the one hand, relB can prevent cell from dormancy by neutralizing relE while on the other hand, relB can easily degraded by Lon. It seems that with the coexpression of Lon, the toxic effect of relE can not be offset by relB. Thus it is a finality that '''''E.Coli''''' will hibernate to survive this stage. | ||
| + | |||
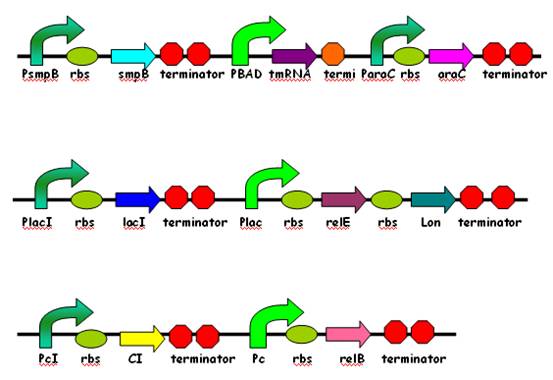
| + | [[Image:SJTU09_Project_Network1.jpg|center|thumb|553px|Fig. 2 Gene network of our project]] | ||
| + | |||
| + | To add suspense to this play, we decide to create a transcriptional regulation pathway to control the rate of synthesis of 3 proteins. We modify 3 genes’ stop codons artificially via primer design in PCR procedure. In detail, we modify the stop codons of '''''relE''''', '''''Lon''''' and '''''relB''''' to UAA, UAG, UGA respectively, which means on the transcriptional level, the mRNA of '''''Lon''''' will be the most severe victim of '''''relE''''' cleavage whereas the mRNA of '''''relB''''' can almost be intact based on a pretty low '''''relE''''' cleavage rate. relE cuts its own mRNA at a medium rate. On this level of competiton, mRNA of '''''relB''''' is the biggest winner, which grants an accordingly high level of relB product via translation. As relB accumulate to an amount surpass that of relE, the physiological state can alter from hibernation to activity, where the metabolism of bacteria are normal and colonies can easily form. | ||
| + | |||
| + | If we put all these stuffs together, who can tell the final champion? relE or relB? At first stage, Once induced by IPTG and arabinose simultaneously, relE, Lon are coexpressed, and relB is hydrolyzed rapidly by Lon once produced. relE is dominant in this phase, the bacteria may keep in low metabolic rate and are in weak capacity to form colonies. | ||
| + | |||
| + | Nevertheless, as relE cleave the 3 kind of mRNAs with different stop codons, relB has the fastest synthesis rate because of a high level of intact mRNAs. In contrast, Lon are cornered to a rather low synthesis rate based on a small amount of available mRNAs. This is a perfect moment for relB to accumulate its products. When relB expression is over relE, the toxic effect of relE can be halted completely. During this stage, the microbe can regain the capacity of colony formation. It’s time to break the nightmare of Hypnos’ Curse!!! | ||
| + | |||
| + | Although some previous job has certify the fact that later induction of relB transcription in RelE-inhibited cells mediated complete recovery of colony formation. | ||
| + | |||
| + | Some may doubt that in the whole '''''E.Coli''''' system there are lots of key housekeeping genes such as those of RNA polymerase, peptide release factors and some catalytic enzymes in respiration, so how can bacteria recover from dormancy without these genes’ proper work, which are also affected by relE toxicity? One requirement for recovery is that ribosomal complexes with peptidyl-tRNA in P site and an mRNA truncated in the A site codon can be brought back initiation. So tmRNA is required for rapid recovery of translation in RelE-inhibited cells. tmRNA is a versatile and highly conserved bacterial molecule endowed with the combined structural and functional properties of both a tRNA and a mRNA. The tmRNA system orchestrates three key biological functions: (1) recognition and rescue of ribosomes stalled on aberrant mRNAs, (2) disposal of the causative defective mRNAs, and (3)addition of a degradation tag to ribosome-associated protein fragments for directed proteolysis. | ||
| + | |||
| + | So tmRNA is also taken into our circuit to assist relB rescue the dormant cells. It does accelerate the rescue efficiency of relB. Overproduction of tmRNA efficiently counteracted relE toxicity. | ||
| + | |||
| + | Thus in our circuit, we create a oscillator in '''''E.coli''''', which function like a biological clock monitoring circadian rhythm in higher animals and plants. As the bacteria alternate their states between dormancy and resuscitation, there seems an inner clock within them. | ||
| + | |||
| + | The duration of a circle from dormancy to resuscitation is expected to be constant, so it may be the definition of 1 day in “'''''E.coli''''' world”. It sounds crazy that we can count the age of '''''E.Coli''''' according to the “days” they have during their life time!! Here we come into another interesting topic-----the life-span of E.coli. With a low metabolic rate in dormancy, the bacteria are much like higher animals who hibernate during winter. As we all know, bears and hedgehogs hibernate to keep a rather low metabolic rate, helping them go through food shortage in winter. So do tortoises, who maintain low metabolic rate to survive a longer life-span. Can our '''''E.Coli''''' with a inner clock extend their life-span via this circadian rhythm? | ||
{{Template:SJTU09_enddiv}} | {{Template:SJTU09_enddiv}} | ||
Revision as of 02:49, 17 October 2009

Project introduction. Inspired by the natural regulator of circadian bioclock exhibited in most eukaryotic organisms, our team has designed an E.coli-based genetic network with the toxin-antitoxin system so that the bacterium oscillates between two states of dormancy and activity (more...)
Background
It is universally acknowledged that bioclock works as a circadian regulator in most eukaryotic multicellular species. This mechanism controls higher plants’ blossom time, brings insects into metamorphosis, and also wakes us up every day.
Then comes up the crazy idea: Why cannot prokaryotes live with a bioclock?
Hence, we constructed our bacteria bioclock by utilizing the toxin-antitoxin system (TA system), which forms an oscillator between two physiological states--dormancy and activity.
The RelE toxin protein is an RNase that preferentially cleaves mRNAs bound to the ribosome at the second position of stop codons, and the order of its cleavage rates on different stop codons is UAG > UAA > UGA. Expression of the RelE gene has been shown to severely inhibit translation and prevent colony formation, whereas expression of the RelB antitoxin reverses these inhibitory effects. The blocked ribosome after RelE-mediated cleavage of the mRNA codon at the A site becomes a substrate for the tmRNA rescue system, which can degrade aberrant proteins made from truncated mRNAs and recycle the stalled ribosomes.
Based on these mechanisms we have designed an ingenious genetic network which functions as a bacterial bioclock oscillating between the two states of dormancy and activity. It is exciting to imagine that we manage to manipulate the lifespan of E.coli by switching the oscillator on, since the metabolic process of microbes is vastly decelerated during the dormancy state, just like bears and hedgehogs in their hibernation.
An example of how this artificial bioclock could be applied might be the preservation of scientifically valuable bacteria which mutate frequently. During the dormancy state bacteria hardly undergo mutation; therefore their genetic characteristics are retained.
Other potential applications such as biologic timing and antibiotic resistance remain intriguing to explore.
Principle & Network
E.coli the napper
Bacteria and at least some archaea harbor ‘suicide systems’, the existence of which is a mystery to those who hear about them for the first time. These systems consist of a lethal toxin that can be neutralized by an antitoxin. The production of antitoxin, which is labile, must at least parallel that of the toxin, which is usually stable, to circumvent the induction of cell poison. To achieve this, both proteins are encoded within a single operon. The toxins seem to target specific, essential cellular processes. Perhaps the best characterized of these system is relBE system, the genes of which are present on the Escherichia coli chromosome. The relE toxin is an RNase that preferentially cleaves mRNAs bound to the ribosome at the second position of stop codons. Stop codons not only signal the end of the protein coding sequence but also serve as the binding site for release factors, which promote release of the nascent polypeptide and facilitate recycling of ribosomes for further rounds of translation. Thus truncated mRNA by cleavage of relE lacks appropriate termination signals, which causes the accumulation of stalled ribosomes and these mRNAs are unable to promote release factor binding, nascent polypeptide release, and ribosome recycling.
As a result, expression of the relE gene has been shown to severely inhibit translation and prevent colony formation.
However, the growth of E.coli can be restored by coexpression of the cognate relB antitoxin, which can form a heterotetrameric (relB-relE)2 structure when binding with relE. Thus,this structure is too large to fit into the A site, so the toxic relE can be neutralized. Overexpression of relB restored protein synthesis and colony formation. From this followed that inhibition of protein synthesis by relE is bacteriostatic, a state of low metabolic rate and cell growth like the hibernation behavior of bears and hedgehogs in chilly winter, rather than bacteriocidal. But the unbound relB is an easy prey for the ATP-dependent Lon protease.
What is most intricating is that relE display codon-specific cleavage of mRNAs in the ribosomal A site, that is to say, among stop codons UAG is cleaved with fast, UAA intermediate and UGA slow rate(UAG>UAA>UGA). The rates of the inhibition reactions in all cases were proportional to the relE concentration and thus determined by kcat/Km. the kcat/Km Values (s-1μM-1) for RelE Cleavage of 3 stop codons in the Ribosomal A Site have been measured previously.(UGA:0.078, UAA:2.2, UAG:26). The physical rationale of the measurement is that kcat/Km is determined by the standard free energy difference between relE in activated complex with a ribosome and a ground state with free relE and ribosome.
In our project, this unique relBE operon is separated into two independent genes, that is , relE and relB are located on two plasmid vectors pSB1A2 and pSB1AK3 respectively. Thus, expression of relE can be induced by substrate IPTG and that of relB by Trp. And the ATP-dependent protease Lon gene follows relE to form a polycistron in the same operon, which means Lon can coexpress with relE once induced by IPTG. When two operons are induced simultaneously, in the protein level, relE, relB, and Lon will fight with each other. On the one hand, relB can prevent cell from dormancy by neutralizing relE while on the other hand, relB can easily degraded by Lon. It seems that with the coexpression of Lon, the toxic effect of relE can not be offset by relB. Thus it is a finality that E.Coli will hibernate to survive this stage.
To add suspense to this play, we decide to create a transcriptional regulation pathway to control the rate of synthesis of 3 proteins. We modify 3 genes’ stop codons artificially via primer design in PCR procedure. In detail, we modify the stop codons of relE, Lon and relB to UAA, UAG, UGA respectively, which means on the transcriptional level, the mRNA of Lon will be the most severe victim of relE cleavage whereas the mRNA of relB can almost be intact based on a pretty low relE cleavage rate. relE cuts its own mRNA at a medium rate. On this level of competiton, mRNA of relB is the biggest winner, which grants an accordingly high level of relB product via translation. As relB accumulate to an amount surpass that of relE, the physiological state can alter from hibernation to activity, where the metabolism of bacteria are normal and colonies can easily form.
If we put all these stuffs together, who can tell the final champion? relE or relB? At first stage, Once induced by IPTG and arabinose simultaneously, relE, Lon are coexpressed, and relB is hydrolyzed rapidly by Lon once produced. relE is dominant in this phase, the bacteria may keep in low metabolic rate and are in weak capacity to form colonies.
Nevertheless, as relE cleave the 3 kind of mRNAs with different stop codons, relB has the fastest synthesis rate because of a high level of intact mRNAs. In contrast, Lon are cornered to a rather low synthesis rate based on a small amount of available mRNAs. This is a perfect moment for relB to accumulate its products. When relB expression is over relE, the toxic effect of relE can be halted completely. During this stage, the microbe can regain the capacity of colony formation. It’s time to break the nightmare of Hypnos’ Curse!!!
Although some previous job has certify the fact that later induction of relB transcription in RelE-inhibited cells mediated complete recovery of colony formation.
Some may doubt that in the whole E.Coli system there are lots of key housekeeping genes such as those of RNA polymerase, peptide release factors and some catalytic enzymes in respiration, so how can bacteria recover from dormancy without these genes’ proper work, which are also affected by relE toxicity? One requirement for recovery is that ribosomal complexes with peptidyl-tRNA in P site and an mRNA truncated in the A site codon can be brought back initiation. So tmRNA is required for rapid recovery of translation in RelE-inhibited cells. tmRNA is a versatile and highly conserved bacterial molecule endowed with the combined structural and functional properties of both a tRNA and a mRNA. The tmRNA system orchestrates three key biological functions: (1) recognition and rescue of ribosomes stalled on aberrant mRNAs, (2) disposal of the causative defective mRNAs, and (3)addition of a degradation tag to ribosome-associated protein fragments for directed proteolysis.
So tmRNA is also taken into our circuit to assist relB rescue the dormant cells. It does accelerate the rescue efficiency of relB. Overproduction of tmRNA efficiently counteracted relE toxicity.
Thus in our circuit, we create a oscillator in E.coli, which function like a biological clock monitoring circadian rhythm in higher animals and plants. As the bacteria alternate their states between dormancy and resuscitation, there seems an inner clock within them.
The duration of a circle from dormancy to resuscitation is expected to be constant, so it may be the definition of 1 day in “E.coli world”. It sounds crazy that we can count the age of E.Coli according to the “days” they have during their life time!! Here we come into another interesting topic-----the life-span of E.coli. With a low metabolic rate in dormancy, the bacteria are much like higher animals who hibernate during winter. As we all know, bears and hedgehogs hibernate to keep a rather low metabolic rate, helping them go through food shortage in winter. So do tortoises, who maintain low metabolic rate to survive a longer life-span. Can our E.Coli with a inner clock extend their life-span via this circadian rhythm?
 "
"