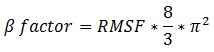Team:EPF-Lausanne/Analysis
From 2009.igem.org
(→Maxwell-Boltzmann Energy Distribution) |
(→Maxwell-Boltzmann Energy Distribution) |
||
| Line 79: | Line 79: | ||
Here we will confirm that the kinetic energy distribution of the atoms in a system corresponds to the Maxwell distribution for a given temperature. | Here we will confirm that the kinetic energy distribution of the atoms in a system corresponds to the Maxwell distribution for a given temperature. | ||
<br> | <br> | ||
| - | + | <br> | |
<b>1.</b> In VMD, load the ''.psf file''. Browse for the restart velocity file (in our case, it was ''2v0w_hydr_wb_i_eq.restart.vel''), the type of file need this time to be selected. In the Determine file type: pull-down menu, choose NAMD Binary Coordinates, and load again. | <b>1.</b> In VMD, load the ''.psf file''. Browse for the restart velocity file (in our case, it was ''2v0w_hydr_wb_i_eq.restart.vel''), the type of file need this time to be selected. In the Determine file type: pull-down menu, choose NAMD Binary Coordinates, and load again. | ||
<br> | <br> | ||
| Line 85: | Line 85: | ||
<br><br> | <br><br> | ||
<b>2.</b> We create a selection with all the atoms in the system, by typing in the TkCon window: | <b>2.</b> We create a selection with all the atoms in the system, by typing in the TkCon window: | ||
| - | + | <br> | |
| - | Open a file energy.dat for writing: | + | <Q> set all [atomselect top all] </Q> |
| + | <br> Open a file energy.dat for writing: | ||
: set fil [open energy.dat w] | : set fil [open energy.dat w] | ||
Revision as of 09:12, 8 September 2009
Contents |
Scripts
As this page is getting crowded, we created another page to explain all the scripts we wrote. The current page has some kind of step by step tutorials, but if you want fast informations, you better go to the script page.
Examples
Maxwell-Boltzmann Energy Distribution
Energies
Here we will calculate the averages of the various energies (kinetic and the different internal energies: bonded (bonds, angles and dihedrals) and non-bonded (electrostatic, van der Waals)) over the course of a simulation.
1. We start with a file obtained from NAMD: http://www.ks.uiuc.edu/Research/namd/utilities/ and download namdstat.tcl
2. In the VMD TkCon window, type
- source namdstats.tcl
- data avg ../namd_log 101 last
The second line will call a procedure which will calculate the average for all output variables in the log file from the first logged timestep after 101 to the end of the simulation.
Temperature distribution
Objective: Simulate our protein in an NVE ensemble, and analyze the temperature distribution.
In order to obtain the data for the temperature from the log file we will again use the script namdstats.tcl, which was already sourced. Type in a terminal window:
- data_time TEMP namd_log
It will store each timestep and its corresponding temperature in the file TEMP.dat.
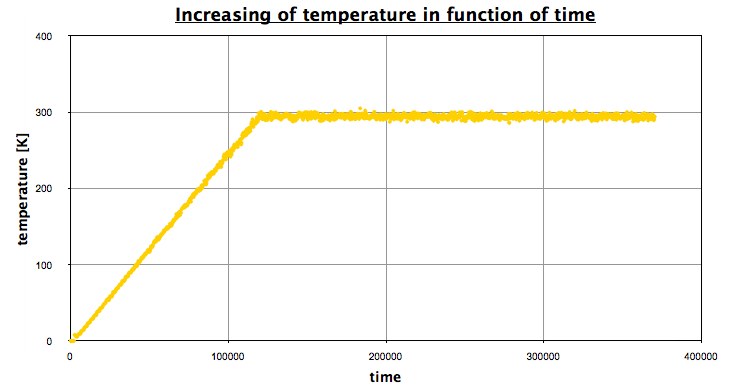
Using EXCEL, we obtain the following graph, which represents the evolution of the temperature in function of time:
The first part corresponds the the heating, then we let the system reach an equilibrium (NPT state), a NVT portion, and finally a NPT portion again.
Density
In order to obtain the data for the volume from the log file we will again use the script namdstats.tcl, which was already sourced. Type in a terminal window:
- data_time VOLUME namd_log
It will store each timestep and its corresponding temperature in the file TEMP.dat.
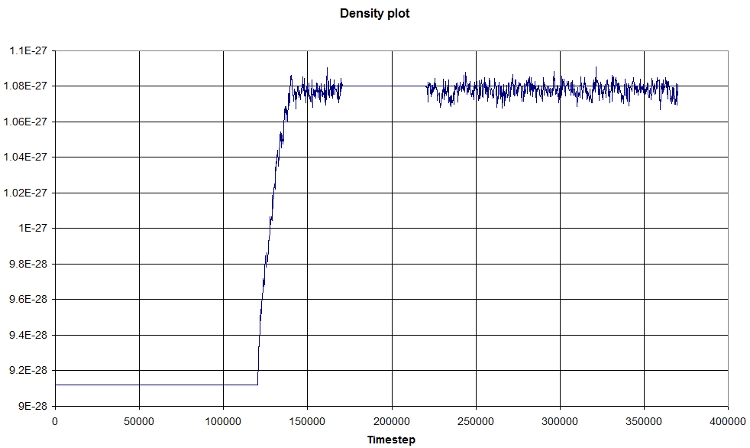
Using EXCEL, we obtain the following graph, which represents the evolution of the density in function of time:
The first part corresponds the the heating, then we let the system reach an equilibrium (NPT state), a NVT portion, and finally a NPT portion again.
Pressure as a function of simulation time
In order to obtain the data for the pressure from the log file we will again use the script namdstats.tcl, which has been updated for the occasion. The file can be found here. Please rename to namdstats.tcl after download.
Here are the steps to use this script:
- source namdstats.tcl
- data_time DATA_NAME LOG_FILE
It will extract data from LOG_FILE, creating a DATA_NAME.dat containing values and time informations.
Available DATA_NAME can be: BOND, ANGLE, DIHED, IMPRP, ELECT, VDW, BOUNDARY, MISC, KINETIC, TOTAL, TEMP, TOTAL2, TOTAL3, TEMPAVG, PRESSURE, GPRESSURE, VOLUME, PRESSAVG, GPRESSAVG.
So, to extract pressure from our first simulation, the command is: data_time PRESSURE namd_log
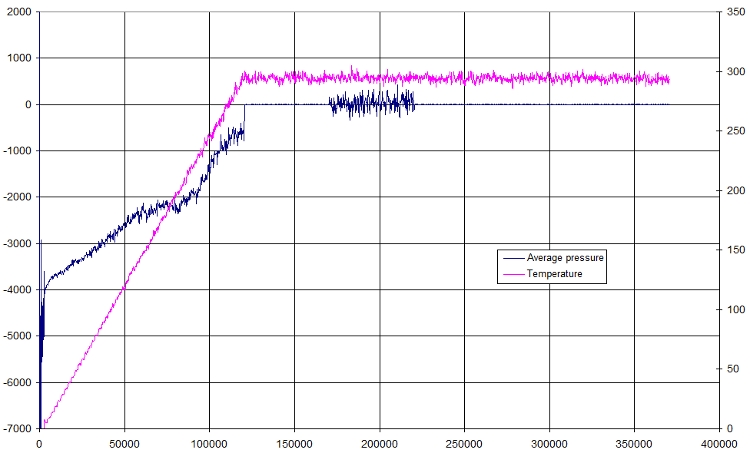
Here is a small plot of pressure and temperature in function of time
RMSD for individual residues
We aim at finding the average RMSD over time for each residue in the protein using VMD.
1. We load the .psf and .dcd files obtained after the first round of simulation.
2. In the TkCon window type:
- source residue_rmsd.tcl
- set sel_resid [[atomselect top "protein and alpha"] get resid]
This will get all the residues number of all alpha-carbons in the protein. Since there is just one and only one α-carbon per residue, it is a good option.
3. Now we will calculate the RMSD values of all atoms in the newly created selection:
- rmsd_residue_over_time top $sel resid
At the end of the calculation, we have a list of the avergae RMSD per residue (in the file residue_rmsd.dat)
- Remark: we updated the script residue_rmsd.tcl to be able to specify on which frames the rsmd has to be computed. Please have a look on this wiki for a more up-to-date version of the file... Command is:
- rmsd_residue_over_time top $sel resid FIRST_FRAME LAST_FRAME
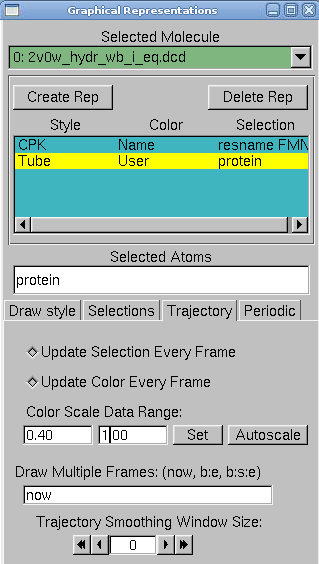
4. In VMD, in Graphics → Representations, do the following actions:
- create two replicates: protein and resname FMN in selected atoms
- for the FMN, choose CPK as a drawing method
- for the protein, choose tube as drawing method, and user as coloring method. Now click on the Trajectory tab, and in the Color Scale Data Range, type 0.40 and 1.00.
The protein is colored according to its average RMSD values. The residues displayed in blue are more mobile while the ones in red move less.
Here is a movie with the protein colored according to average RMSD values.
5. Now we can plot the RMSD value per residue by typing in a Terminal window :
- xmgrace residue rmsd.dat
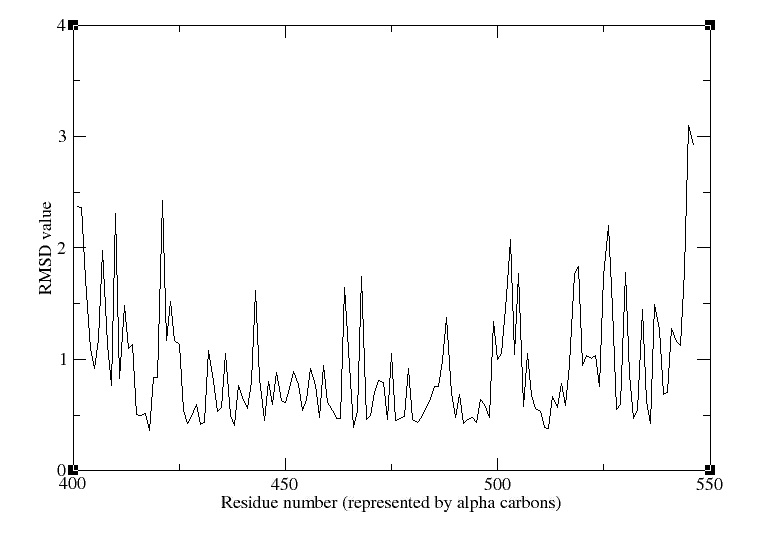
We obtain the following picture:
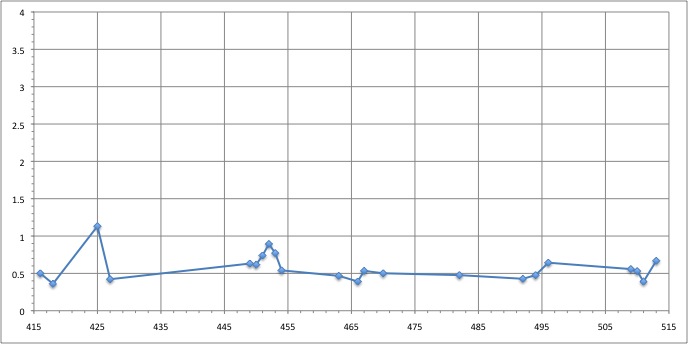
RMSD of residue within 3 angström of the FMN
We can see that the residues that move the most are the residue number: 425, 451, 453
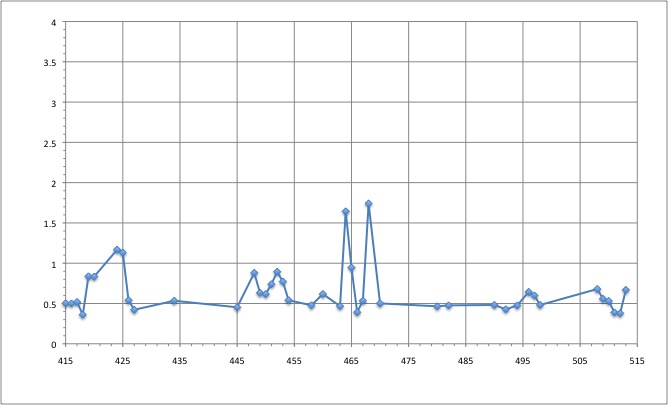
RMSD of residue within 6 angström of the FMN
We can see that the residues that move the most are the residue number: 424, 425, 464, 468
RMSD of selected atoms compared to initial position along time
This script was highly updated, please go to the script page if you encounter a problem!!!
Selections are not precise here!
We made a small TCL script to calculate RMSD from selected atoms compared to their initial position along timestep.
The file can be found here. Please rename to Residue_rmsd_igem09.tcl after download.
Example to run the script:
- load .dcd + .psf on VMD
- source residue_rmsd_igem09.tcl
- set sel_resid [[atomselect top "protein and alpha"] get resid]
- rmsd_residue_over_time top $sel_resid 0 0
We tried to select only backbone from protein + FMN → change
- set sel_resid [[atomselect top "backbone"] get resid]
The script was updated to be able to define reference frame and first frame were RMSD will be calculated. We usually don't need to compute RMSD during heating, for instance. RMSD takes a lot of time. In our first run 1 frame = 100 timesteps * 2 fs*timesteps^-1 = 200 fs
complete form for run is:
- rmsd_residue_over_time top $sel_resid FIRST_FRAME REFERENCE_FRAME
For our first run, if we want to select only the 295°K NPT plateau, and set its first frame as reference, we have to launch:
- rmsd_residue_over_time top $sel_resid 1115 1115
Here is how the script processes:
- calculate how many frames are in .dcd
- for each timestep, the script aligns (best fit) the backbone of the protein to the eference position to minimize RMDS. (Test: "and not mass 1,008000" == and noh was added in selection to remove hydrogen)
- for each residue (selected by sel_resid), RMSD is computed and the sum of all RMSD (one for each residue) is stored for current timestep
- script's output is data_rmsd.dat
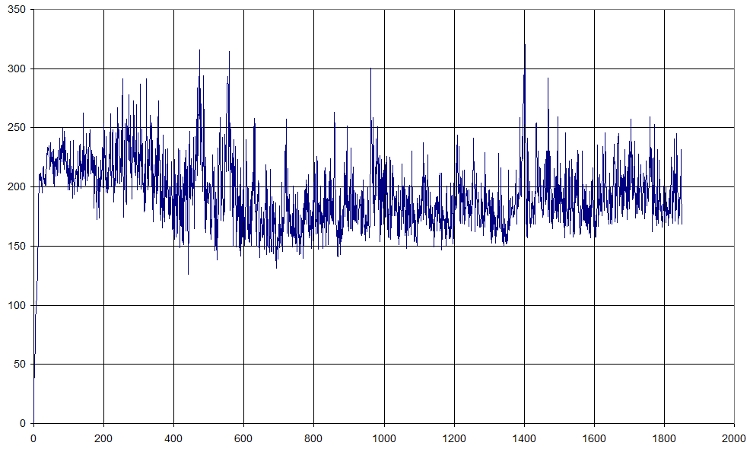
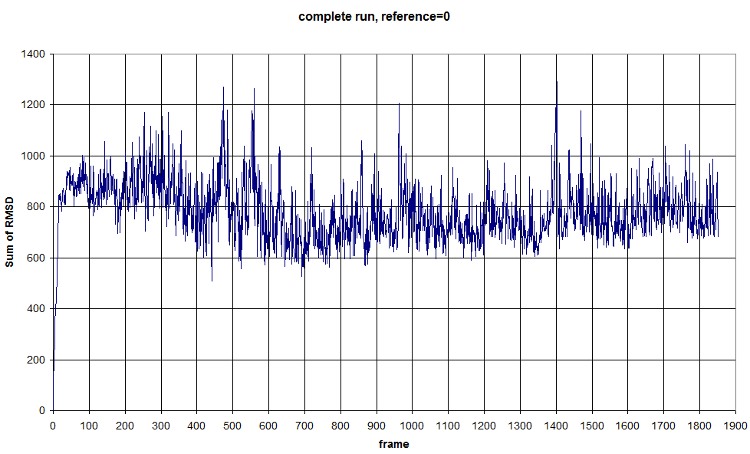
Here is a fast graph of the output of the average RMSD of the atoms in function of time. It seems normal.
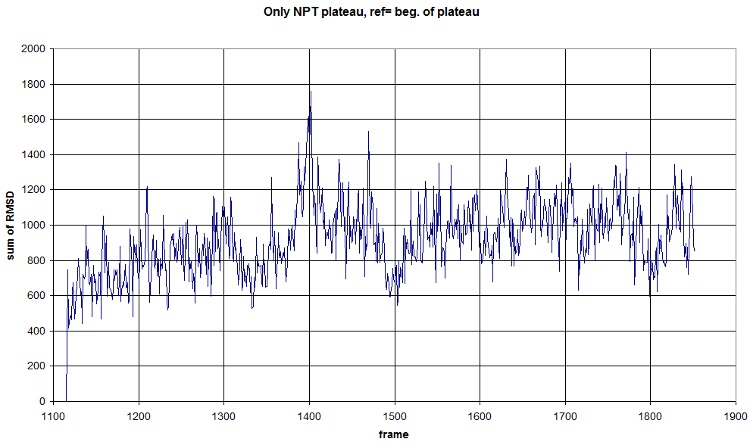
Here is what we got with FIRST_FRAME=1115 REFERENCE_FRAME=1115. Average=921.477, standard deviation=202.1708
FIRST_FRAME=0 REFERENCE_FRAME=0. The difference of the sum probably comes from the new selection of atoms from the backbone. We should compute an average value to normalize amplitude. (fluctuation is conserved, anyway) Average=781.3913, standard deviation=118.1393
Salt bridges
As we wanted to redo the analysis from Schulten's article, we looked for salt bridges. VMD can easily compute this, it even propose an easy GUI. Standard configuration is just fine for now. You'll have a log file containing the list of nitrogen-oxygen susceptible of forming a salt bridge. You'll also get a file for each bridge containing the distance between both atoms along the simulation.
In the light state, we have 9 salt bridges witin the protein and 12 if we consider the protein and the flavin (use "protein or resname FMN" as selection).
- ASP471-ARG467
- GLU409-ARG442
- FMN450-FMN450
- ASP540-LYS544
- ASP432-ARG442
- FMN450-ARG451
- GLU457-LYS489
- GLU444-LYS485
- ASP522-ARG521
- ASP424-ARG451
- GLU475-LYS533
- FMN450-ARG467
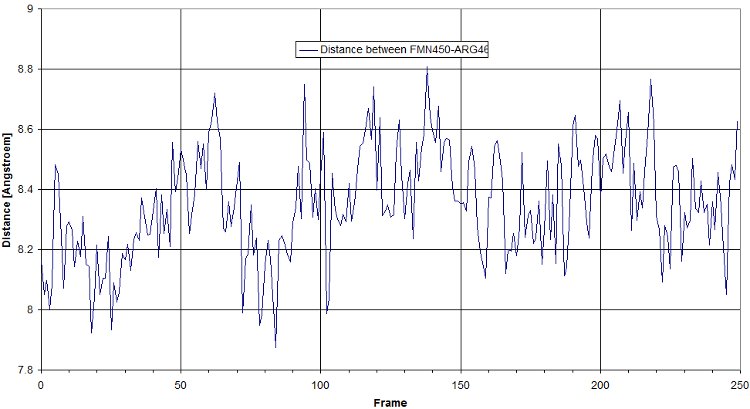
Here is a plot for one of the bridges. We have to look for the max distance for a salt bridge.
RMSF
After changing the script [see here], we perform an interesting analysis from these 2 files. First, we have to correct the RMSF, that can be linked to beta factor using this equation:
If you plot beta factor and RMSF, you get such a thing.
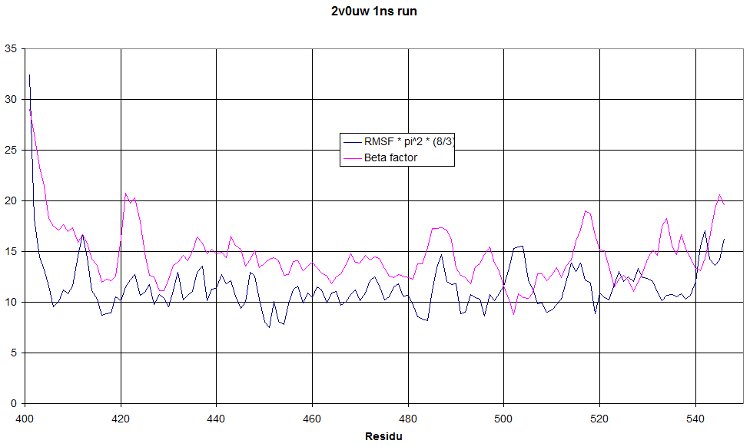
This is a 1 nanosecond NPT run at 300°K. We hope to see a RMSF curve identical to the beta factor. It should only be shifted higher because of the increased temperature. But having a similar tendency would mean our simulation show oscillations similar to what was observed during crystallography. This is really a quite nice validation of our run!
 "
"