Team:Cambridge/Notebook/Week3
From 2009.igem.org
(→Monday) |
(→Monday) |
||
| Line 7: | Line 7: | ||
==Monday== | ==Monday== | ||
| - | + | ===Primer Design=== | |
Primers were designed to convert MelA into a biobrick. Primers A and F are for either end (including prefix and suffix - highlighted in green). The other primers surround the unwanted restiction sites (highlighted in yellow) with the changed base in red. | Primers were designed to convert MelA into a biobrick. Primers A and F are for either end (including prefix and suffix - highlighted in green). The other primers surround the unwanted restiction sites (highlighted in yellow) with the changed base in red. | ||
| Line 17: | Line 17: | ||
[[Image:MelA_primer_map.JPG]] | [[Image:MelA_primer_map.JPG]] | ||
| - | + | ===Orange Pigment=== | |
Used plasmid editor to examine the genes used for the production of orange pigments. The (supposedly) orange-producing pigment from the biobricks: | Used plasmid editor to examine the genes used for the production of orange pigments. The (supposedly) orange-producing pigment from the biobricks: | ||
| Line 27: | Line 27: | ||
[[Image:EBI.JPG]] | [[Image:EBI.JPG]] | ||
| - | + | ===Transformed Pigments=== | |
The MelA plate left on the bench overnight produced a brown-coloured pigment! Growth on media containing IPTG should produce this pigment a lot faster. | The MelA plate left on the bench overnight produced a brown-coloured pigment! Growth on media containing IPTG should produce this pigment a lot faster. | ||
| - | + | ===Amplification system=== | |
:*Possible plan | :*Possible plan | ||
| Line 42: | Line 42: | ||
:*First we need to dig up the 2007 work and recreate their data, this time with a plate reader that can take OD and fluorescence readings simultaneously. | :*First we need to dig up the 2007 work and recreate their data, this time with a plate reader that can take OD and fluorescence readings simultaneously. | ||
| - | + | ====Activators==== | |
:*Found parts submitted by Cambridge '07 team amplifier project. Three translational units (ribosome binding sites and protein coding sequence) for the three activators. Sequence below, 5' to 3': | :*Found parts submitted by Cambridge '07 team amplifier project. Three translational units (ribosome binding sites and protein coding sequence) for the three activators. Sequence below, 5' to 3': | ||
| Line 70: | Line 70: | ||
(delete sequence if you think its unnecessary and just leave part names) | (delete sequence if you think its unnecessary and just leave part names) | ||
| - | + | ====Promoters==== | |
Available as individual parts, with a GFP reporter or as part of a composite with the activator upstream. | Available as individual parts, with a GFP reporter or as part of a composite with the activator upstream. | ||
| Line 81: | Line 81: | ||
PLL promoter, part: I746365 | PLL promoter, part: I746365 | ||
| - | + | ====Degradation Tags==== | |
:*three C-terminal tags are well described in Andersen, J. B., Sternberg, C., Poulsen, L. K., Bjorn, S. P., Givskov, M. & Molin, S. (1998). New unstable variants of green fluorescent protein for studies of transient gene expression in bacteria. Appl Environ Microbiol 64, 2240–2246.[http://aem.asm.org/cgi/reprint/64/6/2240], which detailed tagged-GFP degredation. | :*three C-terminal tags are well described in Andersen, J. B., Sternberg, C., Poulsen, L. K., Bjorn, S. P., Givskov, M. & Molin, S. (1998). New unstable variants of green fluorescent protein for studies of transient gene expression in bacteria. Appl Environ Microbiol 64, 2240–2246.[http://aem.asm.org/cgi/reprint/64/6/2240], which detailed tagged-GFP degredation. | ||
:*the tags as described in the paper that we could use are as follows (all DNA sequences are 5' to 3') | :*the tags as described in the paper that we could use are as follows (all DNA sequences are 5' to 3') | ||
Revision as of 20:10, 27 July 2009
Categories :
Project :
-
Overview
Sensitivity Tuner
--- Characterisation
--- Modelling
Colour Generators
--- Carotenoids (Orange/Red)
--- Melanin (Brown)
--- Violacein (Purple/Green)
The Future
Safety
Notebook :
Team Logistics :
Week 3 - Development
Monday
Primer Design
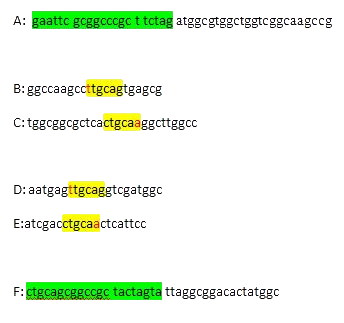
Primers were designed to convert MelA into a biobrick. Primers A and F are for either end (including prefix and suffix - highlighted in green). The other primers surround the unwanted restiction sites (highlighted in yellow) with the changed base in red.
These correspond to the parts on the gene shown below. The stars correspond to the two unwanted restriction sites.
Orange Pigment
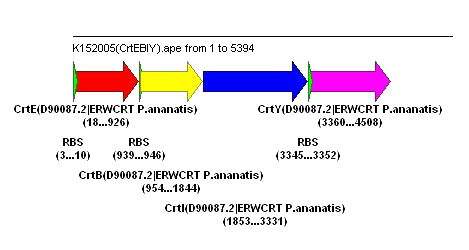
Used plasmid editor to examine the genes used for the production of orange pigments. The (supposedly) orange-producing pigment from the biobricks:
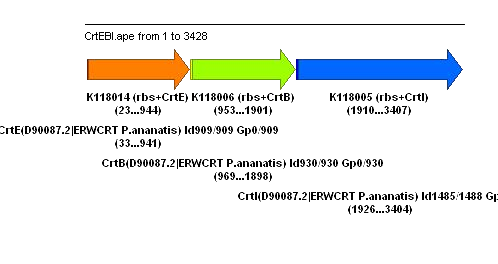
We also tried a preliminary design for a biobrick we could construct, in case the orange biobrick does not function. This uses the same genetic synthetic pathway for gene consruction:
Transformed Pigments
The MelA plate left on the bench overnight produced a brown-coloured pigment! Growth on media containing IPTG should produce this pigment a lot faster.
Amplification system
- Possible plan
- We would want to tag the activator proteins then quantify their degradation in some way
- GFP fusion: one idea was to create a functional activator-GFP fusion, to which we would then add a C-terminal degradation tag. Thus we could express the activator-GFP-tag in E. coli, and could measure flourescence to quantify activator degradation directly. However, this idea depends on us constructing a functional activator-GFP fusion.
- Western blotting: a second idea is to add an N-terminal flag tag to the activator proteins, and then add the C-terminal degradation tag. We could measure the amount of activator using Western blotting and an anti-flag antibody by taking samples over time. However, Western blotting isn't that easy.
- Finally, what we're really doing is solving a problem found in the Cambridge 2007 team's system. If we can show that overnight bacterial growth is less hindered with our tagged activators, then we are effectively showing that because of the degradation tags, the activators are indeed being degraded.
- We could also attempt to see what this effect has on amplification
- Modelling potential - activator half life, change in Pops out?
- First we need to dig up the 2007 work and recreate their data, this time with a plate reader that can take OD and fluorescence readings simultaneously.
Activators
- Found parts submitted by Cambridge '07 team amplifier project. Three translational units (ribosome binding sites and protein coding sequence) for the three activators. Sequence below, 5' to 3':
- Ogr activator from P2 phage: Part I746350
>BBa_I746350 Part-only sequence (237 bp)
aaagaggagaaatactagatgtttcattgtcctttatgccagcatgccgcacatgcgcgtacaagtcgctatatcactgacacgacaaaagagcgttatc
atcagtgccagaacgtgaattgcagcgccacgttcatcacttatgagtcggtacagcgatacatcgtgaagccgggagaagtccacgccgtaaggccgca
cccgttgccatcagggcagcaaattatgtggatgtaa
- pag activator from PSP3 phage: Part I746351
>BBa_I746351 Part-only sequence (237 bp) aaagaggagaaatactagatgatgcactgcccgttatgccaaaacgctgcacatgctcgcactagccggtaccttagcaccgaaacgaaagaacgttatc accagtgccaaaacataaattgcggatgtacatttatcacttttgagacactatcaagattcattgtgaaaccggggactgttgatcctgctccgcccca ccccatcagaaaccaacaacagcaactttggctttga
- delta activator from phiR73 phage: Part I746352
>BBa_I746352 Part-only sequence (264 bp) aaagaggagaaatactagatgatgcgctgccctttctgtcgtcattcagcgcatacccgcaccagccggtatgtgagtgacaatgtcaaagaaagttatc tccagtgccagaatatttactgttcggcgacatttaaaacgcatgagtcaatttgtgccgtgattcgttctccggtcacggaggaaaaaccagcaccggc aagcacagcaccggctgttgtccgaaaagttaaaggctgttacagctcaccattcaaccattaa
(delete sequence if you think its unnecessary and just leave part names)
Promoters
Available as individual parts, with a GFP reporter or as part of a composite with the activator upstream. Five promoters as used by Cambridge '07:
PF promoter, part: I746360 PO promoter, part: I746361 PP promoter, part: I746362 Psid promoter, part: I746364 PLL promoter, part: I746365
Degradation Tags
- three C-terminal tags are well described in Andersen, J. B., Sternberg, C., Poulsen, L. K., Bjorn, S. P., Givskov, M. & Molin, S. (1998). New unstable variants of green fluorescent protein for studies of transient gene expression in bacteria. Appl Environ Microbiol 64, 2240–2246.[http://aem.asm.org/cgi/reprint/64/6/2240], which detailed tagged-GFP degredation.
- the tags as described in the paper that we could use are as follows (all DNA sequences are 5' to 3')
- LAA: amino acid sequence AANDENYALAA, DNA sequence GCAGCAAACGACGAAAACTACGCTTTAGCAGCT
(primer – CGTCGTTTGCTGCTTTTGATGCGAAATCGTCGA) (currently the "planning" biobrick BBa_M0042)
- AAV: amino acid sequence AANDENYAAAV, DNA sequence GCAGCAAACGACGAAAACTACGCTGCAGCAGTT
(primer – CGTCGTTTGCTGCTTTTGATGCGACGTCGTCAA, currently the "planning" biobrick BBa_M0044 with DNA sequence GCAGCAAACGACGAAAACTACGCTGCTGCTGTT)
- NOTE: we need to use the biobrick DNA sequence as the DNA sequence described in the paper has a forbidden PstI site
- ASV: amino acid sequence AANDENYAASV, DNA sequence GCAGCAAACGACGAAAACTACGCTGCATCAGTT
(primer – CGTCGTTTGCTGCTTTTGATGCGACGTAGTCAA, currently the "planning" biobrick BBa_M0046 with DNA sequence GCTGCAAACGACGAAAACTACGCTGCATCAGTT)
 "
"