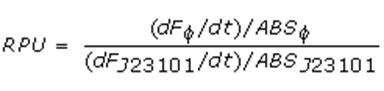Team:UNIPV-Pavia/Parts Characterization
From 2009.igem.org
(→Relative Promoter Units (RPUs)) |
|||
| Line 116: | Line 116: | ||
*it uses a reference standard and so measurements can be compared between different laboratories. | *it uses a reference standard and so measurements can be compared between different laboratories. | ||
| - | The hypotheses on which RPU theory is based can be found in Kelly J. et al., 2008, as well as all the mathematical steps. From our point of view, the main hypotheses that have to be satisfied are the following: | + | The hypotheses on which RPU theory is based can be found in [[Team:UNIPV-Pavia/Project/References#18| Kelly J. et al., 2008]], as well as all the mathematical steps. From our point of view, the main hypotheses that have to be satisfied are the following: |
*the reporter protein must have a half life higher than the experiment duration (we use GFPmut3, <partinfo>BBa_E0240</partinfo>, which has an estimated half life of at least 24 hours, and the experiments duration is always less than 7 hours); | *the reporter protein must have a half life higher than the experiment duration (we use GFPmut3, <partinfo>BBa_E0240</partinfo>, which has an estimated half life of at least 24 hours, and the experiments duration is always less than 7 hours); | ||
*strain, plasmid copy number, antibiotic, growth medium, growth conditions, protein generator assembled downstream of the promoter must be the same in the promoter of interest and in J23101 reference standard. | *strain, plasmid copy number, antibiotic, growth medium, growth conditions, protein generator assembled downstream of the promoter must be the same in the promoter of interest and in J23101 reference standard. | ||
Revision as of 17:40, 21 October 2009

|
|
|
Parts Characterization |
||
|
Here we describe the characterization results of 4 parts of our own design, 2 existing parts re-built because they were inconsistent and 7 existing parts taken from the Registry. When not reported differently, all the experiments have been performed according to Growth conditions and Data analysis sections.
Re-built existing parts (BBa_our part code/BBa_existing part code): Existing parts from the Registry:
Existing parts - sequence debugging:
Growth conditionsMicroplate reader experiments
Fermentation experimentsPROTOCOL#1
PROTOCOL#2
PROTOCOL#3
Data analysisGrowth curvesAll our growth curves have been obtained subtracting for each time sample the broth O.D.600 measurement from that of the culture; broth was considered in the same conditions of the culture (e.g. induced with the same inducer concentration). Doubling timeThe natural logarithm of the growth curves (processed according to the above section) was computed and the linear phase (corresponding to the bacterial exponential growth phase) was isolated by visual inspection. Then the linear regression was performed in order to estimate the slope of the line m. Finally the doubling time was estimated as d=ln(2)/m [minutes]. In the case of multiple growth curves for a strain, the mean value of the processed curves was computed for each time sample before applying the above described procedure. Relative Promoter Units (RPUs)The RPUs are standard units proposed by Kelly J. et al., 2008, in which the transcriptional strength of a promoter can be measured using a reference standard, just like the ground in electric circuits. RPUs have been computed as: in which:
RPU measurement has the following advantages (under suitable conditions)
The hypotheses on which RPU theory is based can be found in Kelly J. et al., 2008, as well as all the mathematical steps. From our point of view, the main hypotheses that have to be satisfied are the following:
Inducible systemsEvery experiment is performed on the following cultures:
For inducible systems several plots are reported. The first plot is a panel containing 4 subplots, numerated this way:
Plot (1) contains growth curves of the cultures, after blank value has been removed. Every curve is calculated averaging on three replicates of the same culture and subtracting the blank for each time sample. Blank is calculated averaging the replicates of blank wells. Plot (2) shows the logarithm of absorbance in exponential phase of bacterial growth, determined by a visual inspection of log-plots. These values are used to evaluate doubling time and R.P.U.. Plot (3) contains (dGFP/dt)/O.D., the value named Scell in Canton procedure for RPU evaluation. In these plots are reported black veritcal lines that define the range of values used to evaluate RPU. It is important to underline, as explained in next paragraph, that RPU are calculated on cultures at the same O.D. level, not at the same time. The second graphic shows Scell VS O.D.. This plot allows the conparison of Scell values between different cultures, that are supposed to reach the same level of growth not at the same time, but at the same O.D. value. The third graphic shows the induction curve. The RPU value is calculated on Scell values corresponding to O.D. values in exponential phase (tpitcally, from 0.05 to 0.16). The curve is obtained averaging in time Scell values corresponding to exponential phase. Error bars rapresent the minimum and maximum value of R.P.U. belonging to the range of O.D. in exponential phase. In RPU evaluation the hypothesis of steady state has to be validated. This hypothesis corresponds to a constant behavior of Scell in time. In exponential phase in several cases it is possible to observe that this variable isn't constant, but grows after exponential phase is over. This behaviour is totally unexpected and can't be justifyed by any biological argument. It is also important to undelrine that the reported methodology has shown how variable R.P.U. value can be. This parameter, in fact, is very sensitive to the respondind O.D. value, as shown from induction curves, where error bars are sometimes wide among the curve. So it is foundamental to define a standardized methodology for RPU evaluation, not sensitive to O.D. or time choose. Materials
|
 "
"