Team:Cambridge/Notebook/Week3
From 2009.igem.org
MikeDavies (Talk | contribs) (→Activators) |
MikeDavies (Talk | contribs) (→Activators) |
||
| Line 45: | Line 45: | ||
*Found parts submitted by Cambridge '07 team amplifier project. Three translational units (ribosome binding sites and protein coding sequence) for the three activators. Sequence below, 5' to 3': | *Found parts submitted by Cambridge '07 team amplifier project. Three translational units (ribosome binding sites and protein coding sequence) for the three activators. Sequence below, 5' to 3': | ||
| - | *'''Ogr activator from P2 phage: Part I746350''' | + | :*'''Ogr activator from P2 phage: Part I746350''' |
| - | + | ||
[http://partsregistry.org/Part:BBa_I746350 BBa_I746350 Part-only sequence (237 bp)] | [http://partsregistry.org/Part:BBa_I746350 BBa_I746350 Part-only sequence (237 bp)] | ||
| - | *'''pag activator from PSP3 phage: Part I746351''' | + | :*'''pag activator from PSP3 phage: Part I746351''' |
| - | + | ||
[http://partsregistry.org/Part:BBa_I746351 BBa_I746351 Part-only sequence (237 bp)] | [http://partsregistry.org/Part:BBa_I746351 BBa_I746351 Part-only sequence (237 bp)] | ||
| - | *'''delta activator from phiR73 phage: Part I746352''' | + | :*'''delta activator from phiR73 phage: Part I746352''' |
| - | + | ||
[http://partsregistry.org/Part:BBa_I746352 BBa_I746352 Part-only sequence (264 bp)] | [http://partsregistry.org/Part:BBa_I746352 BBa_I746352 Part-only sequence (264 bp)] | ||
Revision as of 19:42, 3 August 2009
Categories :
Project :
-
Overview
Sensitivity Tuner
--- Characterisation
--- Modelling
Colour Generators
--- Carotenoids (Orange/Red)
--- Melanin (Brown)
--- Violacein (Purple/Green)
The Future
Safety
Notebook :
Team Logistics :
Week 3 - Development
Monday
Primer Design
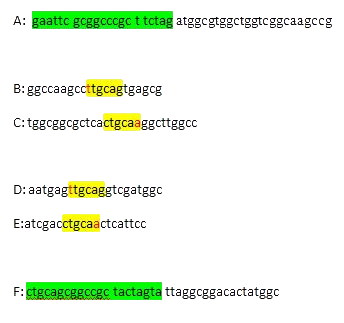
Primers were designed to convert MelA into a biobrick. Primers A and F are for either end (including prefix and suffix - highlighted in green). The other primers surround the unwanted restiction sites (highlighted in yellow) with the changed base in red.
These correspond to the parts on the gene shown below. The stars correspond to the two unwanted restriction sites.
Orange Pigment
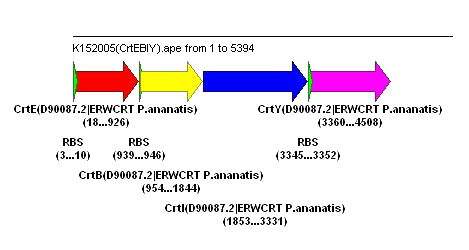
Used plasmid editor to examine the genes used for the production of orange pigments. The (supposedly) orange-producing pigment from the biobricks:
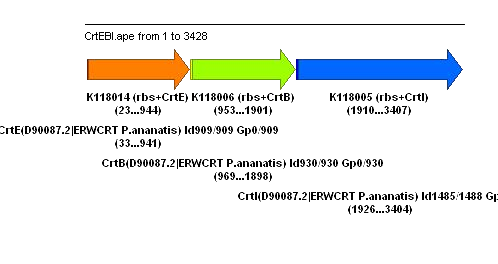
We also tried a preliminary design for a biobrick we could construct, in case the orange biobrick does not function. This uses the same genetic synthetic pathway for gene consruction:
Transformed Pigments
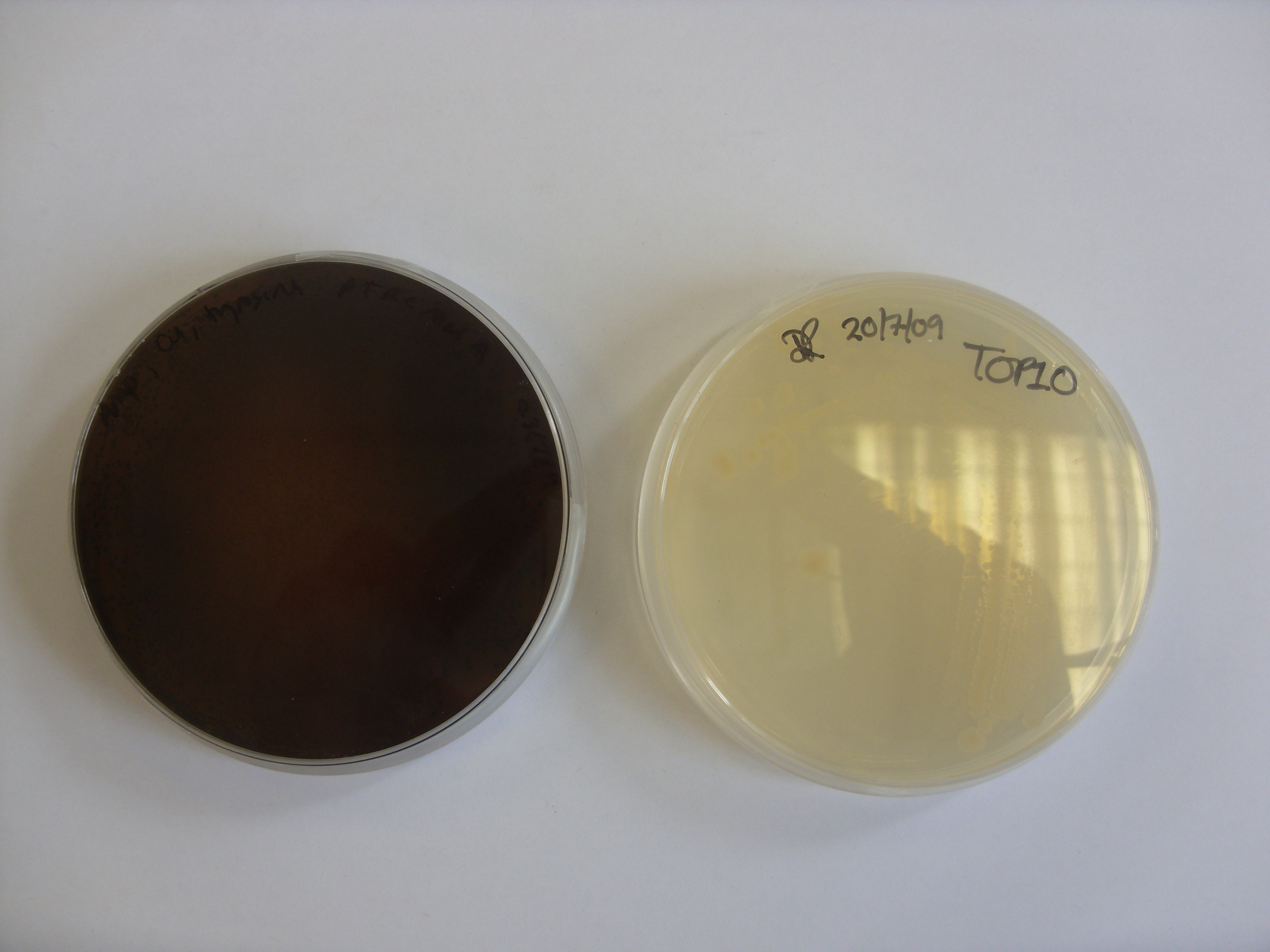
The MelA plate left on the bench overnight produced a brown-coloured pigment! Growth on media containing IPTG should produce this pigment a lot faster.
Amplification system
- Possible plan
- We would want to tag the activator proteins then quantify their degradation in some way
- GFP fusion: one idea was to create a functional activator-GFP fusion, to which we would then add a C-terminal degradation tag. Thus we could express the activator-GFP-tag in E. coli, and could measure flourescence to quantify activator degradation directly. However, this idea depends on us constructing a functional activator-GFP fusion.
- Western blotting: a second idea is to add an N-terminal flag tag to the activator proteins, and then add the C-terminal degradation tag. We could measure the amount of activator using Western blotting and an anti-flag antibody by taking samples over time. However, Western blotting isn't that easy.
- Finally, what we're really doing is solving a problem found in the Cambridge 2007 team's system. If we can show that overnight bacterial growth is less hindered with our tagged activators, then we are effectively showing that because of the degradation tags, the activators are indeed being degraded.
- We could also attempt to see what this effect has on amplification
- Modelling potential - activator half life, change in Pops out?
- First we need to dig up the 2007 work and recreate their data, this time with a plate reader that can take OD and fluorescence readings simultaneously.
Activators
- Found parts submitted by Cambridge '07 team amplifier project. Three translational units (ribosome binding sites and protein coding sequence) for the three activators. Sequence below, 5' to 3':
- Ogr activator from P2 phage: Part I746350
[http://partsregistry.org/Part:BBa_I746350 BBa_I746350 Part-only sequence (237 bp)]
- pag activator from PSP3 phage: Part I746351
[http://partsregistry.org/Part:BBa_I746351 BBa_I746351 Part-only sequence (237 bp)]
- delta activator from phiR73 phage: Part I746352
[http://partsregistry.org/Part:BBa_I746352 BBa_I746352 Part-only sequence (264 bp)]
Promoters
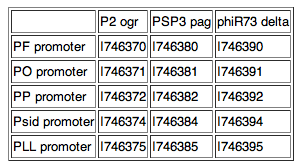
Available as individual parts, with a GFP reporter or as part of a composite with the activator upstream. Five promoters as used by Cambridge '07:
PF promoter, part: I746360 PO promoter, part: I746361 PP promoter, part: I746362 Psid promoter, part: I746364 PLL promoter, part: I746365
Degradation Tags
- Three C-terminal tags are well described in Andersen, J. B., Sternberg, C., Poulsen, L. K., Bjorn, S. P., Givskov, M. & Molin, S. (1998). New unstable variants of green fluorescent protein for studies of transient gene expression in bacteria. Appl Environ Microbiol 64, 2240–2246.[http://aem.asm.org/cgi/reprint/64/6/2240], which detailed tagged-GFP degredation.
- The tags as described in the paper that we could use are as follows (all DNA sequences are 5' to 3')
- LAA: Amino acid sequence AANDENYALAA, DNA sequence GCAGCAAACGACGAAAACTACGCTTTAGCAGCT
(primer – CGTCGTTTGCTGCTTTTGATGCGAAATCGTCGA) (currently the "planning" biobrick BBa_M0042)
- AAV: Amino acid sequence AANDENYAAAV, DNA sequence GCAGCAAACGACGAAAACTACGCTGCAGCAGTT
(primer – CGTCGTTTGCTGCTTTTGATGCGACGTCGTCAA, currently the "planning" biobrick BBa_M0044 with DNA sequence GCAGCAAACGACGAAAACTACGCTGCTGCTGTT)
- NOTE: We need to use the biobrick DNA sequence as the DNA sequence described in the paper has a forbidden PstI site
- ASV: Amino acid sequence AANDENYAASV, DNA sequence GCAGCAAACGACGAAAACTACGCTGCATCAGTT
(primer – CGTCGTTTGCTGCTTTTGATGCGACGTAGTCAA, currently the "planning" biobrick BBa_M0046 with DNA sequence GCTGCAAACGACGAAAACTACGCTGCATCAGTT)
Wet Work
- Made up 4 15ug/mL tetracycline agar plates - argh should have been trimethoprim!
- Electroporated the following plasmids into our TOP10 competent E. coli cells
- K118005
- K118006
- K118013
- K119014
- Orange plasmid from Dupont strain
- Plated each biobrick out onto ampicillin plates, plated the orange plasmid onto a tetracycline plate, left to incubate overnight
- Innoculated 6 single colonies from each of the following cultures, grown over the weekend on plates, in the appropriate antibiotic LB to shake over night in preparation for minipreps and colony PCRs tomorrow
- Violacein plasmid
- Melanin plasmid
- K152005
- R0011
Amplifiers
Basal amplification could potentially be reduced by using a suppressor. The "amplification factors" from Cambridge 2007 appear to be long term, ie steady state. It would be interesting to look at the transient response, as the degradation rate will certainly have an impact on this.
Tuesday
Long Term Aims
Pigments
- Create biobricks
Amplification
- Find biobricks
- Organise system
- Transform
- Test
- Attach to pigments (if time)
Degredation Tags
- Synthesise
- Test
- Attach
Colour Logic
- Get colour biobricks
- Attach inducible promoters
- Test
Results of Melanin transformation
Plate on the left is the melA plasmid plated on LB agar with 15ug/ml copper, 0.2ug/ul tyrosine and 100ug/ml Amp. The plate was incubated at 37 degrees for 24 hours and then left on the bench at room temperature over the weekend. Quicker induction of the melanin would be achieved by growing on plates containing IPTG. The relative concentrations of copper, tyrosine and IPTG could also be optimized. Of particular note is the fact that the pigment appears to have diffused out of the colonies.
The plate on the right is the control plate, containing untransformed TOP10 E. coli. These were the cells used for the transformation. We will also need to do a second control experiment to show that pigmentation is due to the bacteria and not due to auto-oxidation of any of the supplements we added to the media.
Wet Work
Colony PCR
- K152005
- R0011
PCR cycle = two minutes at 98 degrees followed by thirty cycles:
- Denaturation 98 degrees 30 seconds
- Annealing 65 degrees 30 seconds
- Elongation 22 degrees 3 minutes
Followed by a final elongation at 72 degrees for 10 minutes.
After the PCR run the products were run out on a gel (15ul of PCR product with 2ul of 1X loading buffer were added to a 0.8% agarose Invitrogen E-GEL and run for 30 mins).
Mini Prep
- K152005
- R0011 - not a high enough concentration of plasmid DNA, will have to cut out of gel from colony PCR
- Violacein cultures didn't grow overnight, will try again tomorrow with new cultures
- Melanin
- Used nanodrop spectrophotometer to measure DNA concentration.
Preparation for Colony PCRs and Minipreps for Tomorrow
- Overnight cultures of the cultures containing the following plasmids in LA with the appropriate antibiotic
- K118005 (amp)
- K118006 (amp)
- K118013 (amp)
- K118014 (amp)
- Violacein (trimethoprim)
New E. Coli Strains
- Roche parent strain and FPP high flux strain, for orange plasmid
- Prepared to make competent cells tomorrow
Amplification system plan
- Thursday
- Prepare to make arabinose strain (BW27783) competent
- Incubate single colony overnight
- Make more amp plates if necessary
- Friday
- Do competent cell protocol
- Transform all 15 combination constructs.
- Plate (30 amp plates)
- Saturday
- Hope they grew
- Sunday
- Incubate single colonies overnight in culture
- Monday
- Colony PCR of each
- PCR low copy plasmid
- Tuesday
- Use plate reader to gather data for the 15 combination constructs in BW27783 cells on high copy number plasmid after induction with arabinose
- Assembly: restriction digest of combination constructs, ligation into low copy plasmid
- Transform new constructs into Top10
- Plate (30 kan plates)
- Wednesday
- Colony PCR of new constructs the sneaky way
- Incubate single colonies overnight in culture
- Thursday
- Colony PCR of new constructs if yesterdays didn’t work
- Miniprep
- Transform into BW27783 strain
- Plate (30 kan plates)
- Friday
- Do another plate reader analysis after induction with arabinose
Wednesday
Dry Work
Organising a road-map for the procedures, to be presented at the meeting tomorrow
Wet Work
Miniprep
Violacein - got a decent DNA concentration this time.
Making more competant cells
We made the MG1655 Roche and Parent strains competant. The Roche strain has been modified from the parent strain in order to upregulate the pathways that produce the precursors for the orange pigment. We aim to transform both of them with the orange plasmid from Duncan and the orange pigmented biobrick in order to compare the difference in pigment intensity.
We also started the procedure to make strain BW27783 competent. This strain constitutively expresses arabinose transporters and is perhaps better suited to arabinose induction for the amplification project.
Testing DNA ladders
We ran a quick gel to test the ladders in the freezer from the iGEM competition last year (using the 0.8% agarose Bioline E-GELs). All hyperladders were confirmed to be working. It was decided to order more Hyperladder I as this gives the most useful spread of bands for the DNA we will be working with. This was also a test of the E-GEL system confirming the gels worked well.
Picture above shows the gel test run for the ladders. From left: hyperladder I, four samples of hyperladder IV found in the freezer. There were no meaningful results from the PCR run, so we went back to check over the proceedure for errors.
Transformations
We transformed Duncan's orange plasmid into Top10, Roche, and parent strain competent cells. Plated onto chloramphenicol and tetracycline plates, as we're not 100% sure of the antibiotic resistance properties of the plasmid.
Preparing culture
Single cultures of the melA transformed cells and TOP10 cells (for a control) were incubated at 37 degrees in 20 mL of agar overnight. The aim is to try to characterise the melanin pigment using the OD and flourescence plate-reader.
Thursday
Dry Work
Primers
Redesigned Primer A for the melA plasmid to include the native ribosome binding site:
Green is the suffix, purple is the ribosome binding site and the cyan is the start of the gene.
Also characterised the vio operon in terms of the seperate genes:
- vio A = 1256bp (two unwanted restriction sites)
- vio B = 2996bp (six unwanted restriction sites)
- vio C = 1289bp (two unwanted restriction sites)
- vio D = 1121bp (one unwanted restriction sites)
Road maps
Made road maps for melanin, violacein, pseudomonas, detailing the constructions and biobricks we plan to make all the way to making logic circuits.
Notes from 2:30 lab meeting
- Need to further integrate amplification and pigments into a "big idea"
- what is the rate limiting step in pigment formation - number of enzymes or substrate?
- Avenues to consider
- Strains of bacteria that detect concentrations of something and respond by turning a different colour - we could base that on the band pass project and use a system that produces 2 pigments with one pathway, for example violacein or pseudomonas
- for example, as in band pass, at low concentrations of AHL make cyan, have high concentrations of AHL turn on VioC and product violet instead
Wet Work
Results of Previous Colour Transformations
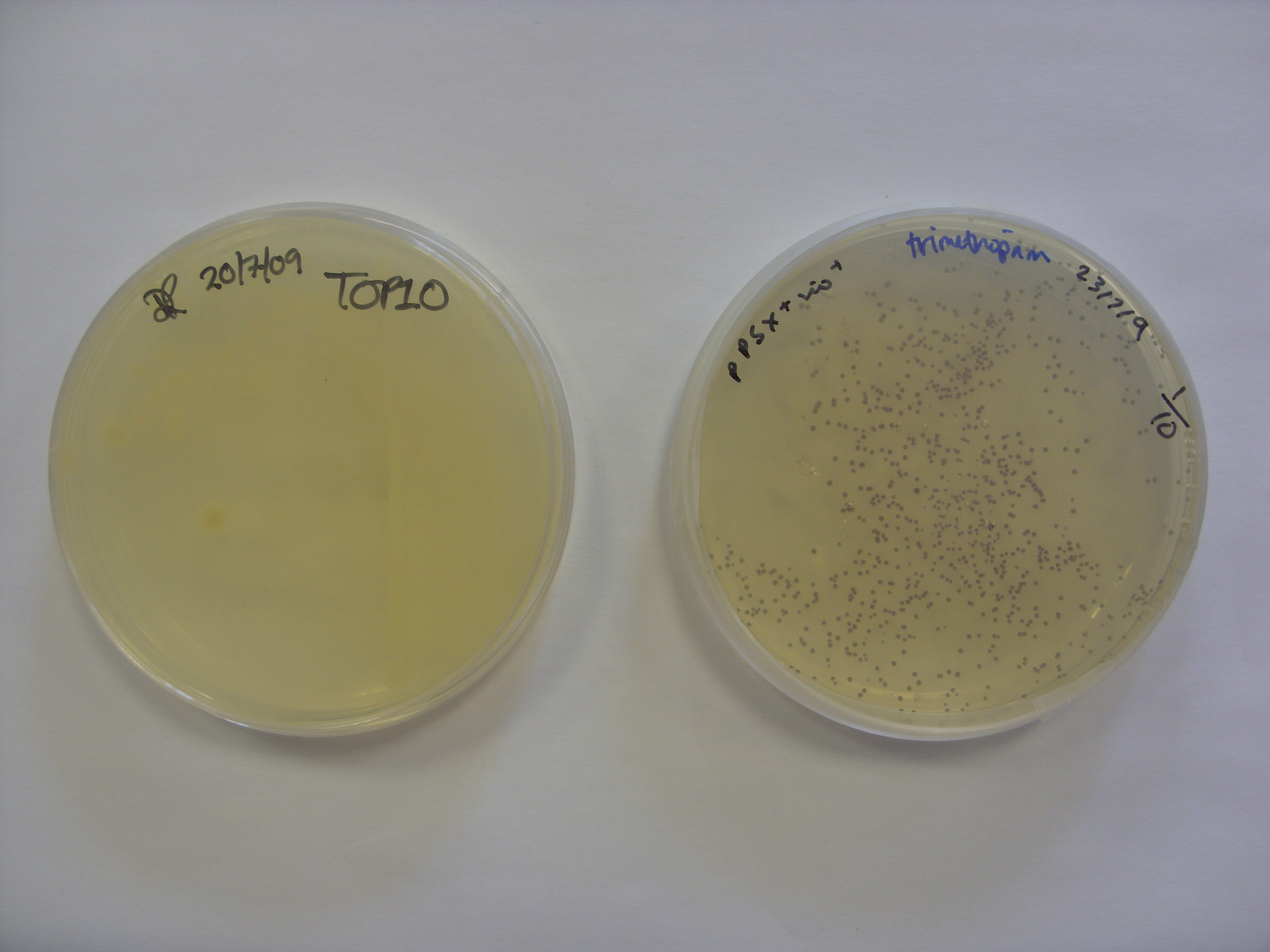
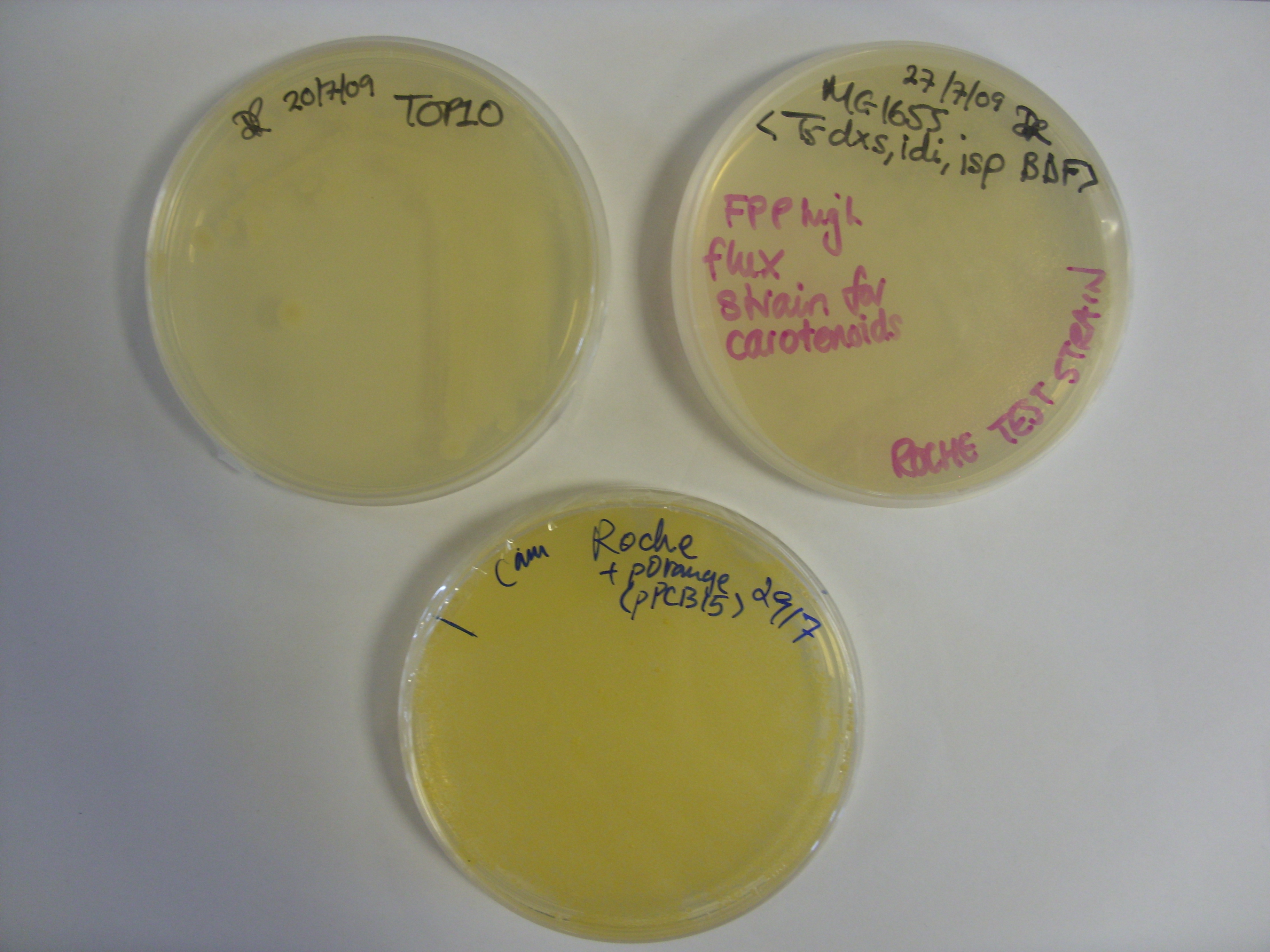
We took the violacein pigment bacteria (right plate in photo) out of the fridge to find that the purple colour had started to develop. They were therefore left at room temperature overnight. The colour appears to be within the bacteria, with little or no bleeding into the media. The control plate (left plate) is the untransformed TOP10 E. coli.
The Orange plasmid transformed into the Roche strain (with upregulated upstream pathways) had also produced colour. The two controls are above the trannsformed strain; the plate on the left shows TOP10 E. coli (i.e 'normal' E. coli colour) while the plate on the right shows the untransformed Roche strain.
Amplification system
- Made arabinose strain electrocompetent
Melanin
- Started using the plate reader to quantify melanin absorption.
Progress on carotenoids
On Wednesday afternoon we transformed pPCB15 (CrtEXYIB, producing beta-carotene) into MG1655 ("Parent"), MG1655 PT5-dxs,idi,ispBDF ("Roche") and TOP10. Results of overnight culture are as follows:
(photos to be uploaded)
All three strains show yellowish/orange pigmentation, indicating production of beta-carotene. We are confident that the Crt enzymes (E, B, I, Y) can function normally in E. coli, and the rate of pigment formation is fast (overnight). Previously, K152005 (CrtEBIY, without promoter) took about 3 days for yellowish/orange pigment to develop, which might be due to the absence of promoter. To confirm this, we will put K152005 under strong promoter (R0011, constitutive), transform the construct into E.coli TOP10 and the two MG1655 strains, and see if there is improvement in the rate of pigment formation.
Preparation for tomorrow:
1. Overnight cultures (5ml each) for K118014 (rbs+CrtE), K118006 (rbs+CrtB), K118005 (rbs+CrtI), K118013 (rbs+CrtY), K152005 (CrtEBIY), R0011, and pPCB15, all in TOP10, with appropriate antibiotics (Amp for biobricks, Cam for pPCB15). For mini-prep tomorrow.
2. Overnight cultures (10ml each) for pPCB15 in MG1655, pPCB15 in MG1655 PT5-dxs,idi,ispBDF, pPCB15 in TOP10, and R0011 in TOP10 (as control). For carotene measurement: try out both plate reader and acetone extraction* (*:need more reading on acetone extraction).
Friday
Dry work
The Big Idea
Developed an outline for our system as a device made up of three parts - a sensor, an adaptor, and a pigment producer. Our idea is to develop different bacterial strains that are sensitive to different concentrations of an inducer. Certain concentrations of the inducer will cause violet pigment production, for example, while other concentrations will cause cyan pigment production. This overall aim requires further attention as to how we will develop multiple sensitivities.
Synthesis and primers
Finalised the sequence for our melanin biobrick on the ApE plasmid editor. Pictorial version shown below:
Restriction sites are all within the registry prefix and suffix and will be used to join the gene to the plasmid backbone.
- Primers have been ordered from Invitrogen and should get here next week.
Also examined the vio genes using the Gene-designer from DNA 2.0 with the aim of optimising them for synthesis. As the vio genes are origonally from the chromobacterium we looked at optimising codon usage for E. coli as well as adding a B0034 ribosome binding site to the beginning of each gene.
Modelling
During the course of this week we have been reading about mathematical modelling of gene circuits. We now have some useful books and several papers. We are looking at using either Matlab, or software related to the Systems Biology Workbench. We may well use Cell Designer to work on our circuits graphically.
We would like to model the amplifier such that we are able to predict transient and steady state responses to various inputs. The amplifier, at a simple level, takes a Polymerase per Second (PoPS) signal input, and produces a larger signal as an output, but this will have an inherent time lag. We are also expecting the amplifier's steady state behaviour not to be linear. This however, could be useful if it introduces semi-discrete behaviour.
We would like to also model the pigment production response curves to a PoPS input at various levels. For this, we would need to look beyond molecular modelling, towards the optical properties of the different pigments at various concentrations.
The nature of biological modelling means that we will introduce assumptions at certain stages in the models, but we should have an idea of what impact these assumptions have. Modelling our iGEM project has the potential to be very interesting!
Wet work
Amplifiers
Transformed the following biobricks from Cambridge 2007 into Top10 competent bacteria:
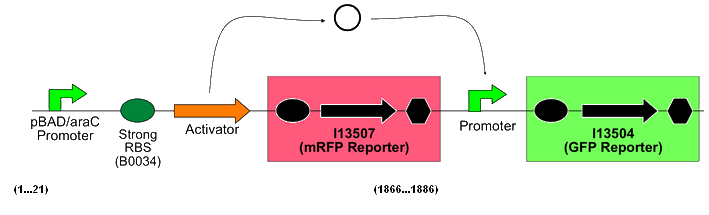
Each of these biobricks is a composite construct in a high copy plasmid vector that follows the following format:
(from the Cambridge 2007 iGEM team's wiki, [http://parts.mit.edu/igem07/index.php/Cambridge/Amplifier_project]
Plated neat and 1/10 dilutions onto amp plates.
Melanin
Plated melanin bacteria on the following plates:
- 100ug/ml Ampicillin, 7.5 ug/ml CuSO4, 0.075 mg/mL tyrosine (my bad, meant to be double that)
- 100ug/ml Ampicillin, 15 ug/ml CuSO4, 0.3 mg/mL tyrosine
- 100ug/ml Ampicillin, 15 ug/ml CuSO4, 0.6 mg/mL tyrosine
- 100ug/ml Ampicillin, 7.5 ug/ml CuSO4, 0.075 mg/mL tyrosine, 0.5mM IPTG (my bad, meant to be double that)
- 100ug/ml Ampicillin, 15 ug/ml CuSO4, 0.3 mg/mL tyrosine, 1mM IPTG
- 100ug/ml Ampicillin, 15 ug/ml CuSO4, 0.6 mg/mL tyrosine, 1mM IPTG
Also, left one 100ug/ml Ampicillin, 15 ug/ml CuSO4, 0.3 mg/mL tyrosine plate out, without any bacteria, just to ensure that the colouration is due to tyrosinase action and not natural oxidation of any of the agar supplements.
Finally, incubated a melanin colony in 5 mL of 100ug/ml Ampicillin, 15 ug/ml CuSO4, 0.3 mg/mL tyrosine, 1mM IPTG over the weekend in preparation for pigment characterisation using the plate reader.
Weekend
Checked to see how the amplification project transformations went: all but I1746570, I13540, and I13520 were growing.
Additionally, the melanin plates were growing rather poorly, probably because they were inoculated with an overnight culture that was a few days old.
Primer order conformation recieved from Invitrogen
 "
"