Team:Cambridge/Notebook/Week5
From 2009.igem.org
(→Violacein gene) |
(→Violacein) |
||
| Line 107: | Line 107: | ||
Removed restriction sites from the new vio plan (shown on yesterdays page) and send it to Jim, along with the ApE plasmid editor file of the operon. After getting the go-ahead from Jim, this plan will be send to DNA2.0 for synthesis. | Removed restriction sites from the new vio plan (shown on yesterdays page) and send it to Jim, along with the ApE plasmid editor file of the operon. After getting the go-ahead from Jim, this plan will be send to DNA2.0 for synthesis. | ||
| + | *Wrote out the design criteria for Violacein synthesis | ||
===Wet Work=== | ===Wet Work=== | ||
Revision as of 14:37, 12 August 2009
Categories :
Project :
-
Overview
Sensitivity Tuner
--- Characterisation
--- Modelling
Colour Generators
--- Carotenoids (Orange/Red)
--- Melanin (Brown)
--- Violacein (Purple/Green)
The Future
Safety
Notebook :
Team Logistics :
Week 5
Monday
Dry Work
Violacein Synthesis
Used GeneDesigner from DNA2.0 to plan what we wanted synthesised, taking into account the different genes in the operon. This would be synthesised with codon optimisation for E. coli. All forbidden restriction sites have been removed (EcoRI, XbaI, SpeI, PstI, NotI)
Wet Work
Protocol de-bugging
For the amplification project, we need to amplify pSB3K3 as a low-copy number plasmid vector in order to transfer the activator constructs into low copy number plasmid. We tried 4 different PCR reaction mixtures, and used a Zymogen PCR wash kit before running them on an 0.8% Agarose E-gel. After looking at the gel, the ladder was quite smeared and no PCR product was visible. It was found that the samples run on the gel had a very low DNA concentration. However, some PCR solution was left before the wash kit was used, and using the Nano-Drop it was determined that it had a high concentration of DNA, so the PCR worked, we just need to refine the protocols after PCR.
Amplification
Mini-prepped I746374, I746375, I746380, and I746391; incubated overnight cultures of the rest of the 11 activator constructs in preparation for miniprep.
Pigments
Melanin
Retransformed melanin plasmid into Top10, and transformed it into MG1655 to (hopefully) achieve better expression. Plated on plates optimized for pigment production, and prepared for solid-phase pigment extraction
Violacein
Retransformed violacein plasmid into Top10, and transformed it into MG1655 to (hopefully) achieve better expression.
Mixing pigments
Plated Top10 transformed with melanin plasmid and violacein plasmid and the Roche strain transformed with Duncan’s orange plasmid on the same plate, optimized for melanin production (as this is the only pigment that requires media supplementation) but without any antibiotic resistance.
Progress on Carotenoids (Monday)
Today we carried out restriction digest of K118006/CrtB (as recipient plasmid vector) using SpeI and PstI (two cuts both within suffix), and of K118005/CrtI (as gene insert) using XbaI and PstI (one cut within prefix, the other cut within suffix).
Recall that the sizes are: K118006/CrtB: insert: 949bp, backbone: 2079bp, total: 3028bp. K118005/CrtI: insert: 1498bp, backbone: 2079bp, total: 3577bp.
Hence we expect one band of about 3000bp for K118006/CrtB, and two bands of about 1500bp and 2100bp for K118005/CrtI, after restriction digest with their respective enzymes.
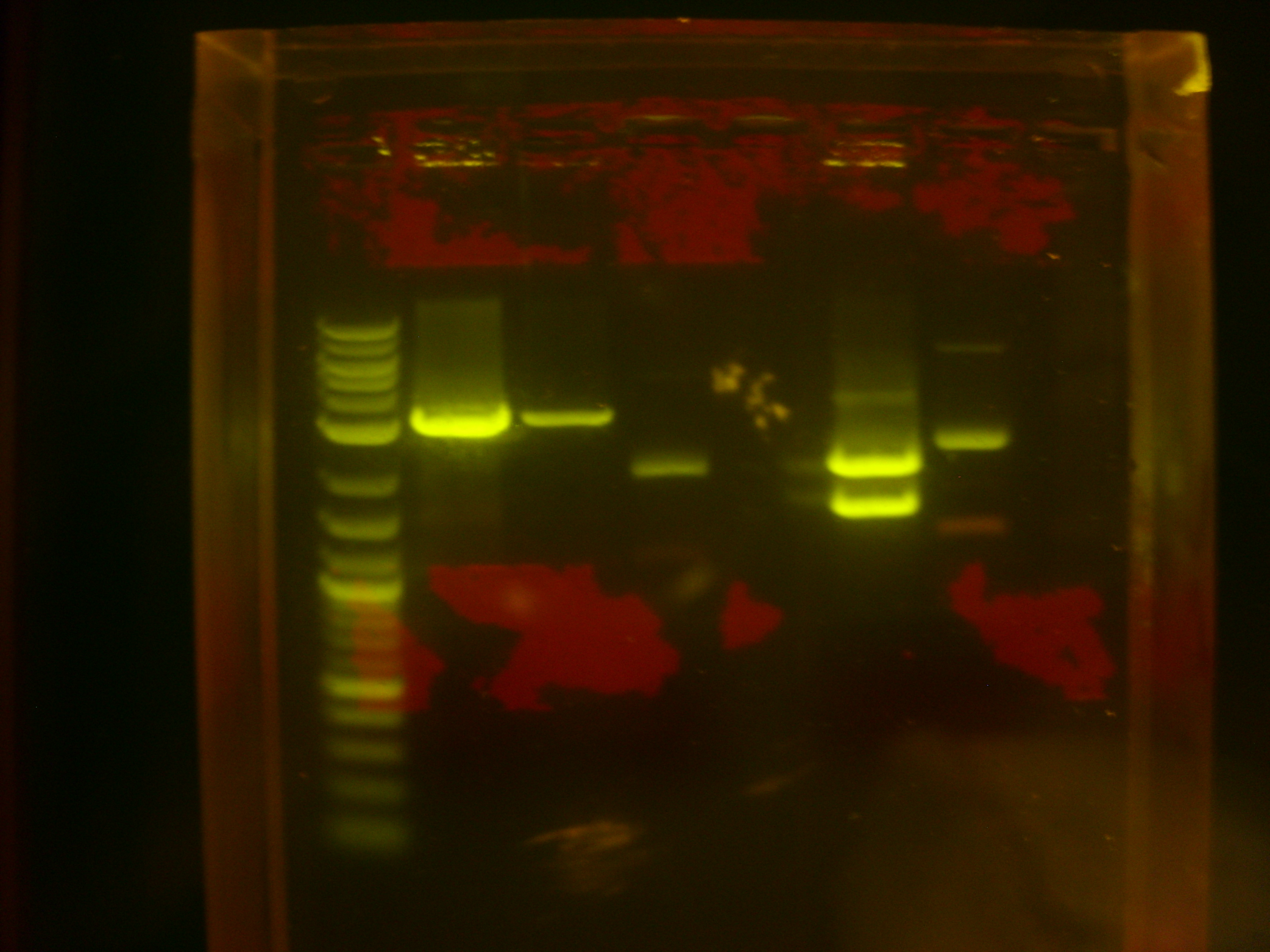
The restriction digest products were run on gel, and the results were shown below:
Lane 1: Log-2 DNA ladder (NEB). Lanes 2 and 3: Restriction digest products of K118006/CrtB (less DNA loaded in lane 3). Lane 4: Undigested K118006/CrtB (negative control). Lane 5 empty. Lane 6: Restriction digest products of K118005/CrtI. Lane 7: Undigested K118005/CrtI (negative control).
This indicated that the restriction digest was successful.
We then carried out gel extraction on Lane 2 and Lane 6, ligation with T4 ligase, and electroporation into E. coli Top10 cells. Results of transformation will be ready for inspection tomorrow.
Tuesday
Dry Work
Violacein gene
Confirmed that correct protein sequence was present for each gene, by using MUSCLE to compare the gene sequence given by GeneDesigner to the sequence in the NCBI database. All protein sequences were identical.
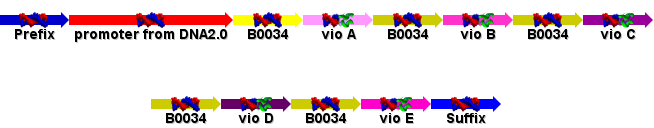
Prepared the GeneDesigner file and the ApE file of the violacein operon to be sent for sequencing. Jim suggested that we refactor the DNA so that each individual ORF is preceeded by an RBS (the biobrick part B0034) and a constituitive promoter added. DNA2.0 kindly agreed to provide the promoter sequence for us, and add it in along with the operon. We also included the vio E gene after reading the Sanchez paper: http://www3.interscience.wiley.com/cgi-bin/fulltext/112732008/HTMLSTART?CRETRY=1&SRETRY=0
- New Vio Plan
Wet Work
Amplification
Continued to refine protocols for successful PCR and purification of pSB3K3.
Miniprepped all remaining amplifier constructs.
Pigments
Yesterdays transformations were successful, continued to incubate to trigger pigment production.
Pics to be put in the right places
Wednesday
Dry Work
Pseudomonas Green and Red
Received confirmation that we will be receiving clones containing the appropriate plasmids (YAY) soon.
Violacein
Removed restriction sites from the new vio plan (shown on yesterdays page) and send it to Jim, along with the ApE plasmid editor file of the operon. After getting the go-ahead from Jim, this plan will be send to DNA2.0 for synthesis.
- Wrote out the design criteria for Violacein synthesis
Wet Work
Pigments
Retransformed melanin and violacein pigment into MG1655 (previous attempt was unsuccessful)
Amplification
Overnight culture of a single colony of J69591, phenotypically confirmed, in preparation to make glycerol stocks, steak out on a plate, and miniprep. This will be the parent colony for all of our characterizations.
 "
"